Chapter 1: Amino Acids, Peptides, and Proteins

Chapter 1: Amino Acids, Peptides, and Proteins
SCIENCE MASTERY ASSESSMENT
Every pre-med knows this feeling: there is so much content I have to know for the MCAT! How do I know what to do first or what's important?
While the high-yield badges throughout this book will help you identify the most important topics, this Science Mastery Assessment is another tool in your MCAT prep arsenal. This quiz (which can also be taken in your online resources) and the guidance below will help ensure that you are spending the appropriate amount of time on this chapter based on your personal strengths and weaknesses. Don't worry though— skipping something now does not mean you'll never study it. Later on in your prep, as you complete full-length tests, you'll uncover specific pieces of content that you need to review and can come back to these chapters as appropriate.
How to Use This Assessment
If you answer 0–7 questions correctly:
Spend about 1 hour to read this chapter in full and take limited notes throughout. Follow up by reviewing all quiz questions to ensure that you now understand how to solve each one.
If you answer 8–11 questions correctly:
Spend 20–40 minutes reviewing the quiz questions. Beginning with the questions you missed, read and take notes on the corresponding subchapters. For questions you answered correctly, ensure your thinking matches that of the explanation and you understand why each choice was correct or incorrect.
If you answer 12–15 questions correctly:
Spend less than 20 minutes reviewing all questions from the quiz. If you missed any, then include a quick read-through of the corresponding subchapters, or even just the relevant content within a subchapter, as part of your question review. For questions you got correct, ensure your thinking matches that of the explanation and review the Concept Summary at the end of the chapter.
- In a neutral solution, most amino acids exist as:
- positively charged compounds.
- zwitterions.
- negatively charged compounds.
- hydrophobic molecules.
- At pH 7, the charge on a glutamic acid molecule is:
- –2.
- –1.
- 0.
- +1.
- Which of the following statements is most likely to be true of nonpolar R groups in aqueous solution?
- They are hydrophilic and found buried within proteins.
- They are hydrophilic and found on protein surfaces.
- They are hydrophobic and found buried within proteins.
- They are hydrophobic and found on protein surfaces.
- Scientists discover a cDNA sequence for an uncharacterized protein. In their initial studies, they use a computer program designed to predict protein structure. Which of the following levels of protein structure can be most accurately predicted?
- Primary structure
- Secondary structure
- Tertiary structure
- Quaternary structure
- How many distinct tripeptides can be formed from one valine molecule, one alanine molecule, and one leucine molecule?
- 1
- 3
- 6
- 27
- Which of the following best describes the change in entropy that occurs during protein folding?
- Entropy of both the water and the protein increase.
- Entropy of the water increases; entropy of the protein decreases.
- Entropy of the water decreases; entropy of the protein increases.
- Entropy of both the water and the protein decrease.
- An α-helix is most likely to be held together by:
- disulfide bonds.
- hydrophobic effects.
- hydrogen bonds.
- ionic attractions between side chains.
- Which of the following is least likely to cause denaturation of proteins?
- Heating the protein to 100 °C
- Adding 8 M urea
- Moving it to a more hypotonic environment
- Adding a detergent such as sodium dodecyl sulfate
- A particular α-helix is known to cross the cell membrane. Which of these amino acids is most likely to be found in the transmembrane portion of the helix?
- Glutamate
- Lysine
- Phenylalanine
- Aspartate
- Which of these amino acids has a chiral carbon in its side chain?
- Serine
- Threonine
- Isoleucine
- I only
- II only
- II and III only
- I, II, and III
- Following translation and folding, many receptor tyrosine kinases exist as monomers in their inactive state on the cell membrane. Upon the binding of a ligand, these proteins dimerize and initiate a signaling cascade. During this process, their highest element of protein structure changes from:
- secondary to tertiary.
- tertiary to quaternary.
- primary to secondary.
- secondary to quaternary.
- Which of these amino acids has a side chain that can become ionized in cells?
- Histidine
- Leucine
- Proline
- Threonine
- In lysine, the pKa of the side chain is about 10.5. Assuming that the pKa of the carboxyl and amino groups are 2 and 9, respectively, the pI of lysine is closest to:
- 5.5.
- 6.2.
- 7.4.
- 9.8.
- Which of the following is a reason for conjugating proteins?
- To direct their delivery to a particular organelle
- To direct their delivery to the cell membrane
- To add a cofactor needed for their activity
- I only
- II only
- II and III only
- I, II, and III
- Collagen consists of three helices with carbon backbones that are tightly wrapped around one another in a “triple helix.” Which of these amino acids is most likely to be found in the highest concentration in collagen?
- Proline
- Glycine
- Threonine
- Cysteine
Answer Key
- B
- B
- C
- A
- C
- B
- C
- C
- C
- C
- B
- A
- D
- D
- B
Chapter 1: Amino Acids, Peptides, and Proteins
CHAPTER 1
AMINO ACIDS, PEPTIDES, AND PROTEINS
In This Chapter
**1.1 Amino Acids Found in Proteins
**
A Note on Terminology
Stereochemistry of Amino Acids
Structures of the Amino Acids
Hydrophobic and Hydrophilic Amino Acids
1.2 Acid–Base Chemistry of Amino Acids
Protonation and Deprotonation
Titration of Amino Acids
**1.3 Peptide Bond Formation and Hydrolysis
**
Peptide Bond Formation
Peptide Bond Hydrolysis
**1.4 Primary and Secondary Protein Structure
**
Primary Structure
Secondary Structure
**1.5 Tertiary and Quaternary Protein Structure
**
Tertiary Structure
Folding and the Solvation Layer
Quaternary Structure
Conjugated Proteins
1.6 Denaturation Concept Summary
CHAPTER PROFILE
The content in this chapter should be relevant to about 20% of all questions about biochemistry on the MCAT.
This chapter covers material from the following AAMC content categories:
1A: Structure and function of proteins and their constituent amino acids
5D: Structure, function, and reactivity of biologically-relevant molecules
Introduction
How important are amino acids? Consider sickle cell disease. People with sickle cell disease have red blood cells that, under certain conditions, can become rigid and sickle-shaped. Those sickle-shaped cells can become stuck in capillaries, blocking them. In severe cases, it can block enough of the blood supply to cause damage to several organs, such as the kidneys, liver, and spleen. This happens because of a mutation in hemoglobin, the protein in red blood cells that transports oxygen. Remarkably, the difference between the normal form of hemoglobin, HbA, and the one that causes sickle cell disease, HbS, is a seemingly minor one. All it takes is a change in a single amino acid on the surface of hemoglobin: the sixth amino acid in two of its four chains is changed from glutamic acid to valine. That minor difference allows the deoxygenated form of HbS to aggregate and precipitate, which leads to the sickled shape—and all the symptoms—of sickle cell disease.
MCAT EXPERTISE
This chapter represents a whopping 20% of all biochemistry questions you will see on Test Day. That makes amino acids one of the highest yield subjects within any of the review books! Make sure to work sufficient study of these materials into your study plan.
In this chapter, we’ll take a look at the basics of proteins by focusing on the amino acids that compose them and how those amino acids contribute to the physical and chemical properties of proteins.
1.1 Amino Acids Found in Proteins
LEARNING OBJECTIVES
After Chapter 1.1, you will be able to:
- Recognize common abbreviations for amino acids, such as Glu and Y
- Distinguish the stereochemistries, typical cellular locations, and reactivities of the 20 major amino acids
- Identify the major amino acids, such as:
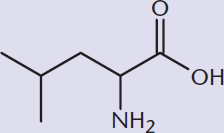
MCAT EXPERTISE
The “High-Yield” badge on this section indicates that the content is frequently tested on the MCAT.
Amino acids are molecules that contain two functional groups: an amino group (–NH2) and a carboxyl group (–COOH). In this chapter, we’ll focus specifically on the α-amino acids, in which the amino group and the carboxyl group are bonded to the same carbon, the α-carbon of the carboxylic acid. Think of the α-carbon as the central carbon of the amino acid, as shown in Figure 1.1.
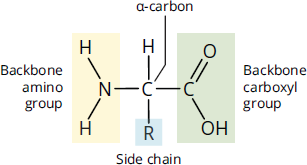
Figure 1.1 Amino Acid Structure
BRIDGE
Remember that for carboxylic acids, the α-carbon is the carbon adjacent to the carboxyl carbon. The structure and nomenclature of carboxylic acids are discussed in Chapter 8 of MCAT Organic Chemistry Review.
In addition to the amino and carboxyl groups, the α-carbon has two other groups attached to it: a hydrogen atom and a side chain, also called an R group, which is specific to each amino acid. The side chains determine the properties of amino acids, and therefore their functions.
KEY CONCEPT
The side chains (R groups) of amino acids determine their chemical properties.
A Note on Terminology
Amino acids do not need to have both the amino and carboxyl groups bonded to the same carbon. For example, the neurotransmitter *γ**-aminobutyric acid(GABA) has the amino group on the gamma (γ) carbon,threecarbons away from the carboxyl group. Similarly, not every amino acid found in the human body is specified by a codon in the genetic code or incorporated into proteins. One example isornithine*, one of the intermediates in the urea cycle, the metabolic process by which the body excretes excess nitrogen. There are also some amino acids that are specifically modified for specialized roles in the body; for example, lysine is sometimes converted into pyrrolysine.
That said, the Association of American Medical Colleges (AAMC) has specifically stated they’ll focus on the 20 α-amino acids encoded by the human genetic code, also called proteinogenic amino acids. So, for the rest of this chapter, we’ll use the term amino acid to refer specifically to these compounds.
MCAT EXPERTISE
While the MCAT could include passages on GABA or other nonstandard amino acids, they are not part of the background information you’re expected to know for Test Day.
Stereochemistry of Amino Acids
For most amino acids, the α-carbon is a chiral (or stereogenic) center, as it has four different groups attached to it. Thus, most amino acids are optically active. The one exception is glycine, which has a hydrogen atom as its R group, making it achiral, as shown in Figure 1.2.
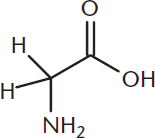
Figure 1.2 Glycine
All chiral amino acids used in eukaryotes are L-amino acids, so the amino group is drawn on the left in a Fischer projection, as demonstrated in Figure 1.3.
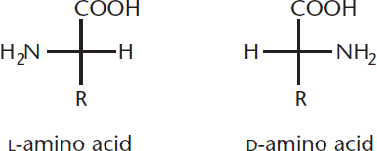
Figure 1.3 L- and D-Amino Acids
KEY CONCEPT
Except for glycine, all amino acids are chiral—and except for cysteine, all of them have an (S) absolute configuration.
REAL WORLD
While L-amino acids are the only ones found in eukaryotic proteins, D-amino acids do exist. One example is gramicidin, an antibiotic produced by a soil bacterium called Bacillus brevis, in which D- and L-amino acids alternate in the primary structure.
In the Cahn–Ingold–Prelog system, this translates to an (S) absolute configuration for almost all chiral amino acids. The only exception is cysteine, which, while still being an L-amino acid, has an (R) absolute configuration, as shown in Figure 1.4, because the –CH2SH group has priority over the –COOH group.
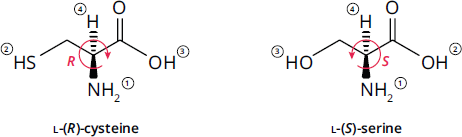
Figure 1.4 Cysteine Has an (R) Absolute Configuration
Structures of the Amino Acids
There are several ways to classify amino acids. In this section, we’ll break them down by the structures of their side chains.
Nonpolar, Nonaromatic Side Chains
Seven amino acids, shown in Figure 1.5a, fall into this class. Glycine, discussed earlier, has a single hydrogen atom as its side chain and is therefore achiral. It is also the smallest amino acid. Four other amino acids—alanine, valine, leucine, and isoleucine—have alkyl side chains containing one to four carbons.
MCAT EXPERTISE
The AAMC’s practice materials have made it clear that test takers are expected to know the structures, names, and three-letter and one-letter abbreviations for the amino acids.
Methionine is one of only two amino acids that contains a sulfur atom in its side chain. Nevertheless, because the sulfur has a methyl group attached, it is considered relatively nonpolar.
Finally, proline is unique in that it forms a cyclic amino acid. In all the other amino acids, the amino group is attached only to the α-carbon. In proline, however, the amino nitrogen becomes a part of the side chain, forming a five-membered ring. That ring places notable constraints on the flexibility of proline, which limits where it can appear in a protein and can have significant effects on proline’s role in secondary structure.
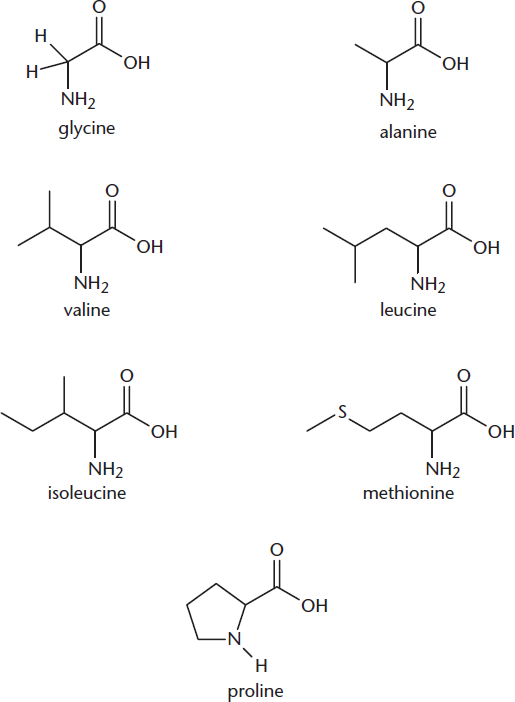
Figure 1.5a Amino Acids with Nonpolar, Nonaromatic Side Chains
MCAT EXPERTISE
The MCAT writers strive to avoid misuse of terminology. Some textbooks problematically describe proline as an imino acid because the amino nitrogen forms two bonds to carbon. The MCAT won’t use this term because an imine is specifically a molecule with a carbon–nitrogen double bond.
Aromatic Side Chains
Three amino acids have uncharged aromatic side chains and are depicted in Figure 1.5b. The largest of these is tryptophan, which has a double-ring heterocyclic system called an indole that contains a nitrogen atom. The smallest is phenylalanine, which has a benzyl side chain (a benzene ring plus a –CH2– group). Adding an –OH group to phenylalanine gives the third member, tyrosine. The –OH group makes tyrosine relatively polar compared to phenylalanine, though still net nonpolar.
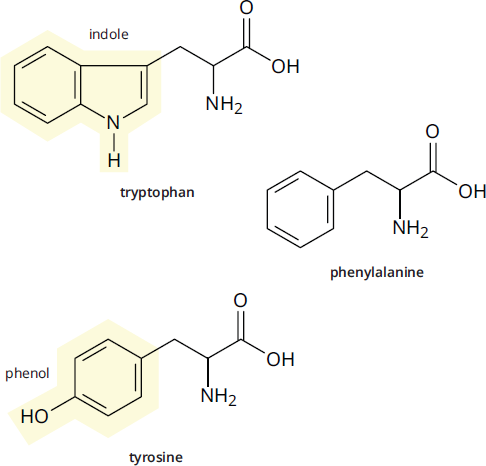
Figure 1.5b Amino Acids with Aromatic Side Chains
Polar Side Chains
Five amino acids, shown in Figure 1.5c, have side chains that are polar but not aromatic. Serine and threonine both have –OH groups in their side chains, which makes them highly polar and able to participate in hydrogen bonding. Asparagine and glutamine have amide side chains. Unlike the amino group common to all amino acids, the amide nitrogens donot gain or lose protons with changes in pH; they do not become charged.
BRIDGE
Make sure you know your carboxylic acid derivatives for Test Day! They are discussed in Chapter 9 of MCAT Organic Chemistry Review.
The last amino acid with a polar side chain is cysteine, which has a thiol (–SH) group in its side chain. Because sulfur is larger than oxygen, the S–H bond is longer and weaker than the O–H bond. In addition, sulfur is more electronegative than oxygen. This leaves the thiol group in cysteine prone to oxidation, a reaction we’ll study later in this chapter.
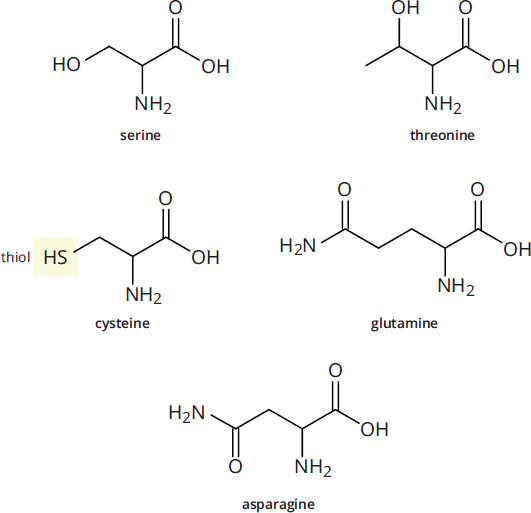
Figure 1.5c Amino Acids with Polar Side Chains
Negatively Charged (Acidic) Side Chains
Only two of the 20 amino acids have negative charges on their side chains at physiological pH (7.4). Those two are aspartic acid (aspartate), which is related to asparagine, and glutamic acid (glutamate), which is related to glutamine. Unlike asparagine and glutamine, aspartate and glutamate have carboxylate (–COO–) groups in their side chains, rather than amides. Note that aspartate is simply the deprotonated form of aspartic acid, and glutamate is the deprotonated form of glutamic acid. These two amino acids are depicted in Figure 1.5d.
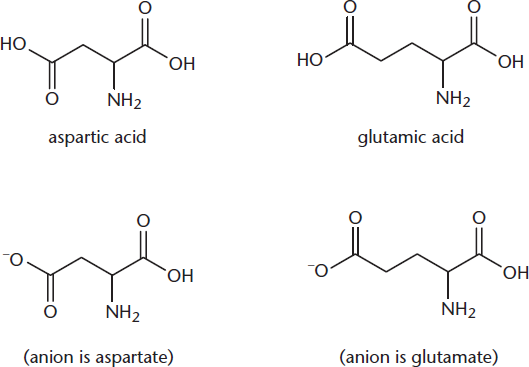
Figure 1.5d Amino Acids with Negatively Charged Side Chains
MCAT EXPERTISE
On the MCAT, you’re likely to see anion names (aspartate and glutamate) instead of acid names (aspartic acid and glutamic acid) because most acids in cells exist in the deprotonated form. This also applies to molecules other than amino acids (malate instead of malic acid).
Positively Charged (Basic) Side Chains
The remaining three amino acids, shown in Figure 1.5e, have side chains that have positively charged nitrogen atoms. Lysine has a terminal primary amino group, while arginine has three nitrogen atoms in its side chain, forming a guanidine; the positive charge is delocalized over all three nitrogen atoms. The final amino acid, histidine, has an aromatic ring with two nitrogen atoms (this ring is called an imidazole). You might be wondering how histidine can acquire a positive charge. The pKa of the side chain is relatively close to 7.4—it’s about 6—so, at physiologic pH, one nitrogen atom is protonated and the other isn’t. Under more acidic conditions, the second nitrogen atom can become protonated, giving the side chain a positive charge.
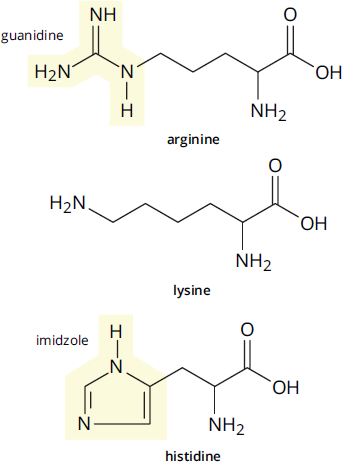
Figure 1.5e Amino Acids with Positively Charged Side Chains
BRIDGE
The catalytic triad found in chymotrypsin makes use of the histidine side chain’s ability to gain a proton. The histidine residue in its active site removes a proton from the –OH group of a serine residue side chain, which facilitates nucleophilic attack by the serine residue on the carbonyl carbon atom of the substrate. Chymotrypsin’s role as a digestive enzyme is discussed in Chapter 9 of MCAT Biology Review.
Hydrophobic and Hydrophilic Amino Acids
Classifying amino acid side chains as hydrophobic or hydrophilic is actually a very complex matter. For example, tyrosine has both an –OH group and an aromatic ring—so which one “wins”?
A few clear conclusions can be drawn, though. First, the amino acids with long alkyl side chains—alanine, isoleucine, leucine, valine, and phenylalanine—are all strongly hydrophobic and thus more likely to be found in the interior of proteins, away from water on the surface of the protein. Second, all the amino acids with charged side chains—positively charged histidine, arginine, and lysine, plus negatively charged glutamate and aspartate—are hydrophilic, as are the amides asparagine and glutamine. The remaining amino acids lie somewhere in the middle and are neither particularly hydrophilic nor particularly hydrophobic.
KEY CONCEPT
The surface of a protein tends to be rich in amino acids with charged side chains. Strongly hydrophobic amino acids tend to be found in the interior of proteins.
Amino Acid Abbreviations
Now that we have explored the structures of the 20 proteinogenic amino acids, it is worth mentioning that you are expected to be able to identify an amino acid on the MCAT not only by name, but also by its three-letter and one-letter abbreviations. These abbreviations are listed in Table 1.1.
MCAT EXPERTISE
It bears repeating: the structures and abbreviations (both three- and one-letter) have been tested heavily on both natural science sections of the MCAT. Make sure you know this page by heart!
Table 1.1 Three- and One-Letter Abbreviations of Amino Acids
AMINO ACID THREE-LETTER ABBREVIATION ONE-LETTER ABBREVIATION
Alanine Ala A
Arginine Arg R
Asparagine Asn N
Aspartic acid Asp D
Cysteine Cys C
Glutamic acid Glu E
Glutamine Gln Q
Glycine Gly G
Histidine His H
Isoleucine Ile I
Leucine Leu L
Lysine Lys K
Methionine Met M
Phenylalanine Phe F
Proline Pro P
Serine Ser S
Threonine Thr T
Tryptophan Trp W
Tyrosine Tyr Y
Valine Val V
Mutagenesis
Mutagenesis describes the process by which mutations are deliberately introduced into a DNA sequence in order to produce a mutant gene or protein. Mutagenesis is commonly used by researchers interested in determining the function of specific amino acids within proteins or of specific nucleotides within DNA or RNA, where the specific sites are mutated to see if the mutation disrupts normal protein or nucleotide function. For instance, a researcher interested in determining if certain amino acids within a protein are phosphorylated may generate mutant proteins that contain alanine mutations (which are not amino acids that undergo phosphorylation) at the putative phosphorylation sites. The researcher can then compare the levels of phosphorylation in the wild-type protein to the mutated proteins. If one of the amino acids that was mutated to an alanine originally was phosphorylated, levels of phosphorylation would be expected to decrease on the mutant protein compared to the wild-type protein. However, if the amino acid that was mutated was never phosphorylated to begin with, there would be no change in the levels of phosphorylation, indicating that amino acid is not a phosphorylation site.
A mainstay of mutagenesis is site-directed mutagenesis, which is a method used to introduce specific mutations into DNA. First, DNA primers are synthesized that contain the mutation of interest. Surrounding the site of mutation are sequences complementary to the template DNA. The primers are then used to introduce the mutation into each newly-synthesized DNA molecule.
As shown in the preceding table, each amino acid has a one-letter code. This one-letter code is commonly used as a shorthand to denote single amino acid mutations: The original amino acid is listed first, which is then followed by a number denoting the position of the mutated amino acid in the primary sequence relative to the N-terminus, before ending with a second letter that denotes the substituted amino acid. For example, S46A indicates that the 46th amino acid, which was originally a serine (S), has been mutated to an alanine (A). This type of shorthand will show up on not just one, but all three, of the science sections of the MCAT on Test Day.
BRIDGE
Solutions to concept checks for a given chapter in MCAT Biochemistry Review can be found near the end of the chapter in which the concept check is located, following the Concept Summary for that chapter.
MCAT CONCEPT CHECK 1.1:
Before you move on, assess your understanding of the material with these questions.
- What are the four groups attached to the central (α) carbon of a proteinogenic amino acid?
________________________________________
- What is the stereochemistry of the chiral amino acids that appear in eukaryotic proteins?
- L or D?
- (R) or (S)? (Exception:)____________
- Which amino acids fit into each of these categories? (Note: The number in parentheses indicates the number of amino acids in that category.)
- Nonpolar, nonaromatic (7):
_________________________________________
- Aromatic (3):
_________________________________________
- Polar (5):
_________________________________________
- Negatively charged/acidic (2):
_________________________________________
- Positively charged/basic (3):
_________________________________________
- Where do hydrophobic amino acids tend to reside within a protein? What about hydrophilic ones?
- Hydrophobic:
_________________________________________
- Hydrophilic:
_________________________________________
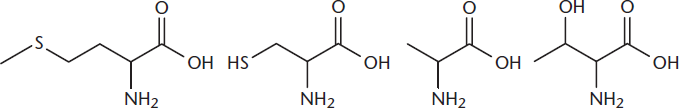
- Identify the amino acids below by their one-letter abbreviation.
1.2 Acid–Base Chemistry of Amino Acids
LEARNING OBJECTIVES
After Chapter 1.2, you will be able to:
- Identify the predominant ion form of the generic amino acid backbone given a pH value
- Calculate pI values for an amino acid given pKa values for backbone and side chains
- Predict the general form of the titration curve for an amino acid
The AAMC loves to use amino acids to test your understanding of acid–base chemistry because they have both an acidic carboxylic acid group and a basic amino group. That makes them amphoteric species, as they can either accept a proton or donate a proton; how they react depends on the pH of their environment. The key to understanding the behavior of amino acids is to remember two facts:
- Ionizable groups tend to gain protons under acidic conditions and lose them under basic conditions. So, in general, at low pH, ionizable groups tend to be protonated; at high pH, they tend to be deprotonated.
- The pKa of a group is the pH at which, on average, half of the molecules of that species are deprotonated; that is, [protonated version of the ionizable group] = [deprotonated version of the ionizable group] or [HA] = [A–]. If the pH is less than the pKa, a majority of the species will be protonated. If the pH is higher than the pKa, a majority of the species will be deprotonated. (These relationships are depicted in Figure 1.6.)
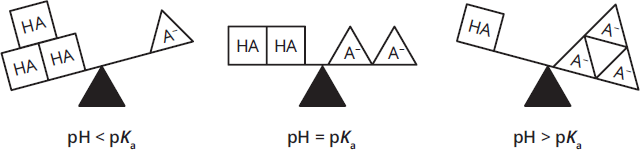
Figure 1.6 Protonation Depends on pH and pKa
MCAT EXPERTISE
You are not expected to know exact values for isoelectric points or side chain pKa values. The MCAT will either provide you with numerical data to use in answering a question or test clear-cut distinctions that can be made without additional information (such as, Which is more hydrophilic, valine or lysine? but not Which is more hydrophobic, valine or alanine?).
Protonation and Deprotonation
Because all amino acids have at least two groups that can be deprotonated, they all have at least two pKa values. The first one, pKa1, is the pKa for the carboxyl group and is usually around 2. For most amino acids, the second pKa value, pKa2, is the pKa for the amino group, which is usually between 9 and 10. For amino acids with an ionizable side chain, there will be three pKa values, but we’ll come back to that later. As an example, let’s take glycine, which doesn’t have an ionizable side chain.
KEY CONCEPT
Like all polyprotic acids, each ionizable proton has its own pKa at which it is “half-deprotonated.”§
Positively Charged under Acidic Conditions
At pH 1 (below even the pH of the stomach), there are plenty of protons in solution. Because we’re far below the pKa of the amino group, the amino group will be fully protonated (–NH3+) and thus positively charged. Because we’re also below the pKa of the carboxylic acid group, it too will be fully protonated (–COOH) and thus neutral. Therefore, at very acidic pH values, amino acids tend to be positively charged, as shown in Figure 1.7.
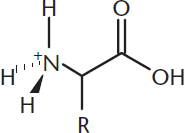
Figure 1.7 Amino Acid Structure at Acidic pH
Zwitterions at Intermediate pH
If we increase the pH of the amino acid solution from pH 1 to pH 7.4, the normal pH of human blood, we’ve moved far above the pKa of the carboxylic acid group. At physiological pH, you will not find amino acids with the carboxylate group protonated (–COOH) and the amino group unprotonated (–NH2). Under these conditions, the carboxyl group will be in its conjugate base form and be deprotonated, becoming –COO–. Conversely, we’re still well below the pKa of the basic amino group, so it will remain fully protonated and in its conjugate acid form (–NH3+). Thus, we have a molecule that has both a positive charge and a negative charge, but overall, the molecule is electrically neutral. We call such molecules dipolar ions, or zwitterions (from the German zwitter, or “hybrid”), as depicted in Figure 1.8. The two charges neutralize one another, and zwitterions exist in water as internal salts.
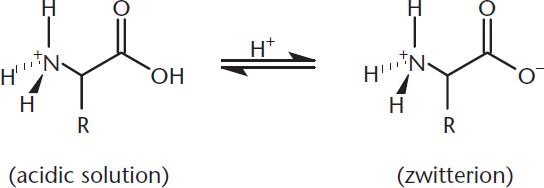
Figure 1.8 Carboxylic Acids Become Deprotonated at Neutral pH, Forming Zwitterions
Negatively Charged under Basic Conditions
Milk of magnesia, which is often used as an antacid, has a pH around 10.5. At that pH, the carboxylate group is already deprotonated and thus remains –COO–. On the other hand, we are now well above the pKa for the amino group, so it deprotonates too, becoming –NH2. So, at highly basic pH, glycine is now negatively charged, as depicted in Figure 1.9.
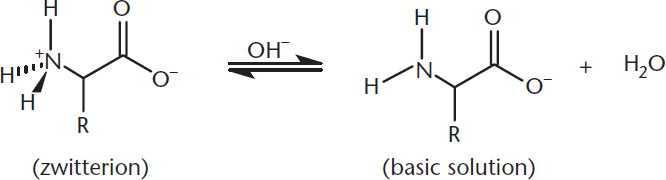
Figure 1.9 Amino Groups Become Deprotonated at Basic pH, Forming an Anion
KEY CONCEPT
At very acidic pH values, amino acids tend to be positively charged. At very alkaline pH values, amino acids tend to be negatively charged.
Titration of Amino Acids
Because of these acid–base properties, amino acids are great candidates for titrations. We assume that the titration of each proton occurs as a distinct step, resembling that of a simple monoprotic acid. Thus, the titration curve looks like a combination of two monoprotic acid titration curves (or three curves, if the side chain is charged). Figure 1.10 shows the titration curve for glycine. After we inspect this curve, we’ll look at the differences for the amino acids with charged side chains.
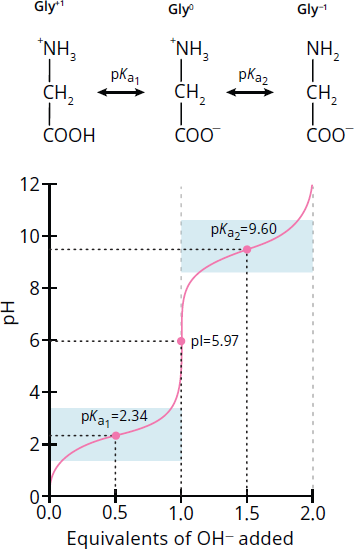
Figure 1.10 Titration Curve for Glycine
Imagine an acidic 1 M glycine solution. At low pH values, glycine exists predominantly as +NH3CH2COOH; it is fully protonated, with a positive charge. As the solution is titrated with NaOH, the carboxyl group will deprotonate first because it is more acidic than the amino group. When 0.5 equivalents of base have been added to the solution, the concentrations of the fully protonated glycine and its zwitterion, +NH3CH2COO–, are equal; that is, [+NH3CH2COOH] = [+NH3CH2COO–]. At this point, the pH equals pKa1. Remember: when the pH is close to the pKa value of a solute, the solution is acting as a buffer and the titration curve is relatively flat, as demonstrated in the blue boxes in the diagram.
KEY CONCEPT
When the pH of a solution is approximately equal to the pKa of the solute, the solution acts as a buffer.
As we add more base, the carboxylate group goes from half-deprotonated to fully deprotonated. The amino acid stops acting like a buffer, and pH starts to increase rapidly during this phase. When we’ve added 1.0 equivalent of base, glycine exists exclusively as the zwitterion form (remember, we started with 1.0 equivalent of glycine). This means that every molecule is now electrically neutral, and thus the pH equals the isoelectric point (pI) of glycine. This is true of all amino acids: the isoelectric point is the pH at which the molecule is electrically neutral. For neutral amino acids, it can be calculated by averaging the two pKa values for the amino and carboxyl groups:
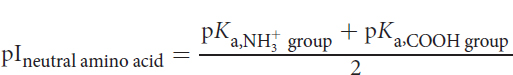
Equation 1.1
For glycine, the pI value is (2.34 + 9.60) ÷ 2 = 5.97. Remember that when the molecule is neutral, it is especially sensitive to pH changes, and the titration curve is nearly vertical.
KEY CONCEPT
When the pH of an amino acid solution equals the isoelectric point (pI) of the amino acid, it exists as electrically neutral molecules. The pI is calculated as the average of the two nearest pKa values. For amino acids with non-ionizable side chains, the pI is usually around 6.
As we continue adding base, glycine passes through a second buffering phase as the amino group deprotonates; again, the pH remains relatively constant. When 1.5 equivalents of base have been added, the concentration of the zwitterion form equals the concentration of the fully deprotonated form; that is, [+NH3CH2COO–] = [NH2CH2COO–], and the pH equals pKa2. Once again, the titration curve is nearly horizontal. Finally, when we’ve added 2.0 equivalents of base, the amino acid has become fully deprotonated, and all that remains is NH2CH2COO–; additional base will only increase the pH further.
Amino Acids with Charged Side Chains
For amino acids with charged side chains, such as glutamic acid and lysine, the titration curve has an extra “step,” but works along the same principles as described above.
Figure 1.11 shows the titration curve for glutamic acid. Because glutamic acid has two carboxyl groups and one amino group, its charge in its fully protonated state at low pHs is +1. As the glutamic acid solution is titrated with NaOH, it undergoes the first deprotonation where, like with glycine, a proton is lost from the main carboxyl group. At that point, the amino acid is electrically neutral with a net charge of 0.
As the pH continues to increase, it loses its second proton so that the net charge of glutamic acid becomes −1, just as it does when glycine loses its second proton. However, unlike glycine, which loses its second proton from the amino group, the second proton that is removed from glutamic acid comes from the side chain carboxyl group! The side chain carboxyl group on glutamic acid is a relatively acidic group, with a pKa of around 4.2. Therefore, the pI of glutamic acid is around 3.2.
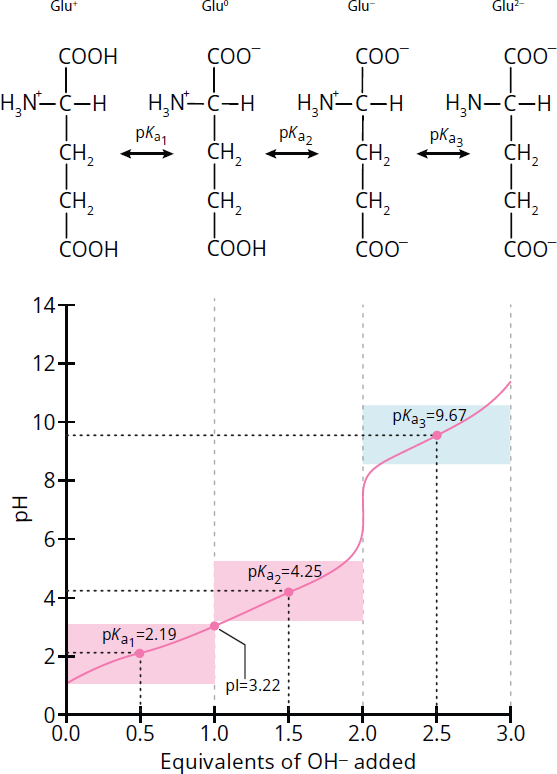
Figure 1.11 Titration Curve for Glutamic Acid
The isoelectric point for an acidic amino acid can be calculated by averaging the pI values of the acidic side chain and carboxylic acid group, as shown below:
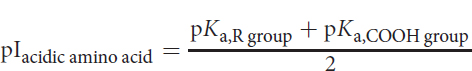
Equation 1.2
Figure 1.12 shows the titration curve for lysine. Lysine has two amino groups and one carboxyl group. Thus, its charge in its fully protonated state is +2, not +1. Losing the carboxyl proton, which still happens around pH 2, brings the charge down to +1. However, lysine does not become electrically neutral until it loses the proton from its main amino group, which happens around pH 9. It gains a negative charge around pH 10.5, at which point it loses the proton on the amino group located on its side chain. Thus, the pI of lysine is around 9.7.
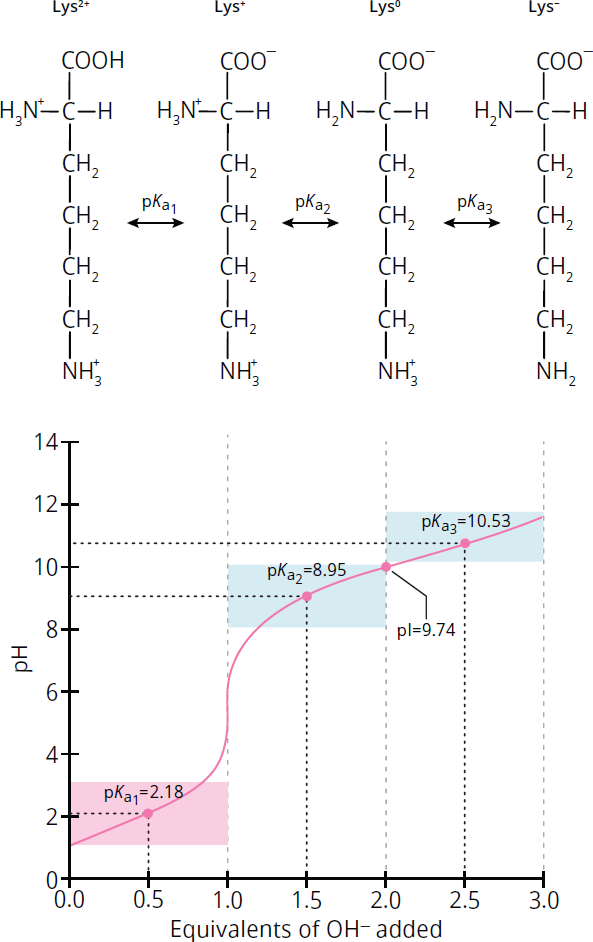
Figure 1.12 Titration Curve for Lysine
The pI for a basic amino acid can be calculated by averaging the pI values of the basic side chain and the amino group, as shown below:
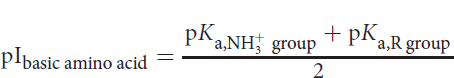
Equation 1.3
KEY CONCEPT
Amino acids with acidic side chains have pI values well below 6; amino acids with basic side chains have pI values well above 6.
The take-home message: amino acids with acidic side chains have relatively low values for pI while those with basic side chains have relatively high ones.
MCAT CONCEPT CHECK 1.2:
Before you move on, assess your understanding of the material with these questions.
- For a generic amino acid, NH2CRHCOOH, with an uncharged side chain, what would be the predominant form at each of the following pH values?
- pH = 1:
_________________________________________
- pH = 7:
_________________________________________
- pH = 11:
_________________________________________
- Given the following pKa values, what is the value of the pI for each of the amino acids listed below?
- Aspartic acid (pKa1 = 1.88, pKa2 = 3.65, pKa3 = 9.60): pI = _________
- Arginine (pKa1 = 2.17, pKa2 = 9.04, pKa3 = 12.48): pI = _________
- Valine (pKa1 = 2.32, pKa2 = 9.62): pI = _________
1.3 Peptide Bond Formation and Hydrolysis
LEARNING OBJECTIVES
After Chapter 1.3, you will be able to:
- Recognize the relationship of nomenclature with length, such as predicting the length of a compound called a “tripeptide”
- Apply the hydrolytic mechanisms of trypsin and chymotrypsin to novel peptide chains
- Predict the products of peptide bond formation and cleavage reactions:
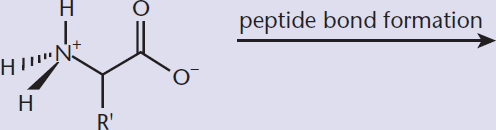
Peptides are composed of amino acid subunits, sometimes called residues, as shown in Figure 1.13. Dipeptides consist of two amino acid residues; tripeptides have three. The term oligopeptide is used for relatively small peptides, up to about 20 residues; while longer chains are called polypeptides.
BRIDGE
For Test Day, you also need to know how peptide bonds are formed in the context of ribosomes, which is covered in Chapter 7 of MCAT Biochemistry Review.
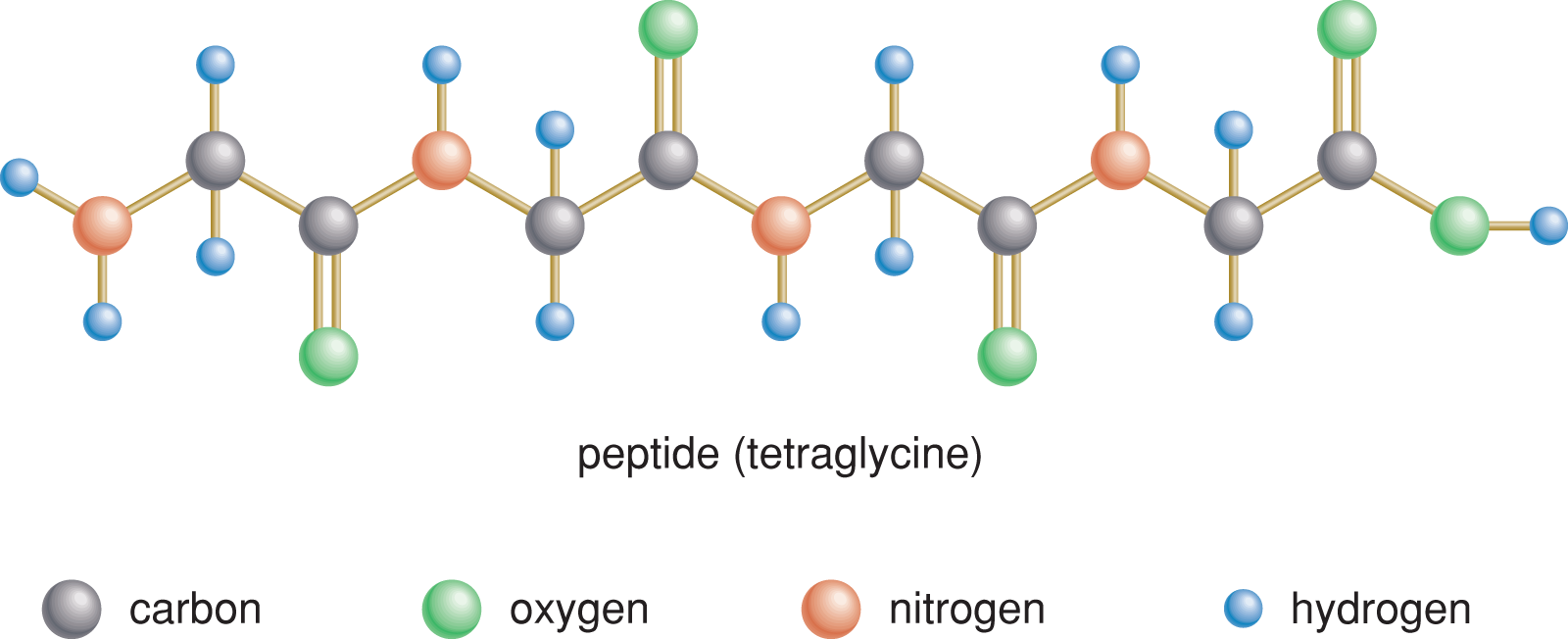
Figure 1.13 Peptide Residues
The residues in peptides are joined together through peptide bonds, a specialized form of an amide bond, which form between the –COO– group of one amino acid and the NH3+ group of another amino acid. This forms the functional group –C(O)NH–. In this section, we’ll look at the key reactions involved in forming and breaking peptide bonds.
Peptide Bond Formation
Peptide bond formation is an example of a condensation or dehydration reaction because it results in the removal of a water molecule (H2O); it can also be viewed as an acyl substitution reaction, which can occur with all carboxylic acid derivatives. When a peptide bond forms, as shown in Figure 1.14, the electrophilic carbonyl carbon on the first amino acid is attacked by the nucleophilic amino group on the second amino acid. After that attack, the hydroxyl group of the carboxylic acid is kicked off. The result is the formation of a peptide (amide) bond.
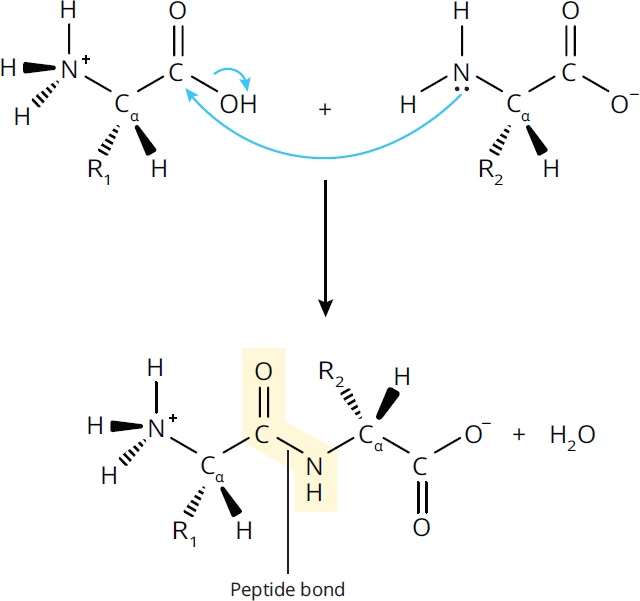
Figure 1.14 Peptide Bond Formation
Because amide groups have delocalizable π electrons in the carbonyl and in the lone pair on the amino nitrogen, they can exhibit resonance; thus, the C–N bond in the amide has partial double bond character, as shown in Figure 1.15.
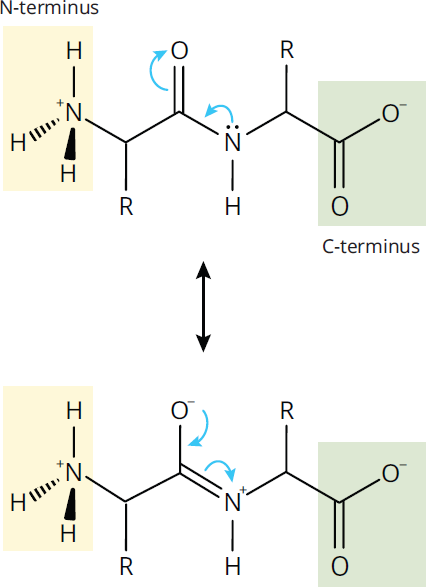
Figure 1.15 Resonance in the Peptide Bond
BRIDGE
The peptide is drawn in the same order that it is synthesized by ribosomes: from the N-terminus to the C-terminus! Translation is covered in Chapter 7 of MCAT Biochemistry Review.
As a result, rotation of the protein backbone around its C–N amide bonds is restricted, which makes the protein more rigid. Rotation around the remaining bonds in the backbone, however, is not restricted, as those remain single (σ) bonds.
When a peptide bond forms, the free amino end is known as the amino terminus or N-terminus, while the free carboxyl end is the carboxy terminus or C-terminus. By convention, peptides are drawn with the N-terminus on the left and the C-terminus on the right; similarly, they are read from N-terminus to C-terminus.
Peptide Bond Hydrolysis
For enzymes to carry out their function, peptides need to be relatively stable in solution. Therefore, they don’t normally fall apart on their own. On the other hand, in order to digest proteins, we need to break them down into their component amino acids. In organic chemistry, amides can be hydrolyzed using acid or base catalysis.
MCAT EXPERTISE
You do not need to memorize the specific amino acids that hydrolytic enzymes recognize or the exact mechanisms for those reactions. On the other hand, you could certainly encounter a passage describing them.
In living organisms, however, hydrolysis is catalyzed by hydrolytic enzymes such as trypsin and chymotrypsin. Both are specific, in that they only cleave at specific points in the peptide chain: trypsin cleaves at the carboxyl end of arginine and lysine, while chymotrypsin cleaves at the carboxyl end of phenylalanine, tryptophan, and tyrosine. While you don’t need to know the exact mechanism of how these enzymes catalyze hydrolysis, you do need to understand the main idea: they break apart the amide bond by adding a hydrogen atom to the amide nitrogen and an OH group to the carbonyl carbon as shown in Figure 1.16. This is the reverse reaction shown before in Figure 1.14.
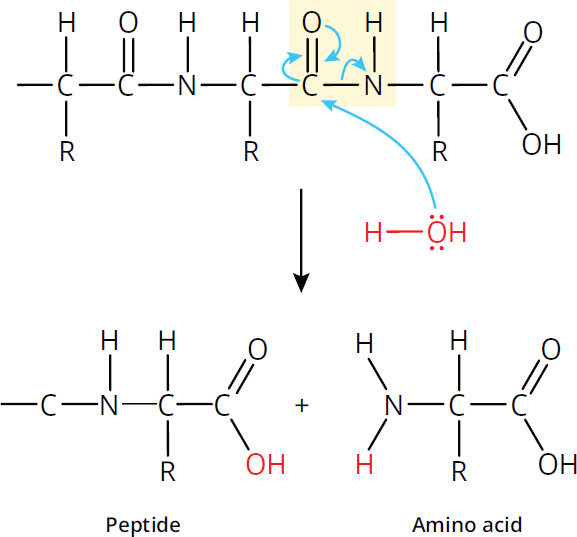
Figure 1.16 Peptide Bond Cleavage
MCAT CONCEPT CHECK 1.3:
Before you move on, assess your understanding of the material with these questions.
- What is the difference between an amino acid, a dipeptide, a tripeptide, an oligopeptide, and a polypeptide?
________________________________________
- What molecule is released during formation of a peptide bond?
________________________________________
- If chymotrypsin cleaves at the carboxyl end of phenylalanine, tryptophan, and tyrosine, how many oligopeptides would be formed in enzymatic cleavage of the following molecule with chymotrypsin?
Val – Phe – Glu – Lys – Tyr – Phe – Trp – Ile – Met – Tyr – Gly – Ala
1.4 Primary and Secondary Protein Structure
LEARNING OBJECTIVES
After Chapter 1.4, you will be able to:
- Describe all four levels of protein structure
- Recognize the unique role of proline in secondary protein structure
- Recall the structural features of α-helices and β-pleated sheets
Proteins are polypeptides that range from just a few amino acids in length up to thousands. They serve many functions in biological systems, functioning as enzymes, hormones, membrane pores and receptors, and elements of cell structure. Proteins are the main actors in cells; the genetic code, after all, is simply a recipe for making thousands of proteins.
Proteins have four levels of structure: primary (1°), secondary (2°), tertiary (3°), and quaternary (4°). In this section, we’ll examine the first two; we’ll discuss tertiary and quaternary structure in the next section.
Primary Structure
The primary structure of a protein is the linear arrangement of amino acids coded in an organism’s DNA. It’s the sequence of amino acids, listed from the N-terminus, or amino end, to the C-terminus, or carboxyl end. So, for example, the first ten amino acids of the β-chain of hemoglobin are normally valine, histidine, leucine, threonine, proline, glutamate, glutamate, lysine, serine, and alanine. Primary structure is stabilized by the formation of covalent peptide bonds between adjacent amino acids.
MCAT EXPERTISE
The MCAT will not expect you to memorize the exact primary sequence of any protein!
The primary structure alone encodes all the information needed for folding at all of the higher structural levels; the secondary, tertiary, and quaternary structures a protein adopts are the most energetically favorable arrangements of the primary structure in a given environment. The primary structure of a protein can be determined by a laboratory technique called sequencing. This is most easily done using the DNA that coded for that protein, although it can also be done from the protein itself.
Secondary Structure
A protein’s secondary structure is the local structure of neighboring amino acids. Secondary structures are primarily the result of hydrogen bonding between nearby amino acids. The two most common secondary structures are α-helices and β-pleated sheets. The key to the stability of both structures is the formation of intramolecular hydrogen bonds between different residues.
KEY CONCEPT
The primary structure of a protein is the order of its amino acids. The two main secondary structures are the α-helix and β-pleated sheet, which both result from hydrogen bonding.
*α*-Helices
The α-helix, shown in Figure 1.17, is a rodlike structure in which the peptide chain coils clockwise around a central axis. The helix is stabilized by intramolecular hydrogen bonds between a carbonyl oxygen atom and an amide hydrogen atom four residues down the chain. One complete turn of the α-helix has 3.6 amino acids residues. The side chains of the amino acids in the α-helical conformation point away from the helix core. The α-helix is an important component in the structure of keratin, a fibrous structural protein found in human skin, hair, and fingernails.
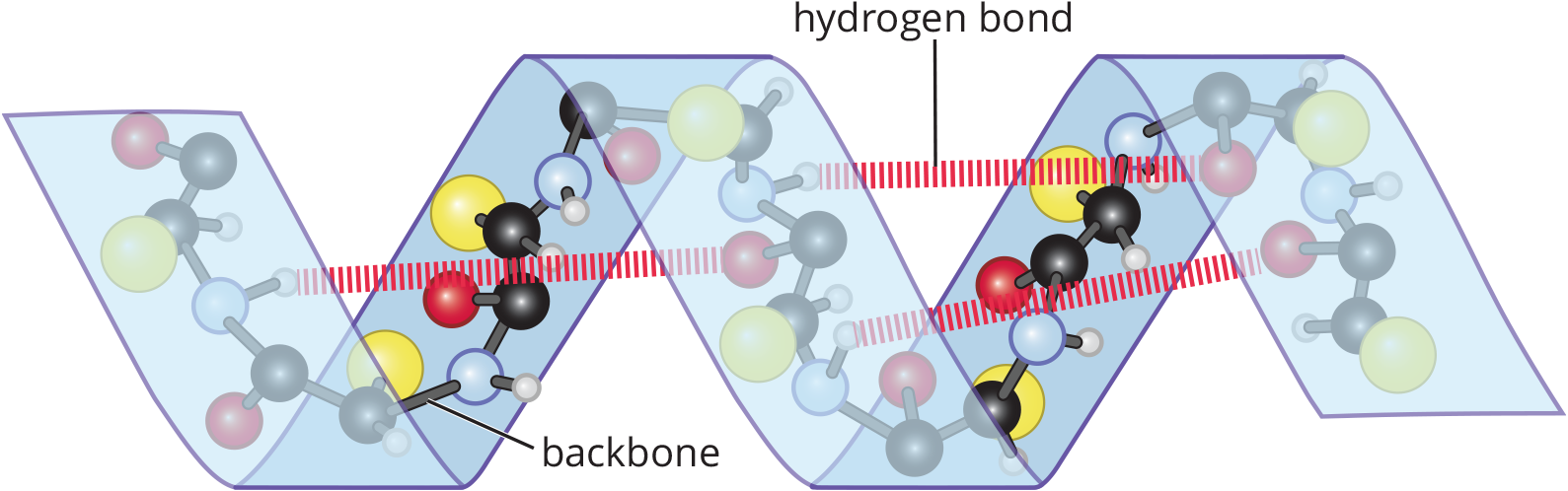
Figure 1.17 Hydrogen Bonding and Spatial Configuration of an α-Helix
*β*-Pleated Sheets
In β-pleated sheets, which can be parallel or antiparallel, the peptide chains lie alongside one another, forming rows or strands held together by intramolecular hydrogen bonds between carbonyl oxygen atoms on one chain and amide hydrogen atoms in an adjacent chain, as shown in Figure 1.18.
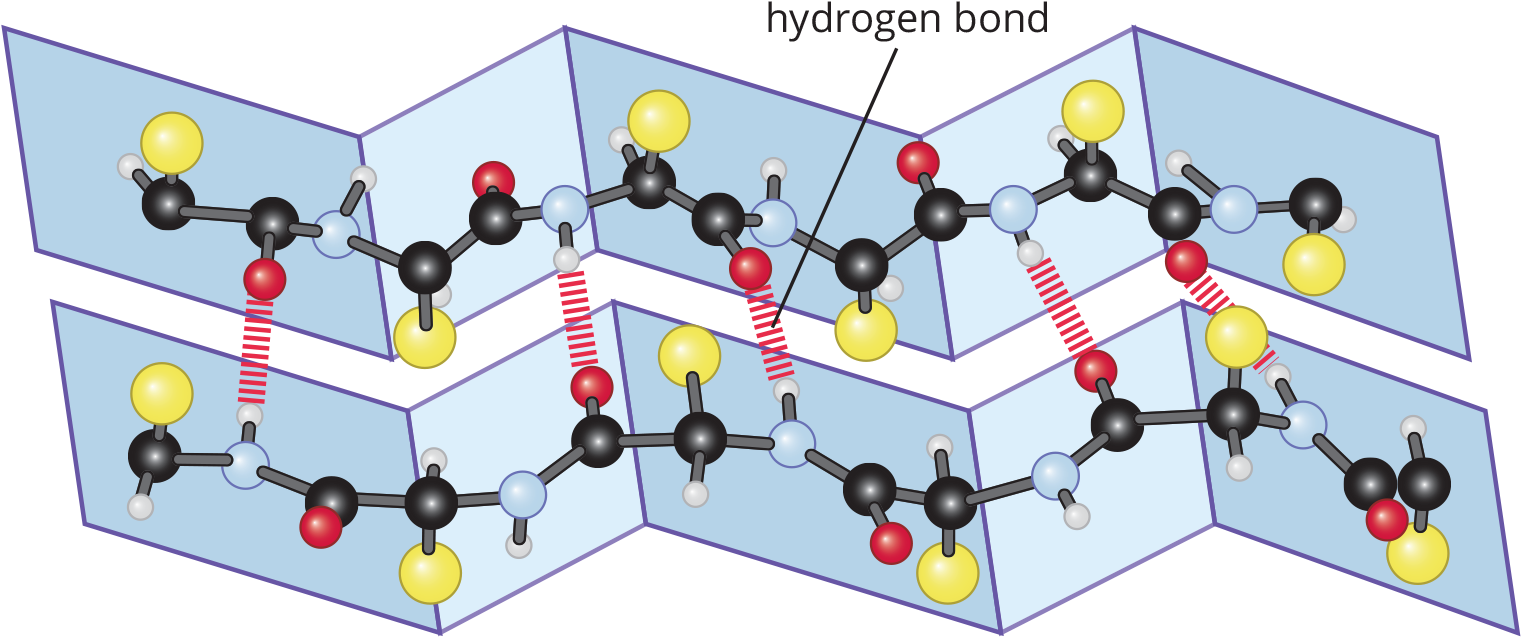
Figure 1.18 Hydrogen Bonding and Spatial Configuration of a β-Pleated Sheet
To accommodate as many hydrogen bonds as possible, the β-pleated sheets assume a pleated, or rippled, shape. The R groups of amino residues point above and below the plane of the β-pleated sheet. The adjacent peptide chains can either form a parallel or antiparallel orientation, as shown in Figure 1.19. Fibroin, the primary protein component of silk fibers, is composed of β-pleated sheets.
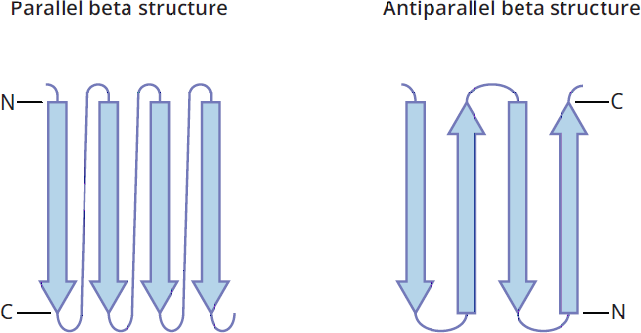
Figure 1.19 Parallel and Antiparallel β-Pleated Sheets
Turns
Turns are secondary structural motifs that lead to a change in the direction of the polypeptide chain. These motifs usually contain between one to five amino acids and join secondary structural elements together, where the first amino acid is labeled i, with subsequent ones labeled i + 1, i + 2, i + 3, and so on. The structure of the turns often allows for the formation of a main chain hydrogen bond between the residues that start and end the turn. They are classified based on the separation between the two end residues of the turn that are joined by a hydrogen bond.
- ***α*-turn:** The hydrogen bond is between end residues i and i + 4.
- ***β*-turn:** The hydrogen bond is between end residues i and i + 3. This is the most common type of turn.
- ***γ*-turn:** The hydrogen bond is between residues i and i + 2.
- ***δ*-turn:** The hydrogen bond is between residues i and i + 1. This type of turn is unlikely to form due to steric unfavorability.
- ***π*-turn:** The hydrogen bond is between residues i and i + 5.
Secondary Structures and Proline
Because of its rigid cyclic structure, proline will introduce a kink in the peptide chain when it is found in the middle of an α-helix (see Figure 1.20). Proline residues are thus rarely found in α-helices, except in helices that cross the cell membrane. Similarly, it is rarely found in the middle of pleated sheets. On the other hand, proline is often found in the turns between the chains of a β-pleated sheet, and it is often found as the residue at the start of an *α**-helix.*
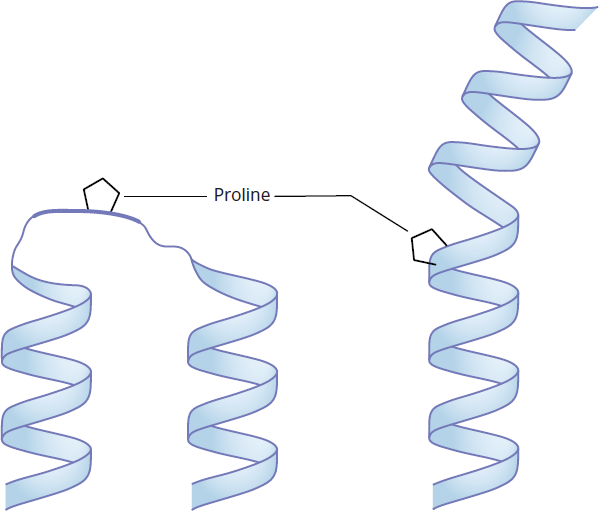
Figure 1.20 Proline in Secondary Structure
MCAT CONCEPT CHECK 1.4:
Before you move on, assess your understanding of the material with these questions.
- What are the definitions of primary and secondary structure, and how do they differ in subtypes and the bonds that stabilize them?
Structural Element Definition Subtypes Stabilizing Bonds Primary structure (1°) Secondary structure (2°)
- What role does proline serve in secondary structure?
________________________________________
- Describe the key structural features of the following secondary structures.
- α-helix:
________________________________________
- β-pleated sheet:
________________________________________
1.5 Tertiary and Quaternary Protein Structure
LEARNING OBJECTIVES
After Chapter 1.5, you will be able to:
- Identify the major structural components of tertiary and quaternary protein structure
- Recognize the relationship between protein folding and the solvation layer
- Recall the nomenclature of proteins with prosthetic groups, such as nucleoproteins
Proteins can be broadly divided into fibrous proteins, such as collagen, that have structures that resemble sheets or long strands, and globular proteins, such as myoglobin, that tend to be spherical (that is, like a globe). These are caused by tertiary and quaternary protein structures, both of which are the result of protein folding.
Tertiary Structure
A protein’s tertiary structure is its three-dimensional shape. Tertiary structures are mostly determined by hydrophilic and hydrophobic interactions between R groups of amino acids. Hydrophobic residues prefer to be on the interior of proteins, which reduces their proximity to water. Hydrophilic N–H and C=O bonds found in the polypeptide chain get pulled in by these hydrophobic residues. These hydrophilic bonds can then form electrostatic interactions and hydrogen bonds that further stabilize the protein from the inside. As a result of these hydrophobic interactions, most of the amino acids on the surface of proteins have hydrophilic (polar or charged) R groups; highly hydrophobic R groups, such as phenylalanine, are almost never found on the surface of a protein.
KEY CONCEPT
The tertiary structure of a protein is primarily the result of moving hydrophobic amino acid side chains into the interior of the protein.
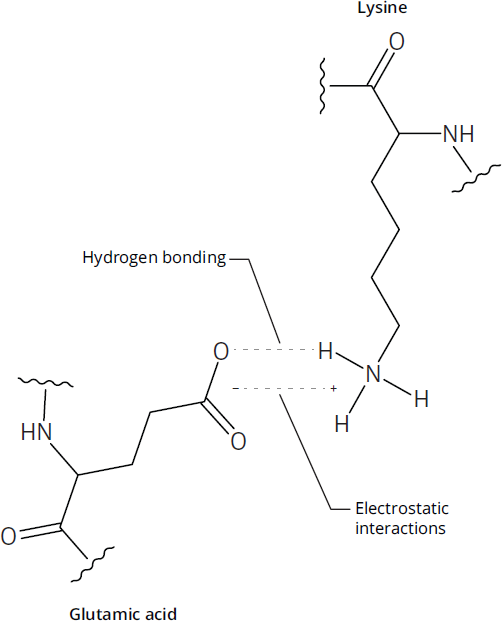
Figure 1.21 Salt Bridge Interaction
The three-dimensional structure can also be determined by hydrogen bonding, as well as acid-base interactions between amino acids with charged R groups. When an interaction consists of both a hydrogen bond and an ionic bond, as shown in Figure 1.21, it is referred to as a salt bridge. A particularly important component of tertiary structure is the presence of disulfide bonds, the bonds that form when two cysteine molecules become oxidized to form cystine, as shown in Figure 1.22. Disulfide bonds create loops in the protein chain. In addition, disulfide bonds determine how wavy or curly human hair is: the more disulfide bonds, the curlier it is. Note that forming a disulfide bond requires the loss of two protons and two electrons (oxidation).
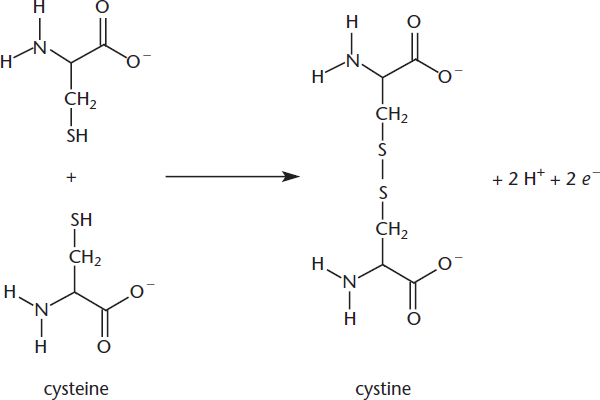
Figure 1.22 Disulfide Bond Formation
The exact details of protein folding are beyond the scope of the MCAT, but the basic idea is that the secondary structures probably form first, and then hydrophobic interactions and hydrogen bonds cause the protein to “collapse” into its proper three-dimensional structure. Along the way, it adopts intermediate states known as molten globules. Protein folding is an extremely rapid process: from start to finish, it typically takes much less than a second.
If a protein loses its tertiary structure, a process commonly called denaturation, it loses its function.
Folding and the Solvation Layer
Why do hydrophobic residues tend to occupy the interior of a protein, while hydrophilic residues tend to accumulate on the exterior portions? The answer can be summed up in one word: entropy.
Whenever a solute dissolves in a solvent, the nearby solvent molecules form a solvation layer around that solute. From an enthalpy standpoint, even hydrocarbons are more stable in aqueous solution than in organic ones (ΔH < 0). However, when a hydrophobic side chain, such as those in phenylalanine and leucine, is placed in aqueous solution, the water molecules in the solvation layer cannot form hydrogen bonds with the side chain. As depicted in Figure 1.23, this forces the nearby water molecules to rearrange themselves into specific arrangements to maximize hydrogen bonding—which means a negative change in entropy, ΔS. Remember that negative changes in entropy represent increasing order (decreasing disorder) and thus are unfavorable. This entropy change makes the overall process nonspontaneous (ΔG > 0).
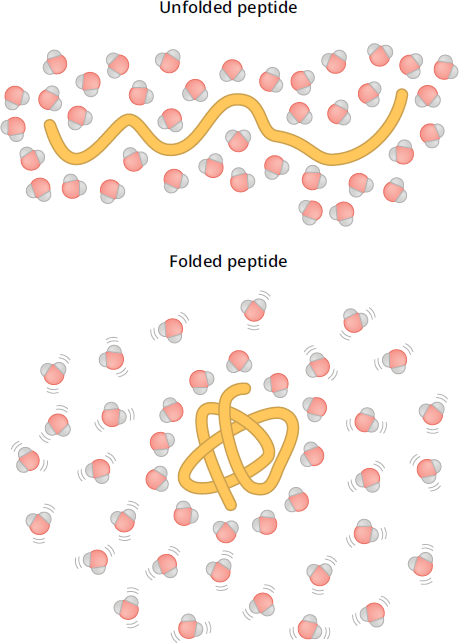
Figure 1.23 Protein Folding and Entropy
BRIDGE
Make sure you understand the basic thermodynamic properties of enthalpy, entropy, and Gibbs free energy, discussed in Chapter 7 of MCAT General Chemistry Review. On Test Day, they can be tested on both natural sciences sections!
On the other hand, putting hydrophilic residues such as serine or lysine on the exterior of the protein allows the nearby water molecules more latitude in their positioning, thus increasing their entropy (ΔS > 0), and making the overall solvation process spontaneous. Thus, by moving hydrophobic residues away from water molecules and hydrophilic residues toward water molecules, a protein achieves maximum stability.
Quaternary Structure
All proteins have elements of primary, secondary, and tertiary structure; not all proteins have quaternary structure. Quaternary structures only exist for proteins that contain more than one polypeptide chain. For these proteins, the quaternary structure is an aggregate of smaller globular peptides, or subunits, and represents the functional form of the protein. The classic examples of quaternary structure are hemoglobin and immunoglobulins, shown in Figures 1.24a and 1.24b. Hemoglobin consists of four distinct subunits, each of which can bind one molecule of oxygen. Similarly, immunoglobulin G (IgG) antibodies also contain a total of four subunits each, held together by disulfide bonds.
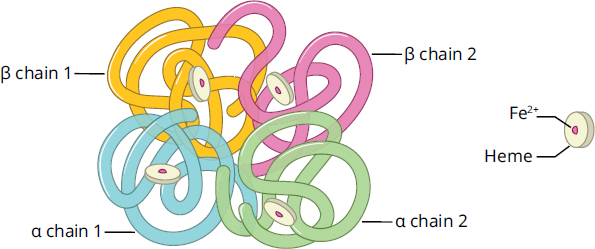
Figure 1.24a Hemoglobin Heme molecules are visible in each chain.
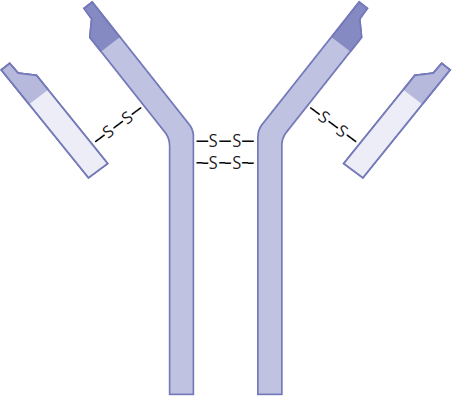
Figure 1.24b Immunoglobulin G1
MNEMONIC
The primary structure of a protein acts like letters. The secondary structure acts like words: only certain orderings of letters make sense (CAT is a word, while CAQ is not). The tertiary structure acts like sentences: words combine to form a functioning whole. The quaternary structure acts like paragraphs: they’re not always present, but subunits can combine to make a cohesive whole.
The formation of quaternary structures can serve several roles. First, they can be more stable, by further reducing the surface area of the protein complex. Second, they can reduce the amount of DNA needed to encode the protein complex. Third, they can bring catalytic sites close together, allowing intermediates from one reaction to be directly shuttled to a second reaction. Finally, and most important, they can induce cooperativity, or allosteric effects. We’ll discuss this much further in the next chapter (especially for hemoglobin), but the basic idea is that one subunit can undergo conformational or structural changes, which either enhance or reduce the activity of the other subunits.
BRIDGE
The reduction of genetic material is crucial for viruses. The genome for most viruses is tiny. Thus, their viral coats typically consist of one small protein repeated dozens or even hundreds of times. Viral structure is discussed in Chapter 1 of MCAT Biology Review.
Conjugated Proteins
Conjugated proteins derive part of their function from covalently attached molecules called prosthetic groups. These prosthetic groups can be organic molecules, such as vitamins, or even metal ions, such as iron. Proteins with lipid, carbohydrate, and nucleic acid prosthetic groups are referred to as lipoproteins, glycoproteins, and nucleoproteins, respectively. These prosthetic groups have major roles in determining the function of their respective proteins. For example, each of hemoglobin’s subunits (as well as myoglobin) contains a prosthetic group called heme. The heme group is composed of four pyrrole rings. Pyrroles are five-membered heterocyclic aromatic compounds, in which the nitrogen of each pyrrole acts to coordinate the central iron ion. The iron atom binds to and carries oxygen; as such, hemoglobin is inactive without the heme group. These groups can also direct the protein to be delivered to a certain location, such as the cell membrane, nucleus, lysosome, or endoplasmic reticulum.
BIOCHEMISTRY GUIDED EXAMPLE WITH EXPERT THINKING
Is R48C/T77C/V76I a suitable candidate for further testing? Why or why not?
This question is asking us to interpret the study results, but we'll want to start with the purpose of the study to be able to go through the data efficiently. The goal is to develop a vaccine for ricin poisoning. The second paragraph starts with what has already been done, which is the creation of a possible working vaccine. Since it’s based on the ricin protein, we can use our background knowledge and critical reasoning to assume that the vaccine is similar enough to the ricin protein to elicit a proper immune response, but does not inhibit ribosome function. Notice the keyword “however” in the second paragraph—this indicates that there is still a problem. In this case, the current vaccine is not stable, which would presumably affect its efficacy. Also, the vaccine seems to have a serious side effect (VLS), which is something the researchers want to eliminate. We’re given a hint about a possible solution to the stability issue—creating a disulfide bond. This means that the disulfide is not native to the protein, but created by mutating select amino acids to cysteines.
Following that possible solution in paragraph 2, the researchers generated a series of protein mutations, which are outlined in the first column of Table 1. Notice the notation: letter-number-letter. We know from the passage that these are mutants and, in looking at this code, we can infer that the first letter is the original amino acid, the number is its position in the sequence, and the second letter is the mutated amino acid. This style of notation is common in scientific papers, so we'll likely want to keep this in mind even on Test Day as a possibility when we see similar notation. For instance, R48C is arginine at position 48 mutated to cysteine. Notice that in each mutation, there is a pair of amino acids mutated to cysteine to create the opportunity for a disulfide bond. The second column tells us about the success of creating a disulfide bond. Since the wild-type protein has one free cysteine, and the mutants introduce two more cysteines for the purpose of a creating a disulfide bond, a successful disulfide bond formation will eliminate two sulfhydryls, leaving the original free cysteine. R48C/T77C/V76I has approximately two (1.89 ± 0.03) free cysteines, suggesting an incomplete formation of the disulfide bond. The third column determines the effect of the disulfide bond formation on stability. Melting temperature (Tm) is a great indication for stability—the higher the temperature, the more energy it takes to break all bonds to unfold the protein. Both wild-type and R48C/T77C/V76I have markedly lower Tm compared to the other constructs, which we can conclude means they have reduced stability. Finally, the last column provides information about whether the construct induced VLS. We can see that R48C/T77C/V76I did not induce VLS, making it 'safer' from the perspective of risk for this dangerous syndrome.
Given the data from the passage, while R48C/T77C/V76I does address the issue of not inducing VLS in animal models, the construct is not stable, and therefore would not be suitable for additional testing.
MCAT CONCEPT CHECK 1.5:
Before you move on, assess your understanding of the material with these questions.
- What are the definitions of tertiary and quaternary structure, and how do they differ in subtypes and the bonds that stabilize them?
Structural Element Definition Subtypes Stabilizing Bonds Tertiary structure (3°) Quaternary structure (4°)
- What is the primary motivation for hydrophobic residues in a polypeptide to move to the interior of the protein?
________________________________________
- List three different prosthetic groups that can be attached to a protein and name the conjugated protein.
________________________________________
1.6 Denaturation
LEARNING OBJECTIVES
After Chapter 1.6, you will be able to:
- Predict the impact of denaturation via heat or solute
In the previous section, we discussed protein folding. The reverse of this process is denaturation, in which a protein loses its three-dimensional structure. Although it is sometimes reversible, denaturation is often irreversible; whether its denaturation is reversible or not, unfolded proteins cannot catalyze reactions. The two main causes of denaturation are heat and solutes.
BRIDGE
Heat denatures proteins for the same reason that raising a reaction’s temperature increases its rate: increased average kinetic energy of molecules means increased molecular motion. The relationship between temperature and kinetic energy is discussed in Chapter 7 of MCAT General Chemistry Review.
As with all molecules, when the temperature of a protein increases, its average kinetic energy increases. When the temperature gets high enough, this extra energy can be enough to overcome the hydrophobic interactions that hold a protein together, causing the protein to unfold. This is what happens when egg whites are cooked: in the uncooked egg whites, albumin is folded, which makes it transparent; cooking them causes the albumin molecules to denature and aggregate, forming a solid, rubbery white mass that will not revert to its transparent form.
KEY CONCEPT
Denatured proteins lose their three-dimensional structure and are thus inactive.
On the other hand, solutes such as urea denature proteins by directly interfering with the forces that hold the protein together. They can disrupt tertiary and quaternary structures by interrupting non-covalent interactions like hydrogen bonds and other side chain interactions that hold α-helices and β-pleated sheets intact. Similarly, detergents such as SDS (sodium dodecyl sulfate, also called sodium lauryl sulfate) can solubilize proteins, disrupting noncovalent bonds and promoting denaturation. Note that both urea and SDS do not interfere with any covalent bonds, including disulfide and peptide bonds.
The stability of proteins to thermal or chemical denaturation can be assessed using denaturation curves, which plot the fraction of unfolded protein against increasing amounts of a denaturant. The denaturation midpoint describes the point at which there are equal amounts of the folded and unfolded proteins. When assessing the effect of temperature on protein unfolding, the midpoint is called the melting point (***Tm), and when using a chemical denaturant to assess protein stability, the midpoint is referred to as amidpoint concentration(C*m**). Changes in the midpoint indicate changes in protein folding stability: For example, Figure 1.25 shows the melting curves of two variants of a protein, Variant A and Variant B. Comparing the two thermal denaturation curves shows that Variant B has a higher Tm, suggesting Variant B requires higher temperatures to denature, while Variant A is less thermally stable and will unfold at lower temperatures.
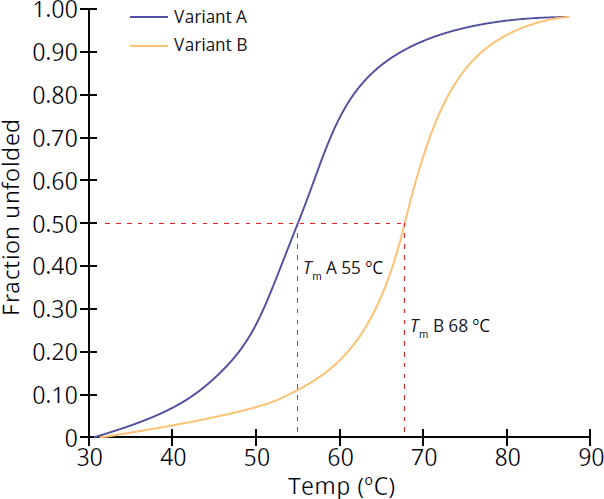
Figure 1.25 Thermal Denaturation Curve Changes in Tm correlate to protein folding stability.
MCAT CONCEPT CHECK 1.6:
Before you move on, assess your understanding of the material with this question.
- Why are proteins denatured by heat and solutes, respectively?
- Heat:
________________________________________
- Solutes:
________________________________________
Conclusion
Nearly every part of a cell involves proteins in some way, from the nucleus to the mitochondria to the cell membrane. The MCAT will test your understanding of key concepts regarding amino acids because the amino acids that compose a protein determine its structure. In the next chapter, we’ll discuss the best-known function of proteins: their role as enzymes.
GO ONLINE!
You've reviewed the content, now test your knowledge and critical thinking skills by completing a test-like passage set in your online resources!
CONCEPT SUMMARY
Amino Acids Found in Proteins
- Amino acids have four groups attached to a central (α) carbon: an amino group, a carboxylic acid group, a hydrogen atom, and an R group.
- The R group determines chemistry and function of that amino acid.
- Twenty amino acids appear in the proteins of eukaryotic organisms.
- The stereochemistry of the α-carbon is L for all chiral amino acids in eukaryotes.
- D-amino acids can exist in prokaryotes.
- All chiral amino acids except cysteine have an (S) configuration.
- All amino acids are chiral except glycine, which has a hydrogen atom as its R group.
- Side chains can be polar or nonpolar, aromatic or nonaromatic, charged or uncharged.
- Nonpolar, nonaromatic: glycine, alanine, valine, leucine, isoleucine, methionine, proline
- Aromatic: tryptophan, phenylalanine, tyrosine
- Polar: serine, threonine, asparagine, glutamine, cysteine
- Negatively charged (acidic): aspartate, glutamate
- Positively charged (basic): lysine, arginine, histidine
- Amino acids with long alkyl chains are hydrophobic, and those with charges are hydrophilic; many others fall somewhere in between.
Acid–Base Chemistry of Amino Acids
- Amino acids are amphoteric; that is, they can accept or donate protons.
- The pKa of a group is the pH at which half of the species are deprotonated; [HA] = [A–].
- Amino acids exist in different forms at different pH values.
- At low (acidic) pH, the amino acid is fully protonated.
- At pH near the pI of the amino acid, the amino acid is a neutral zwitterion.
- At high (alkaline) pH, the amino acid is fully deprotonated.
- The isoelectric point (pI) of an amino acid without a charged side chain can be calculated by averaging the two pKa values.
- Amino acids can be titrated.
- The titration curve is nearly flat at the pKa values of the amino acid.
- The titration curve is nearly vertical at the pI of the amino acid.
- Amino acids with charged side chains have an additional pKa value, and their pI is calculated by averaging the two pKa values that correspond to protonation and deprotonation of the zwitterion.
- Amino acids without charged side chains have a pI around 6.
- Acidic amino acids have a pI well below 6.
- Basic amino acids have a pI well above 6.
Peptide Bond Formation and Hydrolysis
- Dipeptides have two amino acid residues; tripeptides have three. Oligopeptides have a “few” amino acid residues (<20); polypeptides have “many” (>20).
- Forming a peptide bond is a condensation or dehydration reaction (releasing one molecule of water).
- The nucleophilic amino group of one amino acid attacks the electrophilic carbonyl group of another amino acid.
- Amide bonds are rigid because of resonance.
- Breaking a peptide bond is a hydrolysis reaction.
Primary and Secondary Protein Structure
- Primary structure is the linear sequence of amino acids in a peptide and is stabilized by peptide bonds.
- Secondary structure is the local structure of neighboring amino acids, and is stabilized by hydrogen bonding between amino groups and nonadjacent carboxyl groups.
- α-helices are clockwise coils around a central axis.
- β-pleated sheets are rippled strands that can be parallel or antiparallel.
- Proline can interrupt secondary structure because of its rigid cyclic structure.
Tertiary and Quaternary Protein Structure
- Tertiary structure is the three-dimensional shape of a single polypeptide chain, and is stabilized by hydrophobic interactions, acid–base interactions (salt bridges), hydrogen bonding, and disulfide bonds.
- Hydrophobic interactions push hydrophobic R groups to the interior of a protein, which increases entropy of the surrounding water molecules and creates a negative Gibbs free energy.
- Disulfide bonds occur when two cysteine molecules are oxidized and create a covalent bond to form cystine.
- Quaternary structure is the interaction between peptides in proteins that contain multiple subunits.
- Proteins with covalently attached molecules are termed conjugated proteins. The attached molecule is a prosthetic group, and may be a metal ion, vitamin, lipid, carbohydrate, or nucleic acid.
Denaturation
- Both heat and increasing solute concentration can lead to loss of three-dimensional protein structure, which is termed denaturation.
ANSWERS TO CONCEPT CHECKS
1.1
- The four groups are an amino group (–NH2), a carboxylic acid group (–COOH), a hydrogen atom, and an R group.
- All chiral eukaryotic amino acids are L. All chiral eukaryotic amino acids are (S), with the exception of cysteine (because cysteine is the only amino acid with an R group that has a higher priority than a carboxylic acid according to Cahn–Ingold–Prelog rules).
- Nonpolar, nonaromatic: glycine, alanine, valine, leucine, isoleucine, methionine, proline
- Aromatic: tryptophan, phenylalanine, tyrosine
- Polar: serine, threonine, asparagine, glutamine, cysteine
- Negatively charged/acidic: aspartate, glutamate
- Positively charged/basic: lysine, arginine, histidine
- Hydrophobic amino acids tend to reside in the interior of a protein, away from water. Hydrophilic amino acids tend to remain on the surface of the protein, in contact with water.
- From left to right the amino acids are methionine (M), cysteine (C), alanine (A), and threonine (T): MCAT.
**1.2**
- pH = 1:+NH3CRHCOOH; pH = 7:+NH3CRHCOO–; pH = 11: NH2CRHCOO–
Arginine: pI = (9.04 + 12.48) ÷ 2 = 10.76 Valine: pI = (2.32 + 9.62) ÷ 2 = 5.97
- Aspartic acid: pI = (1.88 + 3.65) ÷ 2 = 2.77
**1.3**
- These species differ by the number of amino acids that make them up: amino acid = 1, dipeptide = 2, tripeptide = 3, oligopeptide = “few” (< 20), polypeptide = “many” (> 20)
- Water (H2O)
- 4: Val – Phe; Glu – Lys – Tyr; Ile – Met – Tyr; Gly–Ala. A single amino acid on its own is not considered an oligopeptide.
**1.4**
-
Structural Element Definition Subtypes Stabilizing Bonds Primary structure (1°) Linear sequence of amino acids in chain (none) Peptide (amide) bond
Secondary structure (2°) Local structure determined by nearby amino acids
- α-helix
- β-pleated sheet
Hydrogen bonds
- Proline’s rigid structure causes it to introduce kinks in α-helices or create turns in β-pleated sheets.
- The α-helix is a rod-like structure in which the peptide chain coils clockwise around a central axis. In β-pleated sheets the peptide chains lie alongside one another, forming rows or strands held together by intramolecular hydrogen bonds between carbonyl oxygen atoms on one chain and amide hydrogen atoms in an adjacent chain.
**1.5**
-
Structural Element Definition Subtypes Stabilizing Bonds Tertiary structure (3°) Three-dimensional shape of protein
- Hydrophobic interactions
- Acid–base/salt bridges
- Disulfide links
- van der Waals forces
- Hydrogen bonds
- Ionic bonds
- Covalent bonds
Quaternary structure (4°) Interaction between separate subunits of a multisubunit protein (none) Same as tertiary structure
- Moving hydrophobic residues to the interior of a protein increases entropy by allowing water molecules on the surface of the protein to have more possible positions and configurations. This positive ΔSmakesΔG< 0, stabilizing the protein.
- Examples of common prosthetic groups include lipids, carbohydrates, and nucleic acids; they are known as lipoproteins, glycoproteins, and nucleoproteins, respectively.
**1.6**
- Heat denatures proteins by increasing their average kinetic energy, thus disrupting hydrophobic interactions. Solutes denature proteins by disrupting elements of secondary, tertiary, and quaternary structure.
SCIENCE MASTERY ASSESSMENT EXPLANATIONS
1. B
Most amino acids (except the acidic and basic amino acids) have two sites for protonation: the carboxylic acid and the amine. At neutral pH, the carboxylic acid will be deprotonated (–COO–) and the amine will remain protonated –NH3+. This dipolar ion is a zwitterion, so (B) is the correct answer.
2. B
Glutamic acid is an acidic amino acid because it has an extra carboxyl group. At neutral pH, both carboxyl groups are deprotonated and thus negatively charged. The amino group has a positive charge because it remains protonated at pH 7. Overall, therefore, glutamic acid has a net charge of –1, and (B) is correct. Notice that you do not even need to know the pI values to solve this question; as an acidic amino acid, glutamic acid must have a pI below 7.
3. C
Nonpolar groups are not capable of forming dipoles or hydrogen bonds; this makes them hydrophobic. Burying hydrophobic R groups inside proteins means they don’t have to interact with water, which is polar. This makes (C) correct. (A) and (B) are incorrect because nonpolar molecules are hydrophobic, not hydrophilic; (D) is incorrect because they are not generally found on protein surfaces.
4. A
The cDNA sequence is a DNA copy of the mRNA used to generate a protein. A computer program can quickly identify the amino acid that corresponds to each codon and generate a list of these amino acids. This amino acid sequence is the primary structure of the protein. These observations support (A) as the correct answer. By contrast, the secondary, tertiary, and quaternary structures involve higher level interactions between the backbone and R groups and are increasingly difficult to predict.
5. C
There are three choices for the first amino acid, leaving two choices for the second, and one choice for the third. Multiplying those numbers gives us a total of3 × 2 × 1 = 6distinct tripeptides. (Using the one-letter codes for valine (V), alanine (A), and leucine (L), those six tripeptides are VAL, VLA, ALV, AVL, LVA, and LAV.)
6. B
As the protein folds, it takes on an organized structure and thus its entropy decreases. However, the opposite trend is true for the water surrounding the protein. Prior to protein folding, hydrophobic amino acid residues are exposed and the water molecules must form structured hydration shells around these hydrophobic residues. During folding, these hydrophobic residues are generally buried in the interior of the protein so that the surrounding water molecules gain more latitude in their interactions. Thus, the entropy of the surrounding water increases, making the correct answer (B).
7. C
The α-helix is held together primarily by hydrogen bonds between the carboxyl groups and amino groups of amino acids. Disulfide bridges, (A), and hydrophobic effects, (B), are primarily involved in tertiary structures, not secondary. Even if they were charged, the side chains of amino acids are too far apart to participate in strong interactions in secondary structure.
8. C
High salt concentrations and detergents can denature a protein, as can high temperatures. But moving a protein to a hypotonic environment—that is, a lower solute concentration—should not lead to denaturation.
9. C
An amino acid likely to be found in a transmembrane portion of an α-helix will be exposed to a hydrophobic environment, so we need an amino acid with a hydrophobic side chain. The only choice that has a hydrophobic side chain is (C), phenylalanine. The other choices are all polar or charged.
10. C
Every amino acid except glycine has a chiral α-carbon, but only two of the 20 amino acids—threonine and isoleucine— also have a chiral carbon in their side chains as well. Thus, the correct answer is (C). Just as only one configuration is normally seen at the αcarbon, only one configuration is seen in the side chain chiral carbon.
11. B
In their inactive state, the receptor tyrosine kinases are fully folded single polypeptide chains and thus have tertiary structure. When these monomers dimerize, they become a protein complex and thus have elements of quaternary structure. This change from tertiary to quaternary structure justifies (B).
12. A
Histidine has an ionizable side chain: its imidazole ring has a nitrogen atom that can be protonated. None of the remaining answers have ionizable atoms in their side chains.
13. D
Because lysine has a basic side chain, we ignore the pKa of the carboxyl group, and average the pKa of the side chain and the amino group; the average of 9 and 10.5 is 9.75, which is closest to (D).
14. D
Conjugated proteins can have lipid or carbohydrate “tags” added to them. These tags can indicate that these proteins should be directed to the cell membrane (especially lipid tags) or to specific organelles (such as the lysosome). They can also provide the activity of the protein; for example, the heme group in hemoglobin is needed for it to bind oxygen. Thus, (D) is the correct answer.
15. B
Because collagen has a triple helix, the carbon backbones are very close together. Thus, steric hindrance is a potential problem. To reduce that hindrance, we need small side chains; glycine has the smallest side chain of all: a hydrogen atom.
GO ONLINE
Consult your online resources for additional practice.
EQUATIONS TO REMEMBER
(1.1) Isoelectric point of a neutral amino acid:
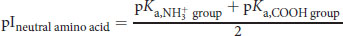
(1.2) Isoelectric point of an acidic amino acid:
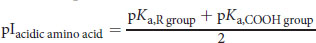
(1.3) Isoelectric point of a basic amino acid:
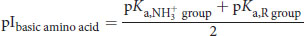
SHARED CONCEPTS
- Biochemistry Chapter 2
- Enzymes
- Biochemistry Chapter 3
- Nonenzymatic Protein Function and Protein Analysis
- Biochemistry Chapter 7
- RNA and the Genetic Code
- Biology Chapter 1
- The Cell
- Biology Chapter 9
- The Digestive System
- General Chemistry Chapter 7
- Thermochemistry
- Organic Chemistry Chapter 8
- Carboxylic Acids
- Organic Chemistry Chapter 9
- Carboxylic Acid Derivatives