Chapter 2: Enzymes
Chapter 2: Enzymes
SCIENCE MASTERY ASSESSMENT
Every pre-med knows this feeling: there is so much content I have to know for the MCAT! How do I know what to do first or what's important?
While the high-yield badges throughout this book will help you identify the most important topics, this Science Mastery Assessment is another tool in your MCAT prep arsenal. This quiz (which can also be taken in your online resources) and the guidance below will help ensure that you are spending the appropriate amount of time on this chapter based on your personal strengths and weaknesses. Don't worry though— skipping something now does not mean you'll never study it. Later on in your prep, as you complete full-length tests, you'll uncover specific pieces of content that you need to review and can come back to these chapters as appropriate.
How to Use This Assessment
If you answer 0–7 questions correctly:
Spend about 1 hour to read this chapter in full and take limited notes throughout. Follow up by reviewing all quiz questions to ensure that you now understand how to solve each one.
If you answer 8–11 questions correctly:
Spend 20–40 minutes reviewing the quiz questions. Beginning with the questions you missed, read and take notes on the corresponding subchapters. For questions you answered correctly, ensure your thinking matches that of the explanation and you understand why each choice was correct or incorrect.
If you answer 12–15 questions correctly:
Spend less than 20 minutes reviewing all questions from the quiz. If you missed any, then include a quick read-through of the corresponding subchapters, or even just the relevant content within a subchapter, as part of your question review. For questions you got correct, ensure your thinking matches that of the explanation and review the Concept Summary at the end of the chapter.
- Consider a biochemical reaction A → B, which is catalyzed by A–B dehydrogenase. Which of the following statements is true?
- The reaction will proceed until the enzyme concentration decreases.
- The reaction will be most favorable at 0 °C.
- A component of the enzyme is transferred from A to B.
- The free energy change (ΔG) of the catalyzed reaction is the same as for the uncatalyzed reaction.
- Which of the following statements about enzyme kinetics is FALSE?
- An increase in the substrate concentration (at constant enzyme concentration) leads to proportional increases in the rate of the reaction.
- Most enzymes operating in the human body work best at a temperature of 37 °C.
- An enzyme–substrate complex can either form a product or dissociate back into the enzyme and substrate.
- Maximal activity of many human enzymes occurs around pH 7.4.
- Some enzymes require the presence of a nonprotein molecule to behave catalytically. An enzyme devoid of this molecule is called a(n):
- holoenzyme.
- apoenzyme.
- coenzyme.
- zymoenzyme.
- Which of the following factors determine an enzyme’s specificity?
- The three-dimensional shape of the active site
- The Michaelis constant
- The type of cofactor required for the enzyme to be active
- The prosthetic group on the enzyme
- Human DNA polymerase is removed from the freezer and placed in a 60 °C water bath. Which of the following best describes the change in enzyme activity as the polymerase sample comes to thermal equilibrium with the water bath?
- Increases then decreases
- Decreases then plateaus
- Increases then plateaus
- Decreases then increases
- In the equation below, substrate C is an allosteric inhibitor to enzyme 1. Which of the following is another mechanism necessarily caused by substrate C?
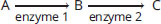
- Competitive inhibition
- Irreversible inhibition
- Feedback enhancement
- Negative feedback
- The activity of an enzyme is measured at several different substrate concentrations, and the data are shown in the table below.
**[S] (mM)**

0.01 1
0.05 9.1
0.1 17
0.5 50
1 67
5 91
10 95
50 99
100 100
Km for this enzyme is approximately:
- 0.5
- 1.0
- 10.0
- 50.0
Questions 8 and 9 refer to the following statement:
Consider a reaction catalyzed by enzyme A with a Km value of 5 × 10–6 M and Vmax of 20
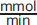
.
- At a concentration of 5 × 10–6 M substrate, the rate of the reaction will be:
- 10
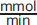
.
- 20
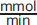
.
- 30
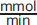
.
- 40
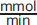
.
- At a concentration of 5 × 10–4 M substrate, the rate of the reaction will be:
- 10
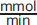
.
- 15
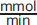
.
- 20
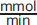
.
- 30
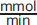
.
- The graph below shows kinetic data obtained for flu virus enzyme activity as a function of substrate concentration in the presence and absence of two antiviral drugs.
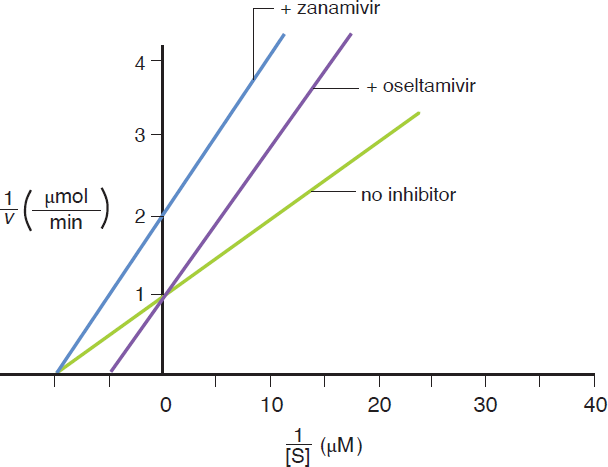
Based on the graph, which of the following statements is correct?
- Both drugs are noncompetitive inhibitors of the viral enzyme.
- Oseltamivir increases the Km value for the substrate compared to Relenza.
- Zanamivir increases the Vmax value for the substrate compared to Tamiflu.
- Both drugs are competitive inhibitors of the viral enzyme.
- The conversion of ATP to cyclic AMP and inorganic phosphate is most likely catalyzed by which class of enzyme?
- Ligase
- Hydrolase
- Lyase
- Transferase
- Which of the following is NOT a method by which enzymes decrease the activation energy for biological reactions?
- Modifying the local charge environment
- Forming transient covalent bonds
- Acting as electron donors or receptors
- Breaking bonds in the enzyme irreversibly to provide energy
- A certain enzyme that displays positive cooperativity has four subunits, two of which are bound to substrate. Which of the following statements must be true?
- The affinity of the enzyme for the substrate has just increased.
- The affinity of the enzyme for the substrate has just decreased.
- The affinity of the enzyme for the substrate is half of what it would be if four sites had substrate bound.
- The affinity of the enzyme for the substrate is greater than with one substrate bound.
- Which of the following is LEAST likely to be required for a series of metabolic reactions?
- Triacylglycerol acting as a coenzyme
- Oxidoreductase enzymes
- Magnesium acting as a cofactor
- Transferase enzymes
- How does the ideal temperature for a reaction change with and without an enzyme catalyst?
- The ideal temperature is generally higher with a catalyst than without.
- The ideal temperature is generally lower with a catalyst than without.
- The ideal temperature is characteristic of the reaction, not the enzyme.
- No conclusion can be made without knowing the enzyme type.
Answer Key
- D
- A
- B
- A
- A
- D
- A
- A
- C
- B
- C
- D
- D
- A
- B
Chapter 2: Enzymes
CHAPTER 2
ENZYMES
In This Chapter
**2.1 Enzymes as Biological Catalysts
**
Enzyme Classifications
Impact on Activation Energy
**2.2 Mechanisms of Enzyme Activity
**
Enzyme–Substrate Binding
Cofactors and Coenzymes
**2.3 Enzyme Kinetics
**
Kinetics of Monomeric Enzymes
Michaelis–Menten Equation
Lineweaver–Burk Plots
Cooperativity
**2.4 Effects of Local Conditions on Enzyme Activity
**
Temperature
pH
Salinity
**2.5 Regulation of Enzyme Activity
**
Feedback Regulation
Reversible Inhibition
Irreversible Inhibition
Regulated Enzymes
Concept Summary
CHAPTER PROFILE
The content in this chapter should be relevant to about 14% of all questions about biochemistry on the MCAT.
This chapter covers material from the following AAMC content categories:
1A: Structure and function of proteins and their constituent amino acids
5E: Principles of chemical thermodynamics and kinetics
Introduction
Obesity is increasingly common in the United States and is paralleled by an increase in high blood pressure, or hypertension. This is extremely relevant to medical students because hypertension increases the risk of stroke, heart failure, and kidney failure.
Each year, physicians encourage millions of Americans to improve their diets, add exercise to their daily regimens, or even take prescription drugs to control their hypertension. Many of these anti-hypertensive medications are called ACE (angiotensin-converting enzyme) inhibitors. In healthy patients, ACE catalyzes a reaction that converts a peptide called angiotensin I to angiotensin II. The angiotensin II peptide then not only directly causes constriction of the blood vessels to raise blood pressure, but also stimulates the release of the hormone aldosterone, which activates the kidneys to reabsorb more water back into the bloodstream. The increase in blood volume also increases blood pressure. Physicians take advantage of this complicated pathway with a straightforward solution: stop the pathway early by inhibiting ACE and blood pressure will decrease.
Enzymes are crucial proteins that dramatically increase the rate of biological reactions. They’re used to regulate homeostatic mechanisms in every organ system and are highly regulated themselves by environmental conditions, activators, and inhibitors. These regulators may be naturally occurring or may be given as a drug, such as the ACE inhibitors used to treat hypertension. Some enzymes are kept in an inactivated form called a zymogen and are only activated as needed. In this chapter, we’ll learn about how enzymes work and how different conditions influence their activity. We’ll also see how enzymes are regulated, which will help us tie together concepts about every organ system and metabolic process we learn about for the MCAT.
2.1 Enzymes as Biological Catalysts
LEARNING OBJECTIVES
After Chapter 2.1, you will be able to:
- Explain the major features of enzyme function, including specificity and catalysis
- Describe all seven classes of enzymes: oxidoreductases, transferases, hydrolases, lyases, isomerases, ligases, and translocases
- Recognize how enzymes affect thermodynamics and kinetics of a reaction
Enzymes are incredibly important as biological catalysts. Catalysts do not impact the thermodynamics of a biological reaction; that is, the ΔHrxn and equilibrium position do not change. Instead, they help the reaction proceed at a much faster rate. As a catalyst, the enzyme is not changed during the course of the reaction. Enzymes increase the reaction rate of a process by a factor of 100, 1000, or even 1,000,000,000,000 (1012) times when compared to the uncatalyzed reaction. Without this increase, we wouldn’t be alive. Table 2.1 summarizes the key points to remember about enzymes.
Table 2.1 Key Features of Enzymes
Lower the activation energy
Increase the rate of the reaction
Do not alter the equilibrium constant
Are not changed or consumed in the reaction (which means that they will appear in both the reactants and products)
Are pH- and temperature-sensitive, with optimal activity at specific pH ranges and temperatures
Do not affect the overall ΔG of the reaction
Are specific for a particular reaction or class of reactions
Enzyme Classifications
Enzymes are picky. The molecules upon which an enzyme acts are called substrates; a given enzyme will only catalyze a single reaction or class of reactions with these substrates, a property known as enzyme specificity. For example, urease only catalyzes the breakdown of urea. Chymotrypsin, on the other hand, can cleave peptide bonds around the amino acids phenylalanine, tryptophan, and tyrosine in a variety of polypeptides. Although those amino acids aren’t identical, they all contain an aromatic ring, which makes chymotrypsin specific for a class of molecules.
Enzymes can be classified into seven categories, based on their function or mechanism. We’ll review each type of enzyme and give examples of those that you are most likely to see on Test Day. If you encounter an unfamiliar enzyme on the MCAT, keep in mind that most enzymes have descriptive names ending in the suffix –ase: lactase, for example, breaks down lactose.
Oxidoreductases
Oxidoreductases catalyze oxidation–reduction reactions; that is, the transfer of electrons between biological molecules. They often have a cofactor that acts as an electron carrier, such as NAD+ or NADP+. In reactions catalyzed by oxidoreductases, the electron donor is known as the reductant, and the electron acceptor is known as the oxidant. Enzymes with dehydrogenase or reductase in their names are usually oxidoreductases. Enzymes in which oxygen is the final electron acceptor often include oxidase in their names. Malate dehydrogenase is an example of an oxidoreductase; it oxidizes malate to oxaloacetate or reduces oxaloacetate to malate, as seen in Figure 2.1.
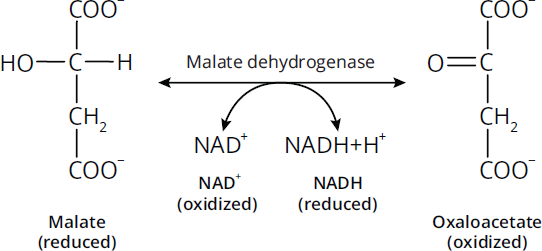
Figure 2.1 Oxidoreductases Catalyze Oxidation-Reduction Reactions
BRIDGE
The convention for naming reductants and oxidants of oxidoreductases is the same as the convention for naming reducing agents and oxidizing agents in general and organic chemistry. This is a good time to brush up on oxidation–reduction reactions if you haven’t seen them in a while—they’re covered in Chapter 11 of MCAT General Chemistry Review and Chapter 4 of MCAT Organic Chemistry Review.
Transferases
Transferases catalyze the movement of a functional group from one molecule to another. For example, in protein metabolism, an aminotransferase can convert aspartate and α-ketoglutarate, as a pair, to glutamate and oxaloacetate by moving the amino group from aspartate to α-ketoglutarate. Most transferases will be straightforwardly named, but remember that kinases are also a member of this class. Kinases catalyze the transfer of a phosphate group, generally from ATP, to another molecule. Figure 2.2 shows an example of the transferase (and kinase) hexokinase transferring ATP onto a glucose molecule.
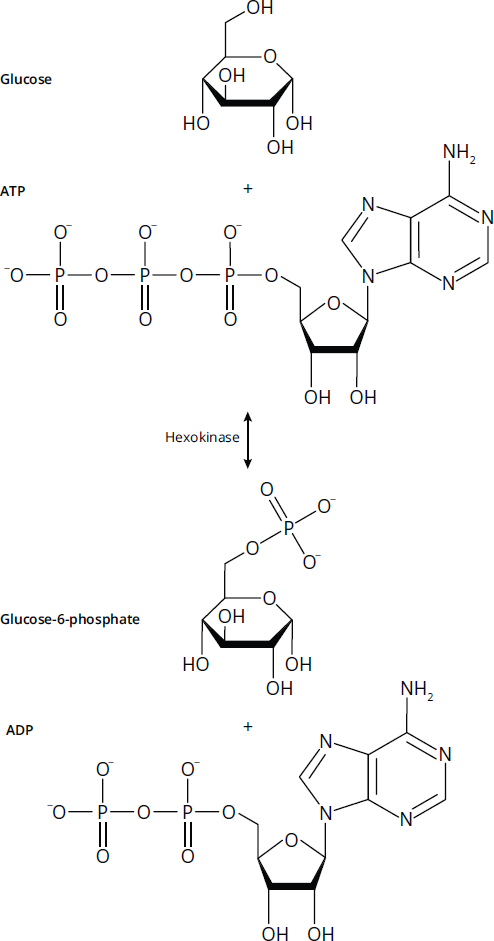
Figure 2.2 Transferases Move Functional Groups between Molecules
Hydrolases
Hydrolases catalyze the breaking of a compound into two molecules using the addition of water, as seen in Figure 2.3. In common usage, many hydrolases are named only for their substrate. For example, one of the most common hydrolases you will encounter on the MCAT is a phosphatase, which cleaves a phosphate group from another molecule. Other hydrolases include peptidases,nucleases, and lipases, which break down proteins, nucleic acids, and lipids, respectively.
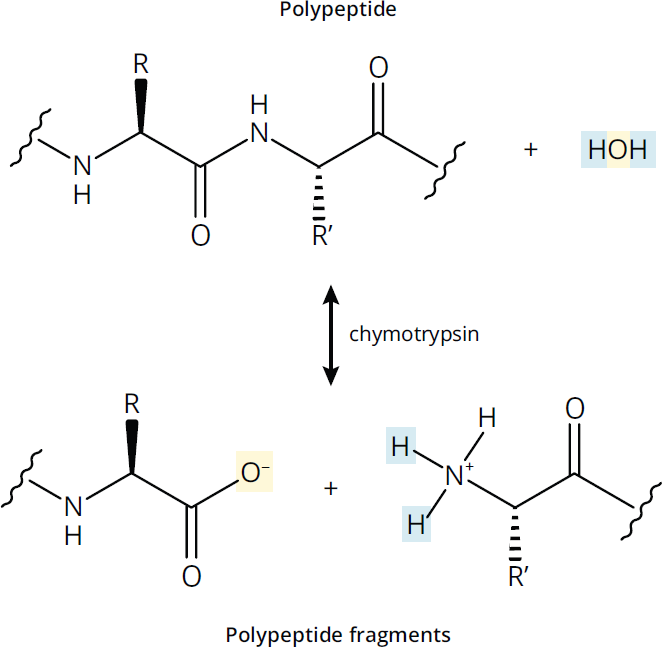
Figure 2.3 Hydrolases Use Water to Cleave Bonds
Lyases
Lyases catalyze the cleavage of a single molecule into two products without the use of water and without acting as oxidoreductases. Adenylyl cyclase (AC), shown converting ATP into cyclic AMP in Figure 2.4, is one example of a lyase. Because most enzymes can also catalyze the reverse of their specific reactions, the synthesis of two molecules into a single molecule may also be catalyzed by a lyase. When fulfilling this function, it is common for them to be referred to as synthases.
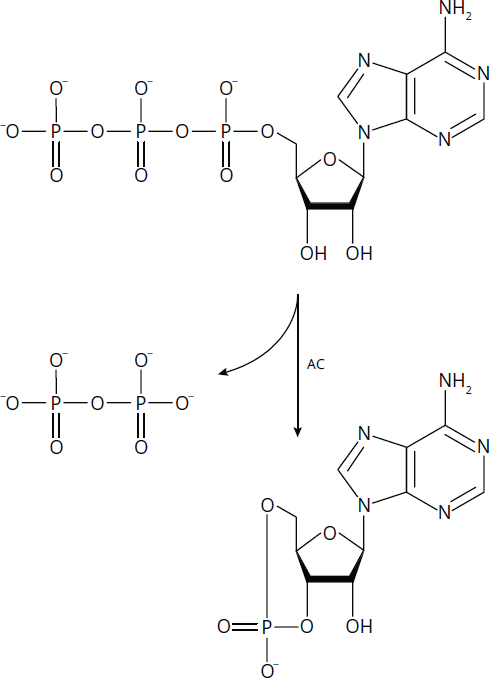
Figure 2.4 Lyases Like Adenylyl Cyclase (AC) Cleave Bonds without Using Water
Isomerases
Isomerases catalyze the rearrangement of bonds within a molecule. Some isomerases can also be classified as oxidoreductases, transferases, or lyases, depending on the mechanism of the enzyme. Keep in mind that isomerases catalyze reactions between stereoisomers as well as constitutional isomers. Figure 2.5 shows the conversion between the isomers citrate and isocitrate, which is catalyzed by the isomerase aconitase.
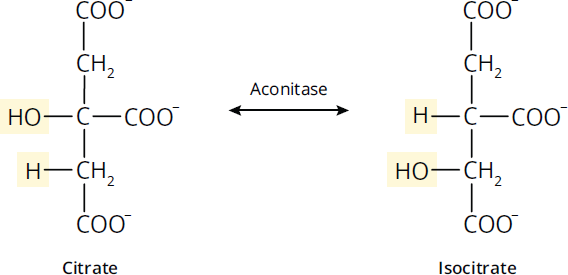
Figure 2.5 Isomerases Rearrange Bonds within Molecules
Ligases
Ligases catalyze addition or synthesis reactions, generally between large, similar molecules, and often require ATP. Synthesis reactions with smaller molecules are generally accomplished by lyases. Ligases are most likely to be encountered in nucleic acid synthesis and repair on Test Day. An example of a ligase is the enzyme acetyl-CoA carboxylase, shown in Figure 2.6, which adds a molecule of CO2 onto acetyl-CoA to create malonyl-CoA.
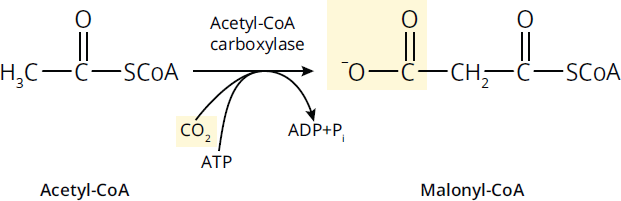
Figure 2.6 Ligases Join Large Molecules Together
MNEMONIC
Major Enzyme Classifications: LI’L HOTT
- Ligase
- Isomerase
- Lyase
- Hydrolase
- Oxidoreductase
- Transferase
- Translocase
Translocases
Enzymes originally were classified into six groups: oxidoreductases, transferases, hydrolases, lyases, isomerases, and ligases, which are described in more detail above. However, certain important enzymes did not properly fit into any of those six classes, so as of 2018, a seventh type was recognized: translocases. Translocases are proteins that help catalyze the transport of molecules across a cell membrane. ATP synthase, shown in Figure 2.7, is an important enzyme that falls under this class. To create ATP, the FO subunit of ATP synthase translocates protons across the inner mitochondrial membrane, where the energy from the electrochemical proton gradient is used by the F1 subunit of ATP synthase to create ATP.
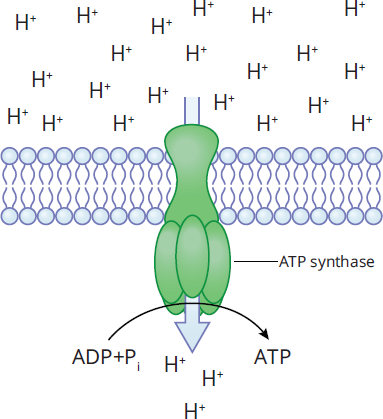
Figure 2.7 Translocases Move Molecules across Membranes
Impact on Activation Energy
Recall that thermodynamics relates the relative energy states of a reaction in terms of its products and reactants. An endergonic reaction is one that requires energy input (ΔG > 0), whereas an exergonic reaction is one in which energy is given off (ΔG < 0). Remember that endo–means “in” and exo–means “out,” so endergonic reactions take in energy as they proceed, whereas exergonic reactions release energy as they proceed. We can look at the reaction diagram in Figure 2.8 to see this demonstrated more clearly.
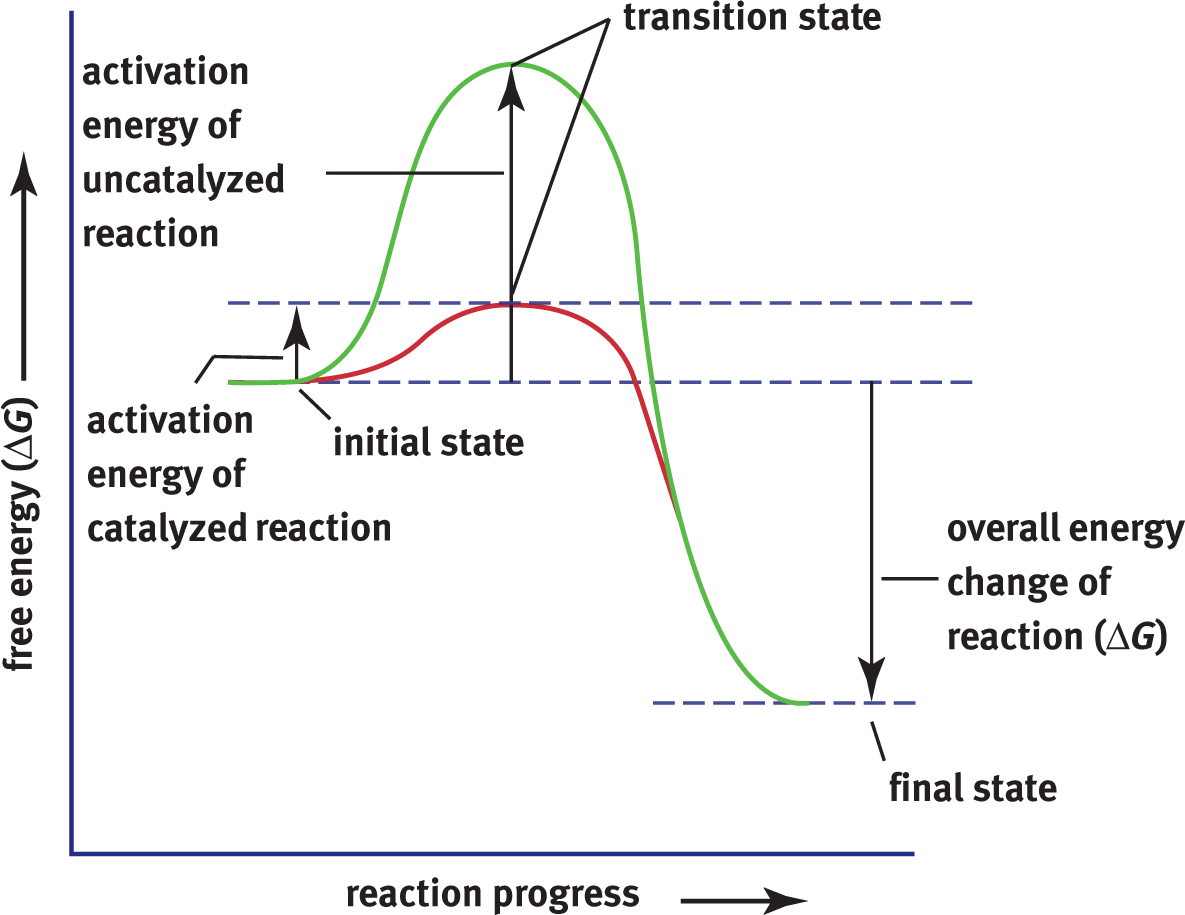
Figure 2.8 Exergonic Reaction Diagram The activation energy required for a catalyzed reaction is lower than that of an uncatalyzed reaction while the ΔG(and ΔH) remains the same.
BRIDGE
Gibbs free energy and endergonic/exergonic reactions are covered in Chapter 7 of MCAT General Chemistry Review.
The reaction shown in Figure 2.8 is spontaneous. Note that the ΔGfor this reaction is negative. A very important characteristic of enzymes is that they do not alter the overall free energy change for a reaction, nor do they change the equilibrium of a reaction. Rather, they affect the rate (kinetics) at which a reaction occurs; thus, they can affect how quickly a reaction gets to equilibrium but not the actual equilibrium state itself. For example, a reaction could take years to approach equilibrium without an enzyme. In comparison, with the enzyme, equilibrium might be attained within seconds. Enzymes ensure that many important reactions can occur in a reasonable amount of time in biological systems. Recall that enzymes, as catalysts, are unchanged by the reaction. What is the functional consequence of this? Far fewer copies of the enzyme are required relative to the overall amount of substrate because one enzyme can act on many, many molecules of substrate over time.
Catalysts exert their effect by lowering the activation energy of a reaction; in other words, they make it easier for the substrate to reach the transition state. Imagine having to walk to the other side of a tall hill. The only way to get there is to climb to the top of the hill and then walk down the other side—but wouldn’t it be easier if the top of the hill was cut off so one wouldn’t have to climb so high? That’s exactly what catalysts do for chemical reactions when they make it easier for substrates to achieve their transition state. Most reactions catalyzed by enzymes are technically reversible, although that reversal may be extremely energetically unfavorable and therefore essentially nonexistent.
MCAT CONCEPT CHECK 2.1:
Before you move on, assess your understanding of the material with these questions.
- How do enzymes function as biological catalysts?
________________________________________
- What is enzyme specificity?
________________________________________
- What are the names and main functions of the seven different classes of enzymes?
Name Function
-
In what ways do enzymes affect the thermodynamics vs. the kinetics of a reaction?
________________________________________
2.2 Mechanisms of Enzyme Activity
LEARNING OBJECTIVES
After Chapter 2.2, you will be able to:
- Differentiate between coenzymes and cofactors
- Compare the induced fit and lock and key models of enzyme function
While enzyme mechanisms will vary depending on the reaction that is being catalyzed, they tend to share some common features. Enzymes may act to provide a favorable microenvironment in terms of charge or pH, stabilize the transition state, or bring reactive groups nearer to one another in the active site. The formation of the enzyme–substrate complex in the active site of an enzyme is the key catalytic activity of the enzyme, which reduces the activation energy of the reaction as described above. This interaction between a substrate and the active site of an enzyme also accounts for the selectivity and some regulatory mechanisms of enzymes.
Enzyme–Substrate Binding
The molecule upon which an enzyme acts is known as its substrate. The physical interaction between these two is referred to as the enzyme–substrate complex. The active site is the location within the enzyme where the substrate is held during the chemical reaction, as shown in Figure 2.9.
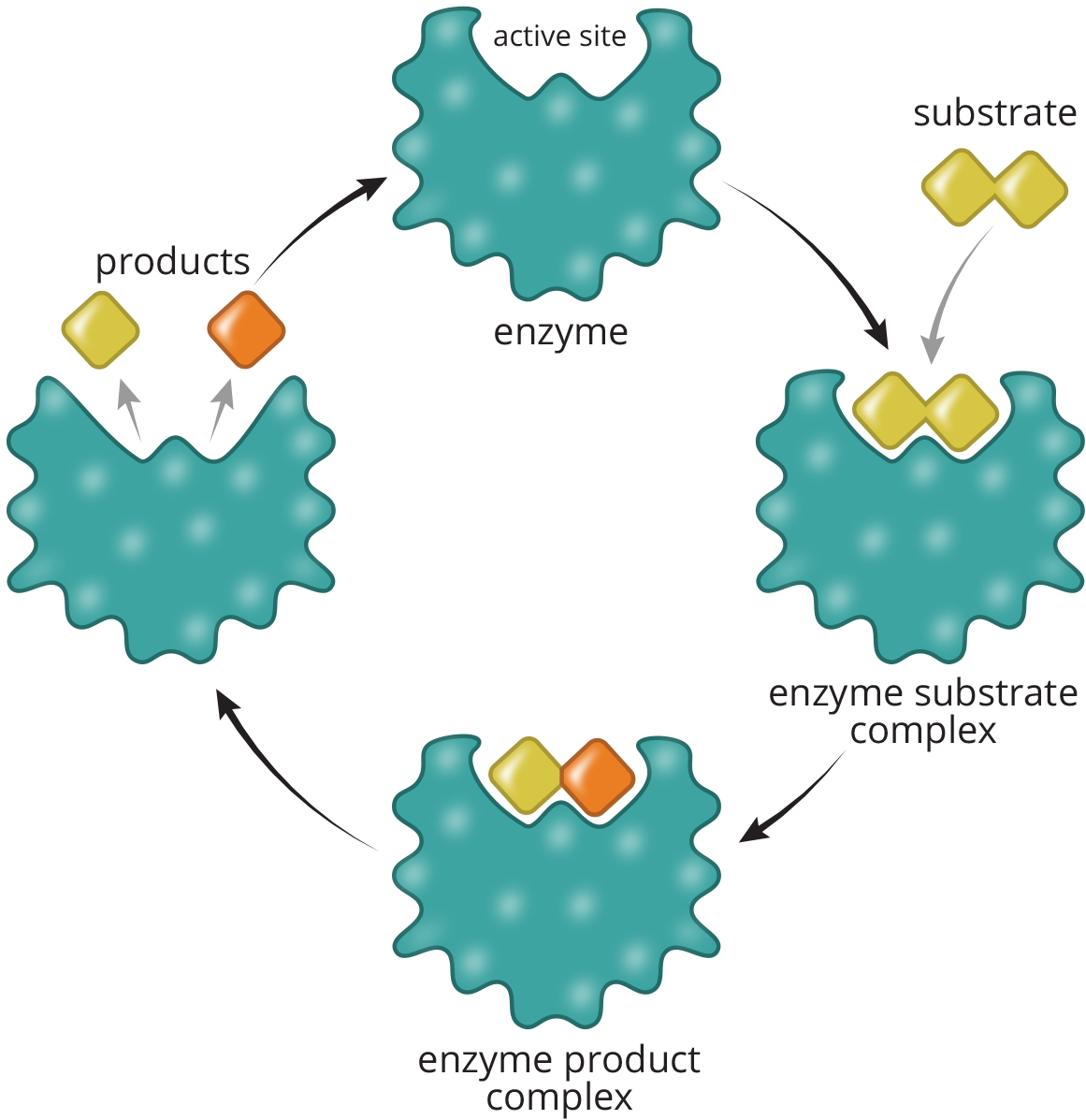
Figure 2.9 Reaction Catalysis in the Active Site of an Enzyme
The active site assumes a defined spatial arrangement in the enzyme–substrate complex, which dictates the specificity of that enzyme for a molecule or group of molecules. Hydrogen bonding, ionic interactions, and transient covalent bonds within the active site all stabilize this spatial arrangement and contribute to the efficiency of the enzyme. Two competing theories explain how enzymes and substrates interact, but one of the two is better supported than the other.
Lock and Key Theory
The lock and key theory is aptly named. It suggests that the enzyme’s active site (lock) is already in the appropriate conformation for the substrate (key) to bind. As shown in Figure 2.10, the substrate can then easily fit into the active site, like a key into a lock or a hand into a glove. No alteration of the tertiary or quaternary structure is necessary upon binding of the substrate.
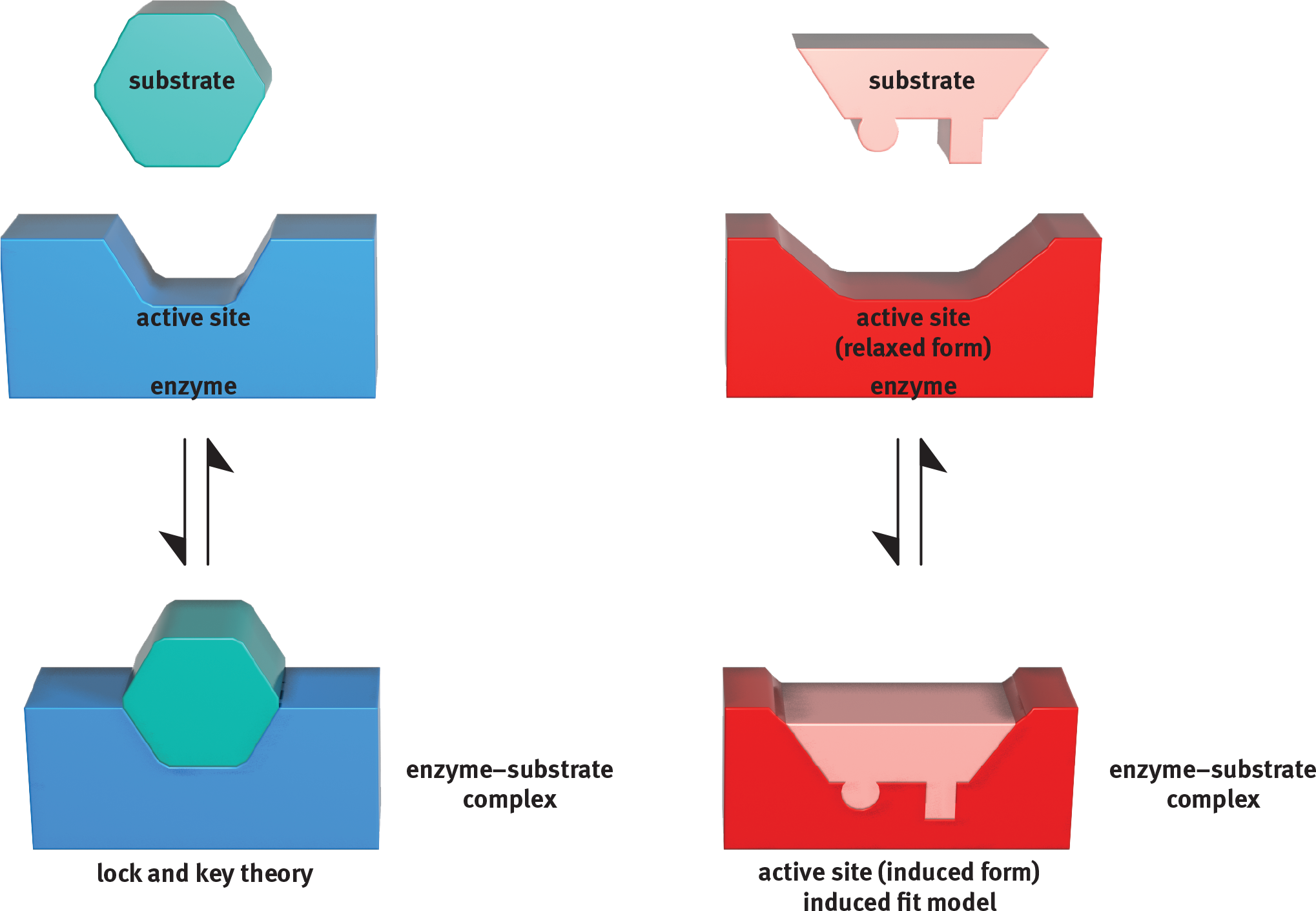
Figure 2.10 Lock and Key Theory vs. Induced Fit Model for Enzyme Catalysis
Induced Fit Model
The more scientifically accepted theory is the induced fit model; this is the one you are more likely to see referenced on Test Day. Imagine that the enzyme is a foam stress ball and the substrate is a frustrated MCAT student’s hand. What’s the desired interaction? The student wants to release some stress and relax. As the student’s hand squeezes the ball, both change conformation. The ball is no longer spherical and the hand is no longer flat because they adjust to fit each other well. In this case, the substrate (the student) has induced a change in the shape of the enzyme (the stress ball). This interaction requires energy, and therefore, this part of the reaction is endergonic. Once the student lets go of the stress ball, we have our desired product: a relaxed, more confident test taker. Letting go of the stress ball is pretty easy and doesn’t require extra energy; so, this part of the reaction is exergonic. Just like enzymes, foam stress balls return to their original shape once their crushers (substrates) let go of them. On a molecular level, demonstrated in Figure 2.10, the induced fit model starts with a substrate and an enzyme active site that don’t seem to fit together. However, once the substrate is present and ready to interact with the active site, the molecules find that the induced form, or transition state, is more comfortable for both of them. Thus, the shape of the active site becomes truly complementary only after the substrate begins binding to the enzyme. Similarly, a substrate of the wrong type will not cause the appropriate conformational shift in the enzyme. Thus, the active site will not be adequately exposed, the transition state is not preferred, and no reaction occurs.
Cofactors and Coenzymes
Many enzymes require nonprotein molecules called cofactors or coenzymes to be effective. These cofactors and coenzymes tend to be small in size so they can bind to the active site of the enzyme and participate in the catalysis of the reaction, usually by carrying charge through ionization, protonation, or deprotonation. Cofactors and coenzymes are usually kept at low concentrations in cells, so they can be recruited only when needed. Enzymes without their cofactors are called apoenzymes, whereas those containing them are holoenzymes. Cofactors are attached in a variety of ways, ranging from weak noncovalent interactions to strong covalent ones. Tightly bound cofactors or coenzymes that are necessary for enzyme function are known as prosthetic groups.
REAL WORLD
Deficiencies in vitamin cofactors can result in devastating disease. Thiamine is an essential cofactor for several enzymes involved in cellular metabolism and nerve conduction. Thiamine deficiency, often a result of excess alcohol consumption and poor diet, results in diseases including Wernicke–Korsakoff syndrome. In this disorder, patients suffer from a variety of neurologic deficits, including delirium, balance problems, and, in severe cases, the inability to form new memories.
Cofactors and coenzymes are topics that we are likely to see on Test Day, so they are important to know. Cofactors are generally inorganic molecules or metal ions, and are often ingested as dietary minerals. Coenzymes are small organic groups, the vast majority of which are vitamins or derivatives of vitamins such as NAD+, FAD, and coenzyme A. The water-soluble vitamins include the B complex vitamins and ascorbic acid (vitamin C), and are important coenzymes that must be replenished regularly because they are easily excreted. The fat-soluble vitamins—A, D, E, and K—are better regulated by partition coefficients, which quantify the ability of a molecule to dissolve in a polar vs. nonpolar environment. Enzymatic reactions are not restricted to a single cofactor or coenzyme. For example, metabolic reactions often require magnesium, NAD+ (derived from vitamin B3), and biotin (vitamin B7) simultaneously.
BRIDGE
Vitamins come in two major classes: fat- and water-soluble. This is important to consider in digestive diseases, where different parts of the gastrointestinal tract may be affected by different disease processes. Because different parts of the gastrointestinal tract specialize in the absorption of different types of biomolecules, loss of different parts of the gastrointestinal tract or its accessory organs may result in different vitamin deficiencies. The digestive system is discussed in Chapter 9 of MCAT Biology Review.
Memorization of the B vitamins is low yield for the MCAT, though testable; however, familiarity with their names may make biochemistry passages easier on Test Day:
- B1: thiamine
- B2: riboflavin
- B3: niacin
- B5: pantothenic acid
- B6: pyridoxal phosphate
- B7: biotin
- B9: folic acid
- B12: cyanocobalamin
MCAT CONCEPT CHECK 2.2:
Before you move on, assess your understanding of the material with these questions.
- How do the lock and key theory and induced fit model differ?
Lock and Key Induced Fit
- What do cofactors and coenzymes do? How do they differ?
________________________________________
2.3 Enzyme Kinetics
LEARNING OBJECTIVES
After Chapter 2.3, you will be able to:
- Predict how changes in enzyme and solute concentration will affect enzyme kinetics
- Define enzyme cooperativity
- Compare Lineweaver–Burk and Michaelis–Menten plots
- Explain key points on a Lineweaver–Burk or Michaelis–Menten plot:
Kinetics of Monomeric Enzymes
Enzyme kinetics is a high-yield topic that can score us several points on Test Day. Just as the relief our student derives from squeezing a stress ball depends on a number of factors, such as size and shape of the ball and the student’s baseline level of stress, enzyme kinetics are dependent on factors like environmental conditions and concentrations of substrate and enzyme.
The concentrations of the substrate, [S], and enzyme, [E], greatly affect how quickly a reaction will occur. Let’s say that we have 100 stress balls (enzymes) and only 10 frustrated students (substrates) to derive stress relief from them. This represents a high enzyme concentration relative to substrate. Because there are many active sites available, we will quickly form products (students letting go and feeling relaxed); in a chemical sense, we would reach equilibrium quickly. As we slowly add more substrate (students), the rate of the reaction will increase; that is, more people will relax in the same amount of time because we have plenty of available stress balls for them to squeeze. However, as we add more and more people (and start approaching 100 students), we begin to level off and reach a maximal rate of relaxation. There are fewer and fewer available stress balls until finally all active sites are occupied. Unlike before, inviting more students into the room will not change the rate of the reaction. It cannot go any faster once it has reached saturation. At this rate, the enzyme is working at maximum velocity, denoted by ***V*max**. The only way to increase Vmax is by increasing the enzyme concentration. In the cell, this can be accomplished by inducing the expression of the gene encoding the enzyme. These concepts are represented graphically in Figure 2.11.
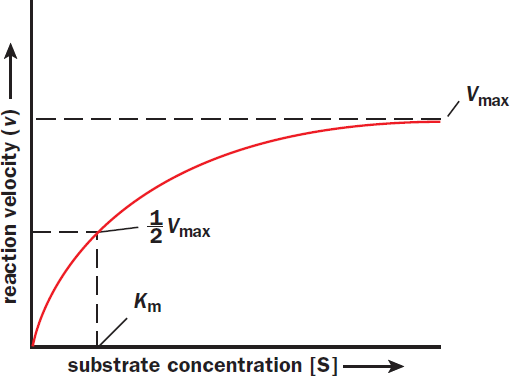
Figure 2.11 Michaelis–Menten Plot of Enzyme Kinetics As the amount of substrate increases, the enzyme is able to increase its rate of reaction until it reaches a maximum enzymatic reaction rate (Vmax); once Vmax is reached, adding more substrate will not increase the rate of reaction.
Michaelis–Menten Kinetics
For most enzymes with one substrate, the Michaelis–Menten equation describes how the rate of the reaction, v, depends on the concentration of both the enzyme, [E], and the substrate, [S], to form the product, [P]. The equation presumes a simple two-step reaction: enzyme–substrate complexes form at a rate k1; then the ES complex can either dissociate at a rate k–1 or turn into E + P at a rate kcat:
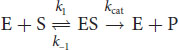
Equation 2.1
Note that in either case, the enzyme is regenerated. Michaelis-Menten kinetics presumes a set of assumptions and conditions:
- Measurements of the initial enzyme velocity (vo) must be at early time points, where [P] = 0.
- The concentration of substrate must be in excess of the concentration of enzyme ([S] >> [E]) so that [S] is effectively constant at early time points.
- The binding of E to S (E + S ⇌ ES) is fast, allowing for [E], [S], and [ES] to reach steady state and therefore equilibrium ratios. The catalytic step (ES → P) is slow and rate-limiting.
- There are only two forms of the enzyme: E and ES. The total concentration of enzyme [ET] is the sum of the free enzyme and enzyme-substrate complex ([ET] = [E] + [ES]).
On Test Day, the concentration of enzyme will be kept constant unless otherwise stated. Under these conditions, we can relate the velocity of the enzyme to substrate concentration using the Michaelis-Menten equation:
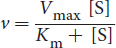
Equation 2.2
Some important and Test Day–relevant math can be derived from this equation. When the reaction rate is equal to half of Vmax, Km = [S]:
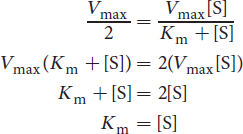
Km can, therefore, be understood to be the substrate concentration at which half of the enzyme’s active sites are full (half the stress balls are in use). Km is the Michaelis constant and is often used to compare enzymes. Under certain conditions, Km is a measure of the affinity of the enzyme for its substrate. When comparing two enzymes, the one with the higher Km has the lower affinity for its substrate because it requires a higher substrate concentration to be half-saturated. The Km value is an intrinsic property of the enzyme–substrate system and cannot be altered by changing the concentration of substrate or enzyme.
KEY CONCEPT
We can assess an enzyme’s affinity for a substrate by noting the Km. A low Km reflects a high affinity for the substrate (low [S] required for 50% enzyme saturation). Conversely, a high Km reflects a low affinity of the enzyme for the substrate.
For a given concentration of enzyme, the Michaelis–Menten relationship generally graphs as a hyperbola, as seen in the Michaelis–Menten plot in Figure 2.11. When substrate concentration is less than Km, changes in substrate concentration will greatly affect the reaction rate. At high substrate concentrations exceeding Km, the reaction rate increases much more slowly as it approaches Vmax, where it becomes independent of substrate concentration.
The variable Vmax represents maximum enzyme velocity and is measured in moles of enzyme per second. Also, Vmax can be mathematically related to kcat, which has units of s–1:
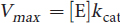
Equation 2.3
Qualitatively speaking, kcat measures the number of substrate molecules “turned over,” or converted to product, per enzyme molecule per second. Most enzymes have kcat values between 101 and 103. The Michaelis–Menton equation above can be restated using kcat:
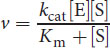
At very low substrate concentrations, where Km >> [S], this derived equation can be further simplified as:
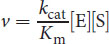
The ratio of kcat/Km is referred to as the catalytic efficiency of the enzyme. A large kcat (high turnover) or a small Km (high substrate affinity) will result in a higher catalytic efficiency, which indicates a more efficient enzyme.
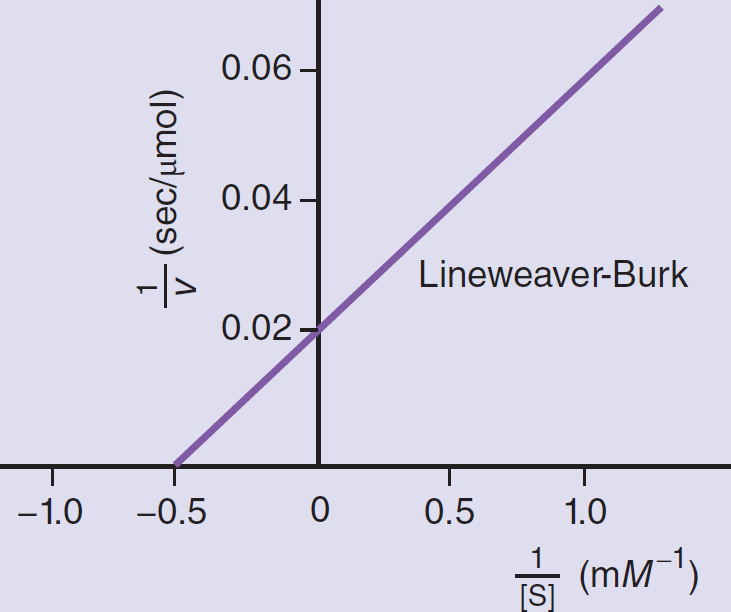
Lineweaver–Burk Plots
The Lineweaver–Burk plot is a double reciprocal graph of the Michaelis–Menten equation. The same data graphed in this way yield a straight line as shown in Figure 2.12. The actual data are represented by the portion of the graph to the right of the y-axis, but the line is extrapolated into the upper left quadrant to determine its intercept with the x-axis. The intercept of the line with the x-axis gives the value of
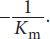
. The intercept of the line with the y-axis gives the value of
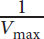
. The Lineweaver–Burk plot is especially useful when determining the type of inhibition that an enzyme is experiencing because Vmax and Km can be compared without estimation.
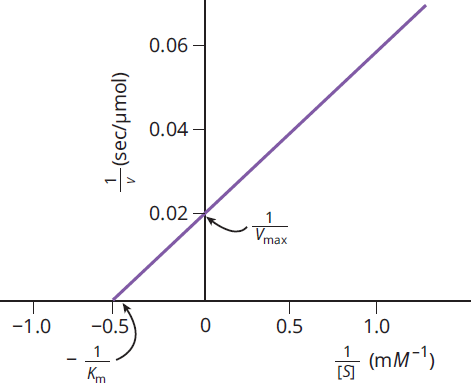
Figure 2.12 Experimentally Determined Lineweaver–Burk (Double Reciprocal) Plot Used to Calculate the Values of Km and Vmax
Cooperativity
Certain enzymes do not show the normal hyperbola when graphed on a Michaelis–Menten plot (vvs. [S]), but rather show sigmoidal (S-shaped) kinetics owing to cooperativity among substrate binding sites, as shown in Figure 2.13. Cooperative enzymes have multiple subunits and multiple active sites. Subunits and enzymes may exist in one of two states: a low-affinity tense state (T) or a high-affinity relaxed state (R). Binding of the substrate encourages the transition of other subunits from the T state to the R state, which increases the likelihood of substrate binding by these other subunits. Conversely, loss of substrate can encourage the transition from the R state to the T state, and promote dissociation of substrate from the remaining subunits. Think of cooperative enzyme kinetics like a party. As more people start arriving, the atmosphere becomes more relaxed and the party seems more appealing, but as people start going home the party dies down and more people are encouraged to leave so the tense hosts can clean up. Enzymes showing cooperative kinetics are often regulatory enzymes in pathways, like phosphofructokinase-1 in glycolysis. Cooperative enzymes are also subject to activation and inhibition, both competitively and through allosteric sites.
MCAT EXPERTISE
The cooperative binding of hemoglobin, which acts as a transport protein rather than an enzyme, results in a characteristic sigmoidal binding curve that is an MCAT favorite.
Cooperativity can also be quantified using a numerical value called Hill's coefficient. The value of Hill's coefficient indicates the nature of binding by the molecule:
- If Hill's coefficient > 1, positively cooperative binding is occurring, such that after one ligand is bound the affinity of the enzyme for further ligand(s) increases.
- If Hill's coefficient < 1, negatively cooperative binding is occurring, such that after one ligand is bound the affinity of the enzyme for further ligand(s) decreases.
- If Hill's coefficient = 1, the enzyme does not exhibit cooperative binding.
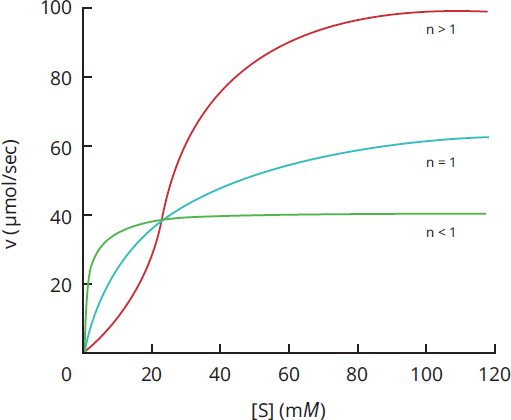
Figure 2.13 Cooperativity in Enzyme Kinetics
BIOCHEMISTRY GUIDED EXAMPLE WITH EXPERT THINKING
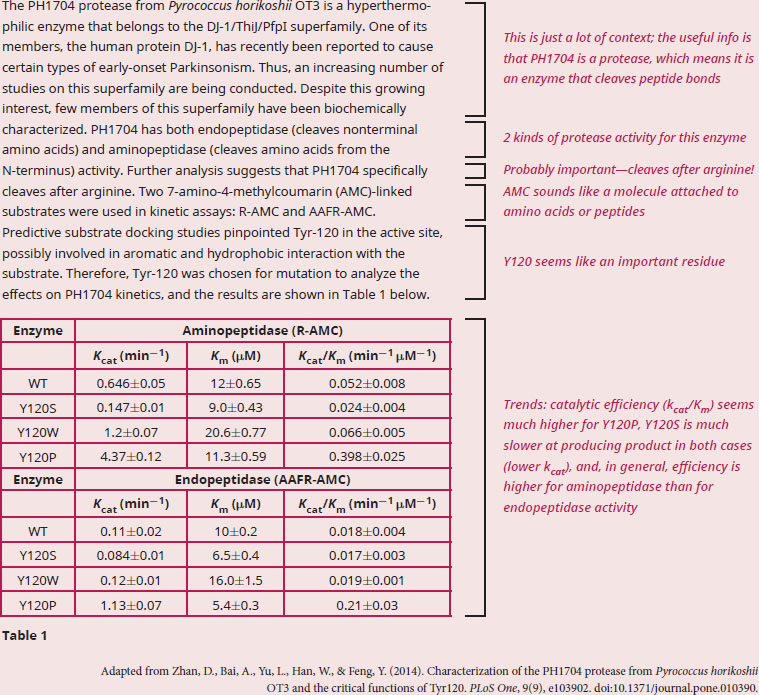
According to the data, does PH1704 have equivalent aminopeptidase and endopeptidase activity? Why or why not?
The question asks us for data analysis, so we know we should start by making sure we understand what the given material is telling us. A quick scan of the passage reveals unfamiliar enzymes, terminology, and a data-packed table, so a careful read of the passage for the big picture will be important in order to relate what the table is trying to prove. We are also definitely going to need to bring in some of our outside knowledge about enzymes. The passage describes an enzyme called PH1704, which has several characteristics outlined in the first half of the paragraph. We’re told it’s a protease, which means it breaks peptide bonds, and it’s hyperthermophilic. Breaking down the term hyperthermophilic, we get hyper-(higher)-thermo-(heat)-philic (love), and we can thus deduce that the enzyme is from a bacterial species that lives in high temperatures. Next, we’re told it’s an endopeptidase (endo- meaning between) and an aminopeptidase with a strong preference for arginine. There are two substrates described in the study. Notice how only R-AMC can be used to discern the aminopeptidase activity, and AAFR-AMC can be used to test endopeptidase activity. Finally, we’re told that Tyr-120 can be important for activity, so researchers created mutations of this residue and the kinetic information of the resulting mutants is described in Table 1.
The question asks us to determine whether PH1704 acts equally as an aminopeptidase and an endopeptidase. In order to answer this question, first note that the mutation data for Y120 is not needed since we are only being asked about the wild-type PH1704 enzyme.
All we really need is a comparison between the WT enzyme with R-AMC as a substrate, and WT enzyme with AAFR-AMC as a substrate. Also, we need to determine whether kcat, km, or kcat/km will be most useful. Recall that kcat provides information about how much product is produced per unit time, and km is a measure of substrate affinity. To have an enzyme that is maximally efficient, you want both a high catalytic turnover (high kcat) and a high substrate affinity (low km), so the best comparison would be the kcat/km data, which is equal to the catalytic efficiency of an enzyme. The catalytic efficiency for PH1704 as an aminopeptidase is 0.052 min–1/μM, and for PH1704 as an endopeptidase efficiency is 0.018 min–1/μM.
Since these values of catalytic efficiency are different when measuring aminopeptidase and endopeptidase activity, we can conclude that PH1704 functions more effectively as an aminopeptidase than as an endopeptidase.
Multi-Substrate Reactions
The Michaelis-Menten model of enzyme kinetics deals with single substrate reactions, but the majority of known enzymatic reactions involve more than one substrate. Some of the multi-substrate reactions are detailed in Figure 2.14.
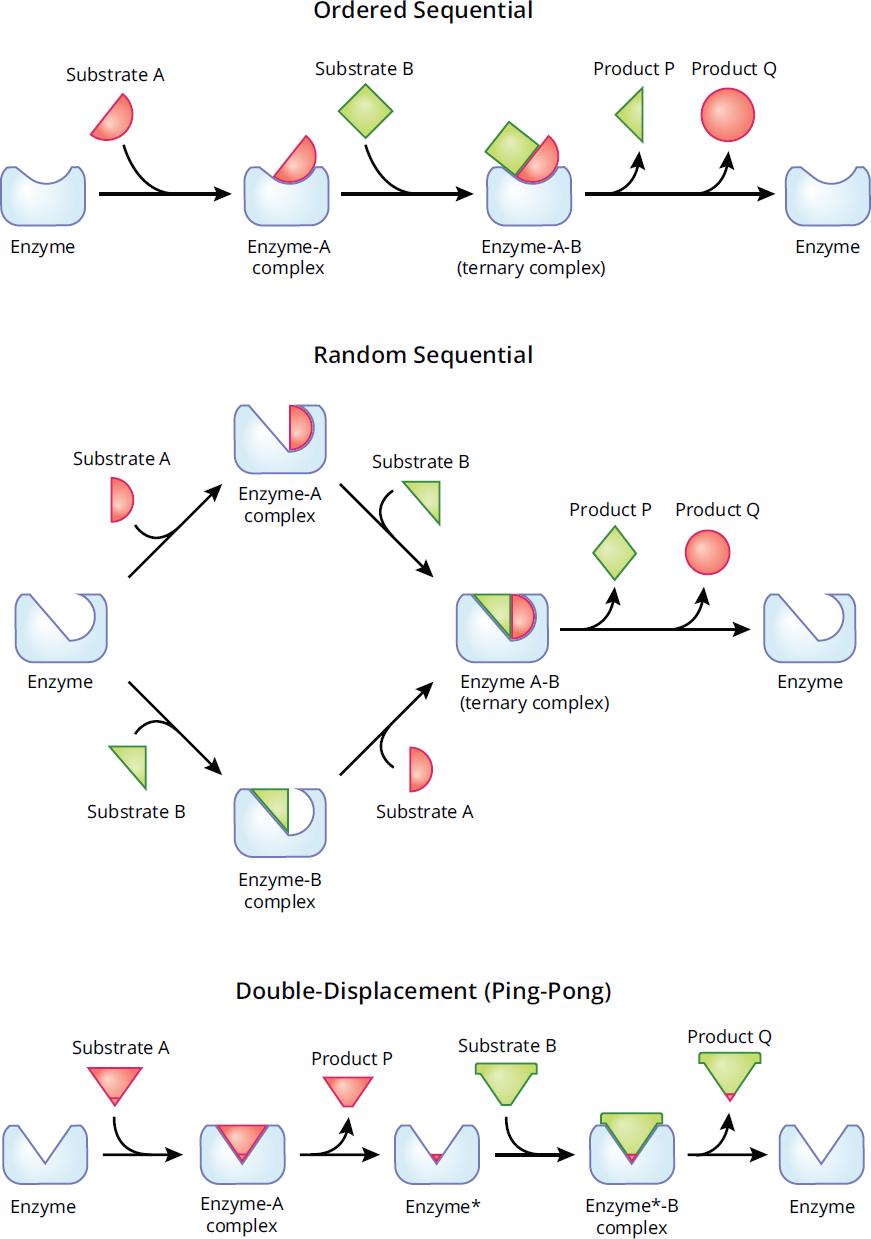
Figure 2.14 Multi-Substrate Reaction Mechanisms
In an ordered sequential reaction, substrates bind to the enzyme in a specific sequence. The conversion of pyruvate into lactate by lactate dehydrogenase demonstrates this type of reaction mechanism: Coenzyme NADH binds to lactate dehydrogenase first, and only then can pyruvate bind for the catalyzation of NADH into NAD+ and pyruvate into lactate.
In contrast to an ordered sequential reaction, a random sequential reaction is unordered, meaning that the substrates and products can bind and release without a preference in order. The enzyme creatinine kinase follows this mechanism, where either ATP or creatinine can first bind the active site to make ADP and phosphocreatine.
In a double-displacement (ping-pong) reaction, one substrate binds first to the enzyme and is turned into product, leaving behind an intermediate form of the enzyme. The modified enzyme can then bind a second substrate and turn it into product. The formation of phosphocholine from choline by choline kinase occurs by this mechanism, where ATP binds to choline kinase, resulting in the formation of ADP and phosphorylation of choline kinase. This modified enzyme then binds to choline after ADP ejection, leading to phosphorylation of choline to create phosphocholine and dephosphorylation of the enzyme to revert it back to its original form.
MCAT CONCEPT CHECK 2.3:
Before you move on, assess your understanding of the material with these questions.
- What are the effects of increasing [S] on enzyme kinetics? What about increasing [E]?
- Increasing [S]:
________________________________________
- Increasing [E]:
________________________________________
- How are the Michaelis–Menten and Lineweaver–Burk plots similar? How are they different?
- Similarities:
________________________________________
- Differences:
________________________________________
-
What does Km represent? What would an increase in Km signify?
________________________________________
________________________________________
-
What do the x- and y-intercepts in a Lineweaver–Burk plot represent?
- x-intercept:
________________________________________
- y-intercept:
________________________________________
-
What is enzyme cooperativity?
________________________________________
2.4 Effects of Local Conditions on Enzyme Activity
LEARNING OBJECTIVES
After Chapter 2.4, you will be able to:
- Predict how changes to the environment will alter enzyme behavior
- Estimate the ideal pH and temperature for enzymes found in the human body
The activity of an enzyme is heavily influenced by its environment; in particular, temperature, acidity or alkalinity (pH), and high salinity have significant effects on the ability of an enzyme to carry out its function. Note that the terms enzyme activity, enzyme velocity, and enzyme rate are all used synonymously on the MCAT.
Temperature
Enzyme-catalyzed reactions tend to double in velocity for every 10 °C increase in temperature until the optimum temperature is reached; for the human body, this is 37 °C (98.6°F or 310 K). After this, activity falls off sharply, as the enzyme will denature at higher temperatures, as shown in Figure 2.15. Some enzymes that are overheated may regain their function if cooled. A real-life example of temperature dependence occurs in Siamese cats. Siamese cats are dark on their faces, ears, tails, and feet but white elsewhere. Why? The enzyme responsible for pigmentation, tyrosinase, is mutated in Siamese cats. It is ineffective at body temperature but at cooler temperatures becomes active. Thus, only the tail, feet, ears, and face (cooled by air passing through the nose and mouth) have an active form of the enzyme and are dark.
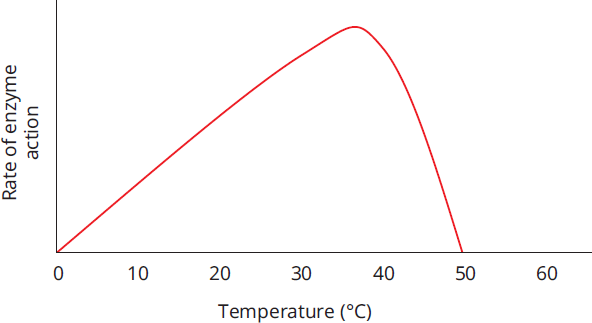
Figure 2.15 Effect of Temperature on the Rate of Enzyme Action
pH
Most enzymes also depend on pH in order to function properly, not only because pH affects the ionization of the active site, but also because changes in pH can lead to denaturation of the enzyme. For enzymes that circulate and function in human blood, this optimal pH is 7.4. A pH < 7.35 in human blood is termed acidemia. Even though it’s more basic than chemically neutral 7.0, it is more acidic than physiologically neutral 7.4. Where might exceptions to this pH 7.4 occur? Both are in our digestive tract. Pepsin, which works in the stomach, has maximal activity around pH 2, whereas pancreatic enzymes, which work in the small intestine, work best around pH 8.5. The graph in Figure 2.16 shows the different optimal pH ranges for the enzymes pepsin, salivary amylase, and trypsin.
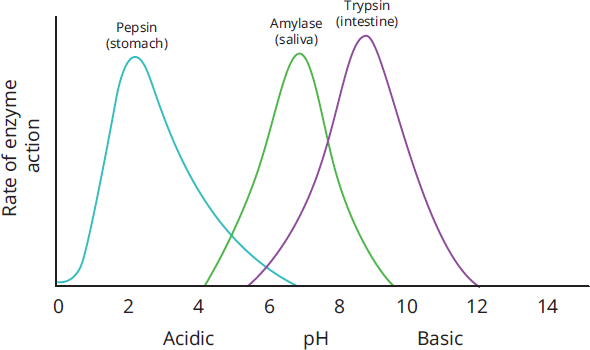
Figure 2.16 Different Enzymes Have Different Optimal pH Ranges
BRIDGE
The pH levels in the stomach and intestine, and their effects on these gastric and pancreatic enzymes, are covered in Chapter 9 of MCAT Biology Review.
Salinity
While the effect of salinity or osmolarity is not generally of physiologic significance, altering the concentration of salt can change enzyme activity in vitro. Increasing levels of salt can disrupt hydrogen and ionic bonds, causing a partial change in the conformation of the enzyme, and in some cases causing denaturation.
MCAT CONCEPT CHECK 2.4:
Before you move on, assess your understanding of the material with these questions.
- What are the effects of temperature, pH, and salinity on the function of enzymes?
- Temperature:
________________________________________
- pH:
________________________________________
- Salinity:
________________________________________
- What is the ideal temperature for most enzymes in the body? The ideal pH?
- Ideal temperature: ______ °C = ____ °F = ______ K
- Ideal pH (most enzymes): ________
- Ideal pH (gastric enzymes): ________
- Ideal pH (pancreatic enzymes): ________
2.5 Regulation of Enzyme Activity
LEARNING OBJECTIVES
After Chapter 2.5, you will be able to:
- Explain feedback inhibition and irreversible inhibition
- Differentiate between the four types of reversible inhibition
- Differentiate between transient and covalent enzyme modifications
- Recall the traits of zymogens
Although enzymes are useful, the body must be able to control when they work; for example, enzymes involved in mitosis should be shut off when cells are no longer dividing (in the G0 phase). This may be accomplished in a variety of ways, as described below.
Feedback Regulation
Enzymes are often subject to regulation by products further down a given metabolic pathway, a process called feedback regulation. Less often, enzymes may be regulated by intermediates that precede the enzyme in the pathway, also called feed-forward regulation. This is clearly evident in the study of metabolism, as discussed in Chapters 9 through 12 of MCAT Biochemistry Review. While there are some examples of feedback activation, feedback inhibition is far more common. Feedback inhibition, or negative feedback, helps maintain homeostasis: once we have enough of a given product, we want to turn off the pathway that creates that product, rather than creating more. In feedback inhibition, the product may bind to the active site of an enzyme or multiple enzymes that acted earlier in its biosynthetic pathway, thereby competitively inhibiting these enzymes and making them unavailable for use. This is schematically represented in Figure 2.17, as we see product D feeding back to inhibit the first enzyme in the pathway.

Figure 2.17 Feedback Inhibition by the Product of a Metabolic Pathway A high concentration of the product, D, inhibits enzyme 1, slowing the entire pathway.
BRIDGE
Negative feedback is an important topic in both enzymology and the endocrine system. Remember that most hormonal feedback loops are also inhibited by negative feedback. The endocrine system is discussed in Chapter 5 of MCAT Biology Review.
Reversible Inhibition
There are four types of reversible inhibition: competitive, noncompetitive, uncompetitive, and mixed. The MCAT typically tests the first three types of inhibition directly, while mixed inhibition is tested through exclusion by way of presenting an inhibitor that does not fall under the three prior categories. Table 2.2 at the end of this section summarizes the features of each type of reversible inhibition.
Competitive Inhibition
Competitive inhibition simply involves occupancy of the active site, as shown in Figure 2.18. Substrates cannot access enzymatic binding sites if there is an inhibitor in the way.
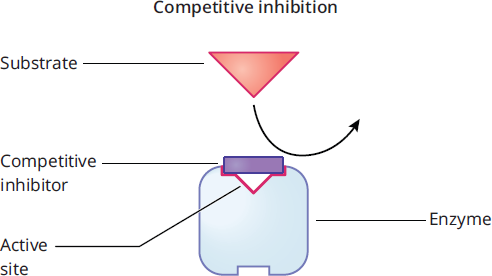
Figure 2.18 Competitive Inhibition
Competitive inhibition can be overcome by adding more substrate so that the substrate-to-inhibitor ratio is higher. If more molecules of substrate are available than molecules of inhibitor, then the enzyme will be more likely to bind substrate than inhibitor (assuming the enzyme has equal affinity for both molecules). Adding a competitive inhibitor does not alter the value of Vmax because if enough substrate is added, it will outcompete the inhibitor and be able to run the reaction at maximum velocity. A competitive inhibitor does increase the measured value of Km. This is because the substrate concentration has to be higher to reach half the maximum velocity in the presence of the inhibitor. A Lineweaver-Burk plot comparing an enzyme with and without a competitive inhibitor is shown in Figure 2.19.
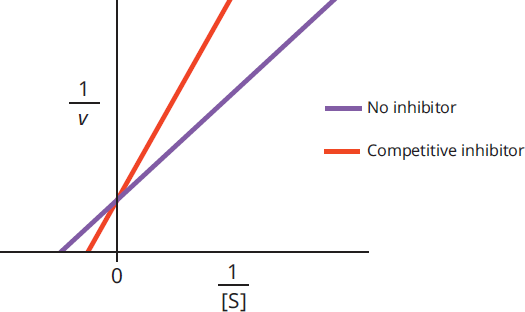
Figure 2.19 Lineweaver–Burk Plot of Competitive Inhibition
REAL WORLD
The concept of competitive inhibition has relevance in the clinical setting. For example, methanol (wood alcohol), if ingested, is enzymatically converted to toxic metabolites, which can cause blindness and even death. Administration of intravenous ethanol is the treatment of choice for a patient suffering from methanol poisoning. Ethanol works by competing with methanol for the active sites of the enzymes involved.
Noncompetitive Inhibition
Noncompetitive inhibitors bind to an allosteric site instead of the active site, which induces a change in enzyme conformation. Allosteric sites are non-catalytic regions of the enzyme that bind regulators. Because the two molecules do not compete for the same site, inhibition is considered noncompetitive and cannot be overcome by adding more substrate. Noncompetitive inhibitors bind equally well to the enzyme and the enzyme–substrate complex, unlike mixed inhibitors. As shown in Figure 2.20, once the enzyme’s conformation is altered by the binding of the inhibitor, no amount of extra substrate will be conducive to forming an enzyme–substrate complex.
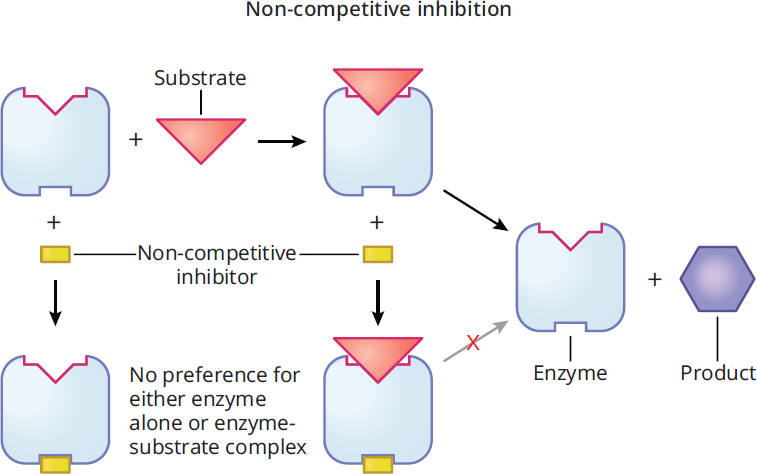
Figure 2.20 Noncompetitive Inhibition
Adding a noncompetitive inhibitor decreases the measured value of Vmax because there is less enzyme available to react; it does not, however, alter the value of Km because any copies of the enzyme that are still active maintain the same affinity for their substrate. A Lineweaver-Burk plot of an enzyme with and without a noncompetitive inhibitor is shown in Figure 2.21.
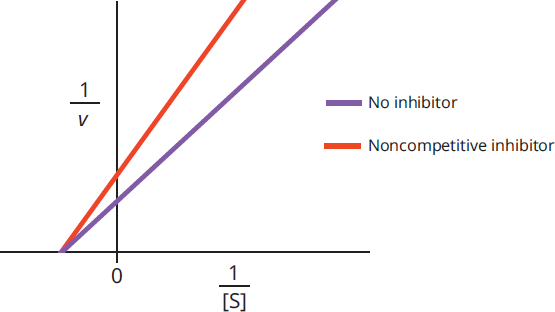
Figure 2.21 Lineweaver–Burk Plot of Noncompetitive Inhibition
Uncompetitive Inhibition
Uncompetitive inhibitors bind only to the enzyme-substrate complex and essentially lock the substrate in the enzyme, preventing its release and inhibiting substrate conversion into product, as shown in Figure 2.22. This can be interpreted as increasing affinity between the enzyme and substrate. Because the enzyme-substrate complex has already formed upon binding, uncompetitive inhibitors must bind at an allosteric site; in fact, it is the formation of the enzyme-substrate complex that creates a conformational change that allows the uncompetitive inhibitor to bind.
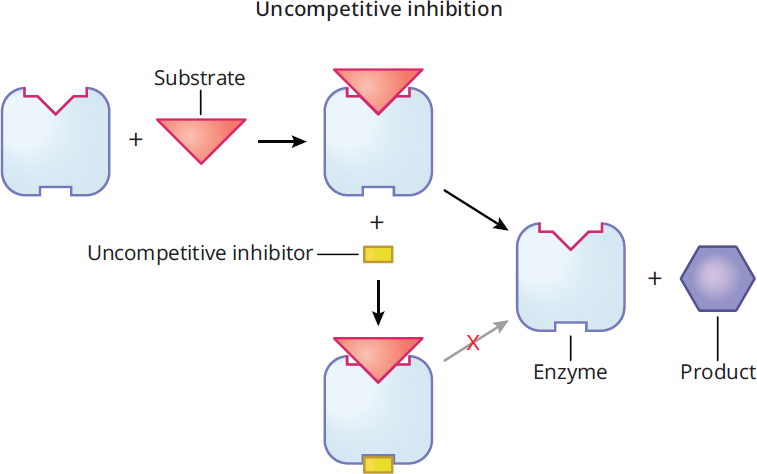
Figure 2.22 Uncompetitive Inhibition
Thus, uncompetitive inhibitors lower Km and Vmax. On a Lineweaver-Burk plot, as shown in Figure 2.23, the curves for activity with and without an uncompetitive inhibitor are parallel.
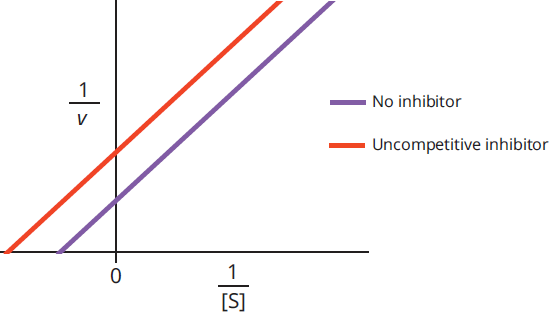
Figure 2.23 Lineweaver-Burk Plot of Uncompetitive Inhibition
Mixed Inhibition
Mixed inhibition results when an inhibitor can bind to either the enzyme or the enzyme-substrate complex, but has different affinity for each. If the inhibitor had the same affinity for both, it would be a noncompetitive inhibitor. As shown in Figure 2.24, mixed inhibitors do not bind at the active site, but at an allosteric site.
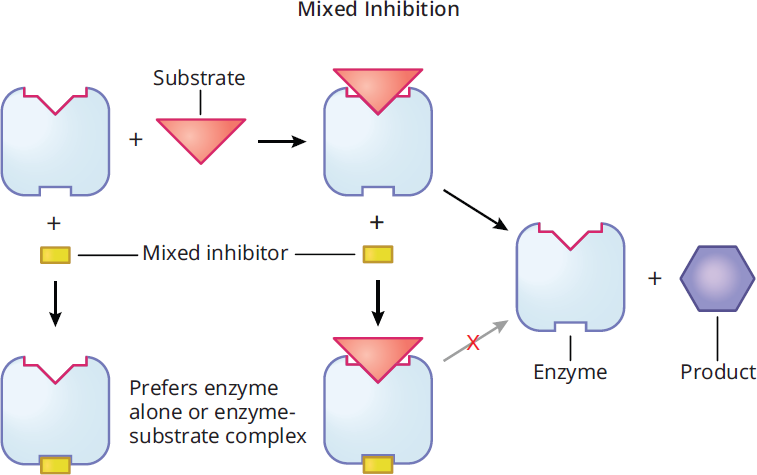
Figure 2.24 Mixed Inhibition
Mixed inhibition alters the experimental value of Km depending on the preference of the inhibitor for the enzyme vs. the enzyme-substrate complex. If the inhibitor preferentially binds to the enzyme, it increases the Km value (lowers affinity); if the inhibitor binds to the enzyme-substrate complex, it lowers the Km value (increases affinity). In either case, Vmax is decreased. On a Lineweaver-Burk plot, as shown in Figure 2.25, the curves for the activity with and without the inhibitor intersect at a point that is not on either axis.
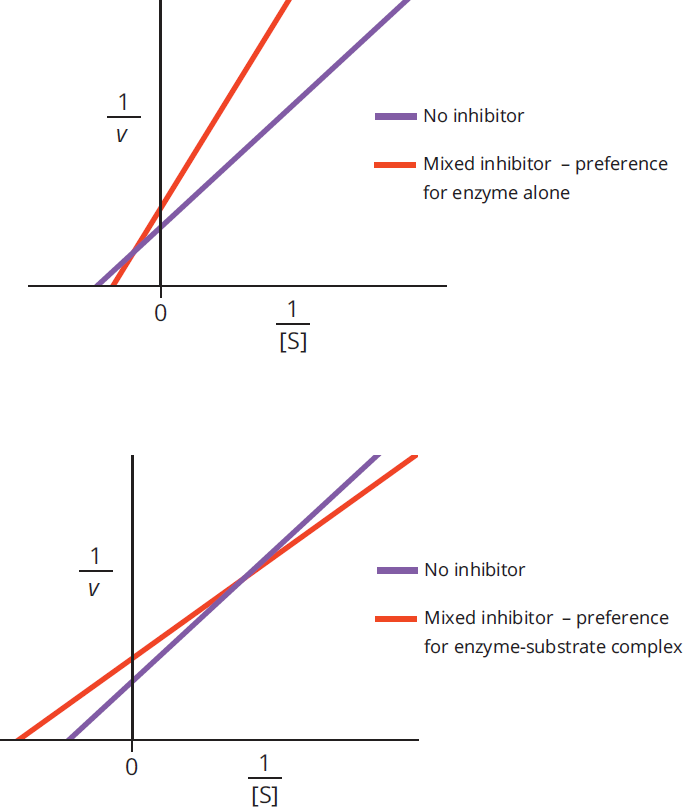
Figure 2.25 Lineweaver-Burk Plots of Mixed Inhibition
Table 2.2 Comparison of Reversible Inhibitors
COMPETITIVE NONCOMPETITIVE UNCOMPETITIVE MIXED
Binding Site Active site Allosteric site Allosteric site Allosteric site
**Impact on Km** Increases Unchanged Decreases Increases or Decreases
**Impact on Vmax** Unchanged Decreases Decreases Decreases
Irreversible Inhibition
In this type of inhibition, the active site is made unavailable for a prolonged period of time, or the enzyme is permanently altered. In other words, this type of inhibition is not easily overcome or reversed. A real-world example is aspirin. Acetylsalicylic acid (aspirin) irreversibly modifies cyclooxygenase-1. The enzyme can no longer bind its substrate (arachidonic acid) to make its products (prostaglandins), which are involved in modulating pain and inflammatory responses. To make more prostaglandins, new cyclooxygenase-1 will have to be synthesized through transcription and translation. Irreversible inhibition is a prime drug mechanism, and something that you will come across often in medical school.
Regulated Enzymes
Allosteric Enzymes
Enzymes that are allosteric have multiple binding sites. The active site is present, as well as at least one other site that can regulate the availability of the active site. These are known as allosteric sites. Allosteric enzymes alternate between an active and an inactive form. The inactive form cannot carry out the enzymatic reaction. Molecules that bind to the allosteric site may be either allosteric activators or allosteric inhibitors. Binding of either causes a conformational shift in the protein; however, the effect differs. An activator will result in a shift that makes the active site more available for binding to the substrate, whereas an inhibitor will make it less available. In addition to being able to alter the conformation of the protein, binding of activators or inhibitors may alter the activity of the enzyme. As shown in Figure 2.13 above, Michaelis–Menten plots of cooperative allosteric enzyme kinetics often have a sigmoidal (S-shaped) curve.
Covalently Modified Enzymes
In addition to transient interactions, enzymes are often subject to covalent modification. For example, enzymes can be activated or deactivated by phosphorylation or dephosphorylation. One cannot predict whether phosphorylation or dephosphorylation will activate an enzyme without experimental determination. Glycosylation, the covalent attachment of sugar moieties, is another covalent enzyme modification. Glycosylation can tag an enzyme for transport within the cell, or can modify protein activity and selectivity. Figure 2.26 shows a protein that has been phosphorylated and glycosylated. There are many other types of covalently-linked modifications, some of which are further detailed in Chapter 7 of MCAT Biochemistry Review, but specific mechanisms for the modifications of enzymes are still being studied and are beyond the scope of the MCAT.
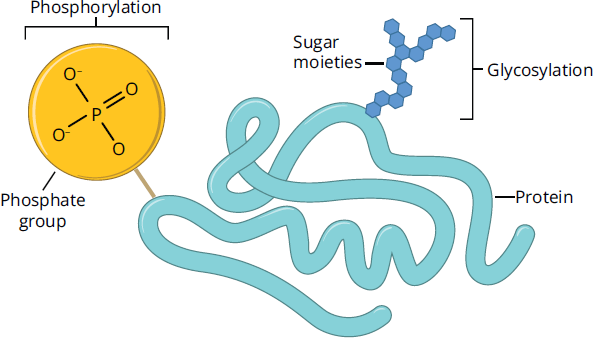
Figure 2.26 Examples of Covalent Protein Modifications
Zymogens
Certain enzymes are particularly dangerous if they are not tightly controlled. These include the digestive enzymes like trypsin, which, if released from the pancreas in an uncontrolled manner, would digest the organ itself. To avoid this danger, these enzymes and many others are secreted as inactive zymogens like trypsinogen. Zymogens contain a catalytic (active) domain and regulatory domain. The regulatory domain must be either removed or altered to expose the active site (see Figure 2.27 for an example). Apoptotic enzymes (caspases) exhibit similar regulation. Most zymogens have the suffix -ogen.
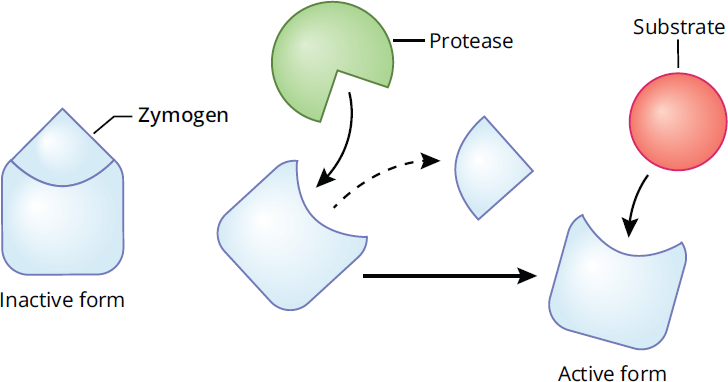
Figure 2.27 Zymogens Are Inactive Enzymes
BRIDGE
Consider that digestive enzymes, discussed in Chapter 9 of MCAT Biology Review, chew up fats, proteins, and carbohydrates—the very same compounds of which our body is made. How do these enzymes know to digest your food but not your body? Simply put, they don’t! So, we regulate their activity in a coordinated manner using feedback mechanisms and other substances.
MCAT CONCEPT CHECK 2.5:
Before you move on, assess your understanding of the material with these questions.
- What is feedback inhibition?
________________________________________
- Of the four types of reversible inhibitors, which could potentially increase Km?
________________________________________
- What is irreversible inhibition?
________________________________________
- What are some examples of transient and covalent enzyme modifications?
- Transient:
________________________________________
- Covalent:
________________________________________
- Why are some enzymes released as zymogens?
________________________________________
Conclusion
Our current chapter focused on the way in which cells are able to carry out the reactions necessary for life. We began with a discussion of the types of enzymes that you are likely to encounter on Test Day before reviewing thermodynamics and kinetics in relation to enzymes, which are biological catalysts. We went on to discuss the analysis of kinetic data with two different types of graphs, and talked about cooperativity. Because catalysts are generally most active in their native environment, we considered the impact of temperature, pH, and salinity on their activity. All of these are likely to appear on Test Day.
Enzymes need to be regulated; we analyzed the basics of feedback mechanisms. We talked about inhibitors of enzymes, which may be reversible or irreversible. The difference between the types of reversible inhibition is a key Test Day concept. Finally, we discussed changes in enzyme activity that may include allosteric activation, covalent modification, or cleavage of inactive zymogens. Let’s move on now to discuss the nonenzymatic functions of proteins. You will notice many parallels between the new material and the concepts described in this chapter, like binding affinity. By the end of the next chapter, you’ll be ready to face any protein question the MCAT can throw at you!
GO ONLINE!
You've reviewed the content, now test your knowledge and critical thinking skills by completing a test-like passage set in your online resources!
CONCEPT SUMMARY
Enzymes as Biological Catalysts
- Enzymes are biological catalysts that are unchanged by the reactions they catalyze and are reusable.
- Each enzyme catalyzes a single reaction or type of reaction with high specificity.
- Oxidoreductases catalyze oxidation–reduction reactions that involve the transfer of electrons.
- Transferases move a functional group from one molecule to another molecule.
- Hydrolases catalyze cleavage with the addition of water.
- Lyases catalyze cleavage without the addition of water and without the transfer of electrons. The reverse reaction (synthesis) is often more important biologically.
- Isomerases catalyze the interconversion of isomers, including both constitutional isomers and stereoisomers.
- Ligases are responsible for joining two large biomolecules, often of the same type.
- Translocases are responsible for transporting molecules across membranes.
- Exergonic reactions release energy; ΔG is negative.
- Enzymes lower the activation energy necessary for biological reactions.
- Enzymes do not alter the free energy (ΔG) or enthalpy (ΔH) change that accompanies the reaction nor the final equilibrium position; rather, they change the rate (kinetics) at which equilibrium is reached.
Mechanisms of Enzyme Activity
- Enzymes act by stabilizing the transition state, providing a favorable microenvironment, or bonding with the substrate molecules.
- Enzymes have an active site, which is the site of catalysis.
- Binding to the active site is explained by the lock and key theory or the induced fit model.
- The lock and key theory hypothesizes that the enzyme and substrate are exactly complementary.
- The induced fit model hypothesizes that the enzyme and substrate undergo conformational changes to interact fully.
- Some enzymes require metal cation cofactors or small organic coenzymes to be active.
Enzyme Kinetics
- Enzymes experience saturation kinetics: as substrate concentration increases, the reaction rate does as well until a maximum value is reached.
- Michaelis–Menten and Lineweaver–Burk plots represent this relationship as a hyperbola and line, respectively.
- Enzymes can be compared on the basis of their Km and Vmax values.
- Cooperative enzymes display a sigmoidal curve because of the change in activity with substrate binding.
Effects of Local Conditions on Enzyme Activity
- Temperature and pH affect an enzyme’s activity in vivo; changes in temperature and pH can result in denaturing of the enzyme and loss of activity due to loss of secondary, tertiary, or, if present, quaternary structure.
- In vitro, salinity can impact the action of enzymes.
Regulation of Enzyme Activity
- Enzyme pathways are highly regulated and subject to inhibition and activation.
- Feedback inhibition is a regulatory mechanism whereby the catalytic activity of an enzyme is inhibited by the presence of high levels of a product later in the same pathway.
- Reversible inhibition is characterized by the ability to replace the inhibitor with a compound of greater affinity or to remove it using mild laboratory treatment.
- Competitive inhibition results when the inhibitor is similar to the substrate and binds at the active site. Competitive inhibition can be overcome by adding more substrate. Vmax is unchanged, Km increases.
- Noncompetitive inhibition results when the inhibitor binds with equal affinity to the enzyme and the enzyme–substrate complex. Vmax is decreased, Km is unchanged.
- Uncompetitive inhibition results when the inhibitor binds only with the enzyme–substrate complex. Km and Vmax both decrease.
- Mixed inhibition results when the inhibitor binds with unequal affinity to the enzyme and the enzyme–substrate complex. Vmax is decreased, Km is increased or decreased depending on if the inhibitor has higher affinity for the enzyme or enzyme–substrate complex.
- Irreversible inhibition alters the enzyme in such a way that the active site is unavailable for a prolonged duration or permanently; new enzyme molecules must be synthesized for the reaction to occur again.
- Regulatory enzymes can experience activation as well as inhibition.
- Allosteric sites can be occupied by activators, which increase either affinity or enzymatic turnover.
- Phosphorylation (covalent modification with phosphate) or glycosylation (covalent modification with carbohydrate) can alter the activity or selectivity of enzymes.
- Zymogens are secreted in an inactive form and are activated by cleavage.
ANSWERS TO CONCEPT CHECKS
2.1
- Catalysts are characterized by two main properties: they reduce the activation energy of a reaction, thus speeding up the reaction, and they are not used up in the course of the reaction. Enzymes improve the environment in which a particular reaction takes place, which lowers its activation energy. They are also regenerated at the end of the reaction to their original form.
- Enzyme specificity refers to the idea that a given enzyme will only catalyze a given reaction or type of reaction. For example, serine/threonine-specific protein kinases will only place a phosphate group onto the hydroxyl group of a serine or threonine residue.
-
Name Function Ligase Addition or synthesis reactions, generally between large molecules, often require ATP
Isomerase Rearrangement of bonds within a compound
Lyase Cleavage of a single molecule into two products, or synthesis of small organic molecules
Hydrolase Breaking of a compound into two molecules using the addition of water
Oxidoreductase Oxidation–reduction reactions (transferring electrons)
Transferase Movement of a functional group from one molecule to another
Translocase Transport of molecules across membranes
- Enzymes have no effect on the overall thermodynamics of the reaction; they have no effect on the ΔG or ΔH of the reaction, although they do lower the energy of the transition state, thus lowering the activation energy. However, enzymes have a profound effect on the kinetics of a reaction. By lowering activation energy, equilibrium can be achieved faster (although the equilibrium position does not change).
**2.2**
-
Lock and Key Induced Fit
- Active site of enzyme fits exactly around substrate
- No alterations to tertiary or quaternary structure of enzyme
- Less accurate model
- Active site of enzyme molds itself around substrate only when substrate is present
- Tertiary and quaternary structure is modified for enzyme to function
- More accurate model
- Cofactors and coenzymes both act as activators of enzymes. Cofactors tend to be inorganic (minerals), while coenzymes tend to be small organic compounds (vitamins). In both cases, these regulators induce a conformational change in the enzyme that promotes its activity. Tightly bound cofactors or coenzymes that are necessary for enzyme function are termed prosthetic groups.
**2.3**
Increasing [E] will always increase Vmax, regardless of the starting concentration of enzyme.
- Increasing [S] has different effects, depending on how much substrate is present to begin with. When the substrate concentration is low, an increase in [S] causes a proportional increase in enzyme activity. At high [S], however, when the enzyme is saturated, increasing [S] has no effect on activity because Vmax has already been attained.
- Both the Michaelis–Menten and Lineweaver–Burk relationships account for the values of Km and Vmax under various conditions. They both provide simple graphical interpretations of these two variables and are derived from the Michaelis–Menten equation. However, the axes of these graphs and visual representation of this information is different between the two. The Michaelis–Menten plot is v vs. [S], which creates a hyperbolic curve for monomeric enzymes. The Lineweaver–Burk plot, on the other hand, is
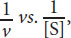
which creates a straight line.
- Km is a measure of an enzyme’s affinity for its substrate, and is defined as the substrate concentration at which an enzyme is functioning at half of its maximal velocity. As Km increases, an enzyme’s affinity for its substrate decreases.
- The x-intercept represents
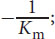
the y-intercept represents
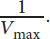
- Cooperativity refers to the interactions between subunits in a multisubunit enzyme or protein. The binding of substrate to one subunit induces a change in the other subunits from the T (tense) state to the R (relaxed) state, which encourages binding of substrate to the other subunits. In the reverse direction, the unbinding of substrate from one subunit induces a change from R to T in the remaining subunits, promoting unbinding of substrate from the remaining subunits.
**2.4**
- As temperature increases, enzyme activity generally increases (doubling approximately every 10 °C). Above body temperature, however, enzyme activity quickly drops off as the enzyme denatures. Enzymes are maximally active within a small pH range; outside of this range, activity drops quickly with changes in pH as the ionization of the active site changes and the protein is denatured. Changes in salinity can disrupt bonds within an enzyme, causing disruption of tertiary and quaternary structure, which leads to loss of enzyme function.
- Ideal temperature: 37 °C = 98.6°F = 310 K
Ideal pH for most enzymes is 7.4; for gastric enzymes, around 2; for pancreatic enzymes, around 8.5.
**2.5**
- Feedback inhibition refers to the product of an enzymatic pathway turning off enzymes further back in that same pathway. This helps maintain homeostasis: as product levels rise, the pathway creating that product is appropriately downregulated.
- A competitive inhibitor increases Km because the substrate concentration has to be higher to reach half the maximum velocity in the presence of the inhibitor. A mixed inhibitor will increase Km only if the inhibitor preferentially binds to the enzyme over the enzyme–substrate complex.
- Irreversible inhibition refers to the prolonged or permanent inactivation of an enzyme, such that it cannot be easily renatured to gain function.
- Examples of transient modifications include allosteric activation or inhibition. Examples of covalent modifications include phosphorylation and glycosylation.
- Zymogens are precursors of active enzymes. It is critical that certain enzymes (like the digestive enzymes of the pancreas) remain inactive until arriving at their target site.
SCIENCE MASTERY ASSESSMENT EXPLANATIONS
1. D
Enzymes catalyze reactions by lowering their activation energy, and are not changed or consumed during the course of the reaction. While the activation energy is lowered, the free energy of the reaction, ΔG, remains unchanged in the presence of an enzyme. A reaction will continue to occur in the presence or absence of an enzyme; it simply runs slower without the enzyme, eliminating (A). Most physiological reactions are optimized at body temperature, 37 °C, eliminating (B). Finally, dehydrogenases catalyze oxidation–reduction reactions, not transfer reactions, eliminating (C).
2. A
Most enzymes in the human body operate at maximal activity around a temperature of 37 °C and a pH of 7.4, which is the pH of most body fluids. In addition, as characterized by the Michaelis–Menten equation, enzymes form an enzyme–substrate complex, which can either dissociate back into the enzyme and substrate or proceed to form a product. So far, we can eliminate (B), (C), and (D), so let’s check (A). An increase in the substrate concentration, while maintaining a constant enzyme concentration, leads to a proportional increase in the rate of the reaction only initially. However, once most of the active sites are occupied, the reaction rate levels off, regardless of further increases in substrate concentration. At high concentrations of substrate, the reaction rate approaches its maximal velocity and is no longer changed by further increases in substrate concentration.
3. B
An enzyme devoid of its necessary cofactor is called an apoenzyme and is catalytically inactive.
4. A
An enzyme’s specificity is determined by the three-dimensional shape of its active site. Regardless of which explanation for enzyme specificity we are discussing (lock and key or induced fit), the active site determines which substrate the enzyme will react with.
5. A
As the temperature of the DNA polymerase sample increases from 0 °C to the usual physiological temperature, i.e. 37 °C, the enzyme's activity will increase. However, at temperatures above 37 °C, the enzyme's activity will rapidly decline due to denaturation.
6. D
By limiting the activity of enzyme 1, the rest of the pathway is slowed, which is the definition of negative feedback. (A) is incorrect because there is no competition for the active site with allosteric interactions. While many products do indeed competitively inhibit an enzyme in the pathway that creates them, this is an example of an allosterically inhibited enzyme. There is not enough information for (B) to be correct because we aren’t told whether the inhibition is reversible. In general, allosteric interactions are temporary. (C) is incorrect because it is the opposite of what occurs when enzyme 1 activity is reduced.
7. A
While the equations given in the text are useful, recognizing relationships is even more important. You can see that as substrate concentration increases significantly, there is only a small change in the rate. This occurs as we approach Vmax. Because the Vmax is near
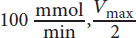
equals
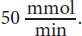
The substrate concentration giving this rate is 0.5 mM and corresponds to Km; therefore, (A) is correct.
8. A
As with the last question, relationships are important. At a concentration of 5 × 10–6 M, enzyme A is working at one-half of its Vmax because the concentration is equal to the Km of the enzyme. Therefore, one-half of
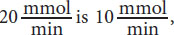
which corresponds to (A).
9. C
At a concentration of 5 × 10–4 M, there is 100 times more substrate than present at half maximal velocity. At high values (significantly larger than the value of Km), the enzyme is at or near its Vmax, which is 20
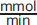
.
10. B
Based on the graph, when the substrate is present, oseltamivir results in the same Vmax and a higher Km compared to when no inhibitor is added. These are hallmarks of competitive inhibitors. Noncompetitive inhibitors result in decreased Vmax and the same Km as the uninhibited reaction, which is shown by the zanamivir line in the graph. Because the question is only comparing the values between the two inhibitors, and not the enzyme without inhibitor, the mechanism of inhibition is less important to determine than the values of Km and Vmax. This is a great example of why previewing the answer choices works well in the sciences.
11. C
Lyases are responsible for the breakdown of a single molecule into two molecules without the addition of water or the transfer of electrons. Lyases often form cyclic compounds or double bonds in the products to accommodate this. Water was not a reactant, and no cofactor was mentioned; thus lyase, (C), remains the best answer choice.
12. D
Enzymes are not altered by the process of catalysis. A molecule that breaks intramolecular bonds to provide activation energy would not be able to be reused.
13. D
Cooperative enzymes demonstrate a change in affinity for the substrate depending on how many substrate molecules are bound and whether the last change was accomplished because a substrate molecule was bound or left the active site of the enzyme. Because we cannot determine whether the most recent reaction was binding or dissociation, (A) and (B) are eliminated. We can make absolute comparisons though. For enzymes expressing positive cooperativity, the unbound enzyme has the lowest affinity for substrate, and the enzyme with all but one subunit bound has the highest. The increase in affinity is not necessarily linear. Furthermore, if all four sites have substrate bound, the enzyme cannot bind to any more substrate. Therefore, (C) is not true. An enzyme with two subunits occupied must have a higher affinity for the substrate than the same enzyme with only one subunit occupied; thus, (D) is correct.
14. A
Triglycerides are unlikely to act as coenzymes for a few reasons, including their large size, neutral charge, and ubiquity in cells. Cofactors and coenzymes tend to be small in size, such as metal ions like (C) or small organic molecules. They can usually carry a charge by ionization, protonation, or deprotonation. Finally, they are usually in low, tightly regulated concentrations within cells. Metabolic pathways would be expected to include both oxidation–reduction reactions and movement of functional groups, thus eliminating (B) and (D).
15. B
The rate of reaction increases with temperature because of the increased kinetic energy of the reactants, but reaches a peak temperature because the enzyme denatures with the disruption of hydrogen bonds at excessively high temperatures. In the absence of enzyme, this peak temperature is generally much hotter. Heating a reaction provides molecules with an increased chance of achieving the activation energy, but the enzyme catalyst would typically reduce activation energy. Keep in mind that thermodynamics and kinetics are not interchangeable, so we are not considering the impact of heat on the equilibrium position.
GO ONLINE
Consult your online resources for additional practice.
EQUATIONS TO REMEMBER
(2.1) Michaelis–Menten rates:
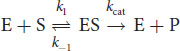
(2.2) Michaelis–Menten equation:
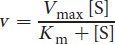
(2.3) **Turnover number (kcat):**
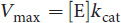
SHARED CONCEPTS
- Biochemistry Chapter 1
- Amino Acids, Peptides, and Proteins
- Biochemistry Chapter 12
- Bioenergetics and Regulation of Metabolism
- Biology Chapter 5
- The Endocrine System
- Biology Chapter 9
- The Digestive System
- General Chemistry Chapter 5
- Chemical Kinetics
- General Chemistry Chapter 7
- Thermochemistry
- General Chemistry Chapter 11
- Oxidation–Reduction Reactions