Chapter 3: Nonenzymatic Protein Function and Protein Analysis

Chapter 3: Nonenzymatic Protein Function and Protein Analysis
SCIENCE MASTERY ASSESSMENT
Every pre-med knows this feeling: there is so much content I have to know for the MCAT! How do I know what to do first or what's important?
While the high-yield badges throughout this book will help you identify the most important topics, this Science Mastery Assessment is another tool in your MCAT prep arsenal. This quiz (which can also be taken in your online resources) and the guidance below will help ensure that you are spending the appropriate amount of time on this chapter based on your personal strengths and weaknesses. Don't worry though— skipping something now does not mean you'll never study it. Later on in your prep, as you complete full-length tests, you'll uncover specific pieces of content that you need to review and can come back to these chapters as appropriate.
How to Use This Assessment
If you answer 0–7 questions correctly:
Spend about 1 hour to read this chapter in full and take limited notes throughout. Follow up by reviewing all quiz questions to ensure that you now understand how to solve each one.
If you answer 8–11 questions correctly:
Spend 20–40 minutes reviewing the quiz questions. Beginning with the questions you missed, read and take notes on the corresponding subchapters. For questions you answered correctly, ensure your thinking matches that of the explanation and you understand why each choice was correct or incorrect.
If you answer 12–15 questions correctly:
Spend less than 20 minutes reviewing all questions from the quiz. If you missed any, then include a quick read-through of the corresponding subchapters, or even just the relevant content within a subchapter, as part of your question review. For questions you got correct, ensure your thinking matches that of the explanation and review the Concept Summary at the end of the chapter.
- At what pH can protein A best be obtained through electrophoresis? (Note: MM = molar mass)
Protein pI MM Protein A 4.5 25,000
Protein B 6.0 10,000
Protein C 9.5 12,000
- 2.5
- 3.5
- 4.5
- 5.5
- What is the function of sodium dodecyl sulfate (SDS) in SDS-PAGE?
- SDS stabilizes the gel matrix, improving resolution during electrophoresis.
- SDS solubilizes proteins to give them uniformly negative charges, so the separation is based purely on size.
- SDS raises the pH of the gel, separating multiunit proteins into individual subunits.
- SDS solubilizes proteins to give them uniformly positive charges, so separation is based purely on pH.
- Which of the following is NOT involved in cell migration?
- Dynein
- Flagella
- Actin
- Centrioles
- Which of the following proteins is most likely to be found extracellularly?
- Tubulin
- Myosin
- Collagen
- Actin
- Hormones are found in the body in very low concentrations, but tend to have a strong effect. What type of receptor are hormones most likely to act on?
- Ligand-gated ion channels
- Enzyme-linked receptors
- G protein-coupled receptors
- I only
- III only
- II and III only
- I, II, and III
- Which of the following is most likely to be found bound to a protein in the body?
- Sodium
- Potassium
- Chloride
- Calcium
- Which of the following characteristics is NOT attributed to antibodies?
- Antibodies bind to more than one distinct antigen.
- Antibodies label antigens for targeting by other immune cells.
- Antibodies can cause agglutination by interaction with antigen.
- Antibodies have two heavy chains and two light chains.
- Which ion channels are responsible for maintaining the resting membrane potential?
- Ungated channels
- Voltage-gated channels
- Ligand-gated channels
- No ion channels are involved in maintenance of the resting membrane potential.
- Which of the following is NOT a component of all trimeric G proteins?
- Gα
- Gβ
- Gγ
- Gi
- Which of the following methods would be best to separate large quantities of the following proteins? (Note: MM = molar mass)
Protein pI MM Protein A 6.5 28,000
Protein B 6.3 70,000
Protein C 6.6 200,000
- Ion-exchange chromatography
- Size-exclusion chromatography
- Isoelectric focusing
- Native PAGE
- Which amino acids contribute most significantly to the pI of a protein?
- Lysine
- Glycine
- Arginine
- I only
- I and II only
- I and III only
- II and III only
- How does the gel for isoelectric focusing differ from the gel for traditional electrophoresis?
- Isoelectric focusing uses a gel with much larger pore sizes to allow for complete migration.
- Isoelectric focusing uses a gel with SDS added to encourage a uniform negative charge.
- Isoelectric focusing uses a gel with a pH gradient that encourages a variable charge.
- The gel is unchanged in isoelectric focusing; the protein mixture is treated before loading.
- Which protein properties allow UV spectroscopy to be used as a method of determining concentration?
- Proteins have partially planar characteristics in peptide bonds.
- Globular proteins cause scattering of light.
- Proteins contain aromatic groups in certain amino acids.
- All organic macromolecules can be assessed with UV spectroscopy.
- A protein collected through affinity chromatography displays no activity even though it is found to have a high concentration using the Bradford protein assay. What best explains these findings?
- The Bradford reagent was prepared incorrectly.
- The active site is occupied by free ligand.
- The protein is bound to the column.
- The protein does not catalyze the reaction of interest.
- What property of protein-digesting enzymes allows for a sequence to be determined without fully degrading the protein?
- Selectivity
- Sensitivity
- Turnover
- Inhibition
Answer Key
- D
- B
- D
- C
- C
- D
- A
- A
- D
- B
- C
- C
- C
- B
- A
Chapter 3: Nonenzymatic Protein Function and Protein Analysis
CHAPTER 3
NONENZYMATIC PROTEIN FUNCTION AND PROTEIN ANALYSIS
In This Chapter
3.1 Cellular Functions
Structural Proteins
Motor Proteins
Binding Proteins
Cell Adhesion Molecules
Immunoglobulins
3.2 Biosignaling
Ion Channels
Enzyme-Linked Receptors
G Protein-Coupled Receptors
**3.3 Protein Isolation
**
Electrophoresis
Chromatography
3.4 Protein Analysis
Protein Structure
Amino Acid Composition
Activity Analysis
Concentration Determination
Concept Summary
CHAPTER PROFILE
The content in this chapter should be relevant to about 4% of all questions about biochemistry on the MCAT.
This chapter covers material from the following AAMC content categories:
1A: Structure and function of proteins and their constituent amino acids
3A: Structure and functions of the nervous and endocrine systems, and ways in which these systems coordinate the organ systems
5C: Separation and purification methods
5D: Structure, function, and reactivity of biologically relevant molecules
Introduction
Did you ever wonder why some athletes eat raw eggs when training? It doesn’t help with digestion or increase enzymatic activity throughout the body; rather, all that protein is used for muscle building. While muscle building does increase cytoplasm and all of the enzymes found therein, the biggest increase is seen in structural and motor proteins like actin and myosin. Valid concerns about bacteria in and on eggs have stopped most athletes from drinking raw eggs, but protein shakes and massive containers of whey protein have replaced them on the shelves of health-food stores, pharmacies, and supermarkets.
In this chapter, we will examine some of the structural and motor proteins that athletes love to build, as well as more complex protein functions in humans, including biosignaling. Finally, we’ll learn how to determine which proteins—and their concentrations—are in those eggs through separation and quantitative analysis.
3.1 Cellular Functions
LEARNING OBJECTIVES
After Chapter 3.1, you will be able to:
- Compare and contrast cytoskeletal proteins and motor proteins
- Associate collagen, elastin, keratins, actin, and tubulin with their major functions
- Describe the adhesive properties of cadherins, integrins, and selectins
- Predict the possible outcomes of an antibody binding to its antigen
Typical functions provided by proteins within the cell include supporting cellular shape and organization and acting as enzymes. The enzymatic functions of proteins were detailed in Chapter 2 of MCAT Biochemistry Review and we are now ready to look at other protein functions. Structural and motor proteins are found in abundance within individual cells and are also found in the extracellular matrix.
Structural Proteins
The cytoskeleton can be thought of as a three-dimensional web or scaffolding system for the cell. It is comprised of proteins that are anchored to the cell membrane by embedded protein complexes. In addition to intracellular support, extracellular matrices composed of proteins also support the tissues of the body. Tendons, ligaments, cartilage, and basement membranes are all proteinaceous. The primary structural proteins in the body are collagen, elastin, keratin, actin, and tubulin. Structural proteins generally have highly repetitive secondary structure and a supersecondary structure—a repetitive organization of secondary structural elements together sometimes referred to as a motif. This regularity gives many structural proteins a fibrous nature.
Collagen
Collagen has a characteristic trihelical fiber (three left-handed helices woven together to form a secondary right-handed helix) and makes up most of the extracellular matrix of connective tissue. It is found throughout the body and is important in providing strength and flexibility.
REAL WORLD
The importance of the structure of collagen is highlighted in the disorder osteogenesis imperfecta, also referred to as brittle bone disease. Collagen—a major component of bone—forms a unique and specific secondary helical structure based on the abundance of the amino acid glycine. The replacement of glycine with other amino acids can cause improper folding of the collagen protein and cell death, leading to bone fragility.
Elastin
Elastin is another important component of the extracellular matrix of connective tissue. Its primary role is to stretch and then recoil like a spring, which restores the original shape of the tissue.
Keratins
Keratins are intermediate filament proteins found in epithelial cells. Keratins contribute to the mechanical integrity of the cell and also function as regulatory proteins. Keratin is the primary protein that makes up hair and nails.
Actin
Actin is a protein that makes up microfilaments and the thin filaments in myofibrils. It is the most abundant protein in eukaryotic cells. Actin proteins have a positive side and a negative side; this polarity allows motor proteins to travel unidirectionally along an actin filament, like a one-way street.
Tubulin
Tubulin is the protein that makes up microtubules. Microtubules are important for providing structure, chromosome separation in mitosis and meiosis, and intracellular transport with kinesin and dynein, described in the next section. Like actin, tubulin has polarity: the negative end of a microtubule is usually located adjacent to the nucleus, whereas the positive end is usually in the periphery of a cell.
Motor Proteins
Some structural proteins also have motor functions in the presence of motor proteins. The motile cilia and flagella of bacteria and sperm are prime examples, as is the contraction of the sarcomere in muscle. Motor proteins also display enzymatic activity, acting as ATPases that power the conformational change necessary for motor function. Motor proteins have transient interactions with either actin or microtubules.
Myosin is the primary motor protein that interacts with actin. In addition to its role as the thick filament in a myofibril, myosin can be involved in cellular transport. Each myosin subunit has a single head and neck; movement at the neck is responsible for the power stroke of sarcomere contraction.
BRIDGE
Motor proteins are responsible for muscle contraction and cellular movement. Take a moment to review sarcomere structure in Chapter 11 of MCAT Biology Reviewas another example of the interaction between motor proteins and the cytoskeleton.
Kinesins and dyneins are the motor proteins associated with microtubules. They have two heads, at least one of which remains attached to tubulin at all times. Kinesins play key roles in aligning chromosomes during metaphase and depolymerizing microtubules during anaphase of mitosis. Dyneins are involved in the sliding movement of cilia and flagella. Both proteins are important for vesicle transport in the cell, but have opposite polarities: kinesins bring vesicles toward the positive end of the microtubule, and dyneins bring vesicles toward the negative end. In neurons, we see a classic example of these motor proteins’ polarities. Kinesins bring vesicles of neurotransmitter to the positive end of the axonal microtubules (toward the synaptic terminal). In contrast, dyneins bring vesicles of waste or recycled neurotransmitter back toward the negative end of the microtubule (toward the soma) through retrograde transport. The activity of kinesins is illustrated in Figure 3.1 below.
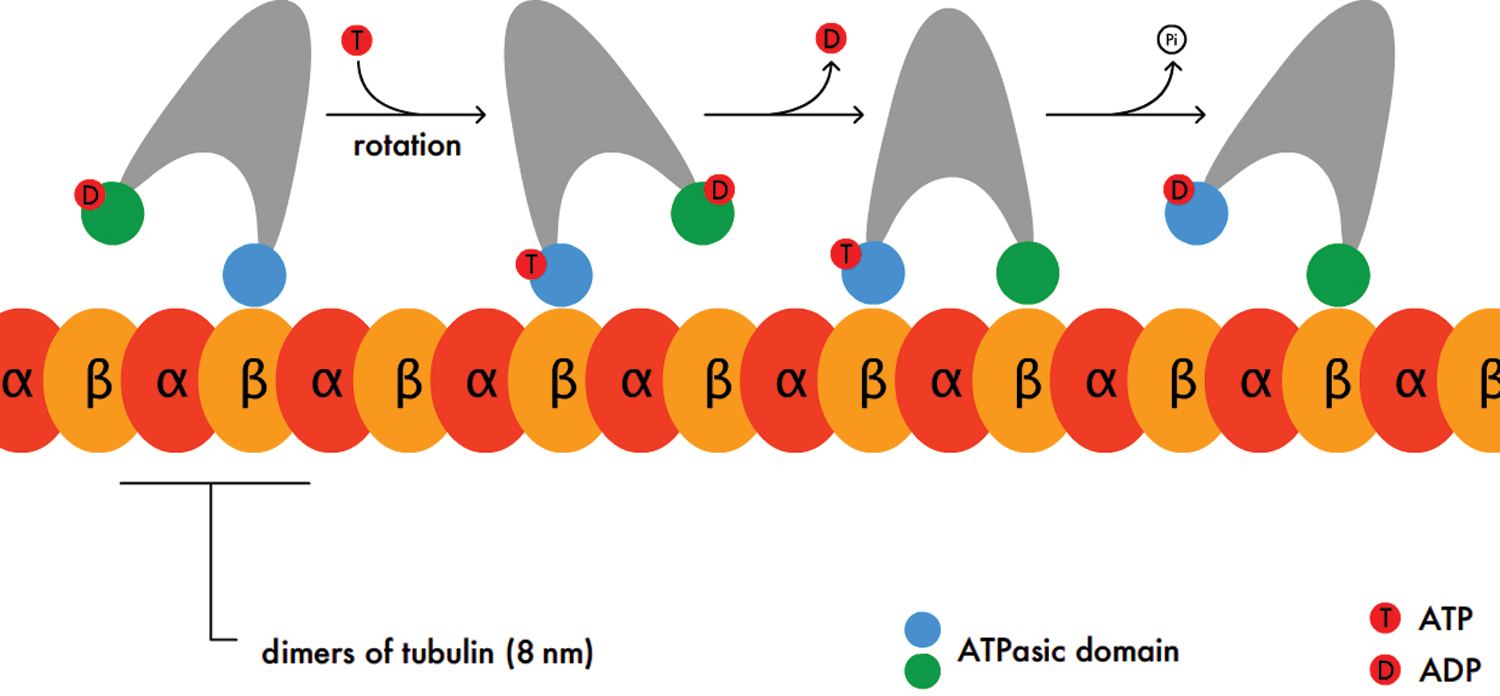
Figure 3.1 Stepwise Activity of Kinesins Kinesins move along microtubules in a stepping motion such that one or both heads remain attached at all times.
Binding Proteins
While proteins primarily exert enzymatic or structural functions within the cell, they also can have stabilizing functions in individual cells and the body. Proteins that act in this way transport or sequester molecules by binding to them. Binding proteins include hemoglobin, calcium-binding proteins, DNA-binding proteins (often transcription factors), and others. Each binding protein has an affinity curve for its molecule of interest; the oxyhemoglobin dissociation curve is one well-known example. This curve differs depending on the goal of the binding protein. When sequestration of a molecule is the goal, the binding protein usually has high affinity for its target across a large range of concentrations so it can keep it bound at nearly 100 percent. A transport protein, which must be able to bind or unbind its target to maintain steady-state concentrations, is likely to have varying affinity depending on the environmental conditions.
Cell Adhesion Molecules
Cell adhesion molecules (CAMs) are proteins found on the surface of most cells and aid in binding the cell to the extracellular matrix or other cells. While there are a number of different types of CAMs, they are all integral membrane proteins. Adhesion molecules can be classified into three major families: cadherins, integrins, and selectins.
Cadherins
Cadherins are a group of glycoproteins that mediate calcium-dependent cell adhesion. Cadherins often hold similar cell types together, such as epithelial cells. Different cells usually have type-specific cadherins; for example, epithelial cells use E-cadherin while nerve cells use N-cadherin.
Integrins
Integrins are a group of proteins that all have two membrane-spanning chains called α and β. These chains are very important in binding to and communicating with the extracellular matrix. Integrins also play a very important role in cellular signaling and can greatly impact cellular function by promoting cell division, apoptosis, or other processes. For example, integrin αIIbβ3 allows platelets to stick to fibrinogen, a clotting factor, which causes activation of platelets to stabilize the clot. Other integrins are used for white blood cell migration, stabilization of epithelium on its basement membrane, and other processes.
Selectins
Selectins are unique because they bind to carbohydrate molecules that project from other cell surfaces. These bonds are the weakest formed by the CAMs discussed here. Selectins are expressed on white blood cells and the endothelial cells that line blood vessels. Like integrins, they play an important role in host defense, including inflammation and white blood cell migration, as shown in Figure 3.2.
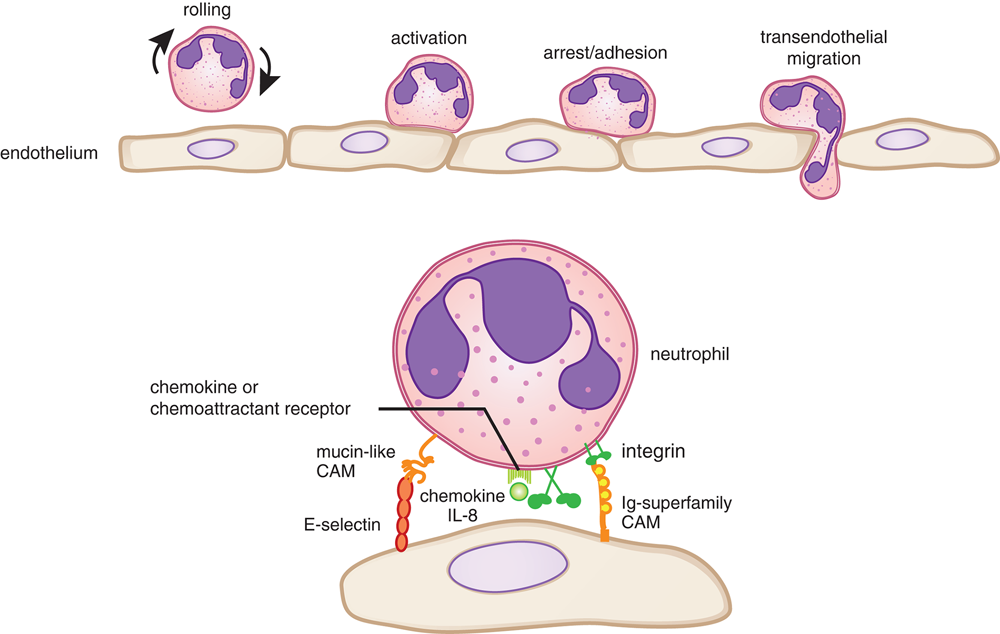
Figure 3.2 White Blood Cell Migration Using Selectins and Integrins Many other proteins are involved in white blood cell migration, but are outside the scope of the MCAT.
REAL WORLD
Many medications target selectins and integrins. For example, research has shown that the ability of cancer cells to metastasize (break away from a tumor and invade other distant tissues) is associated with unique expression patterns of CAMs. By targeting these CAMs, metastasis may be avoided. To stop the clotting process during heart attacks, other medications target CAMs used by platelets.
Immunoglobulins
The immune system is very complex and is made up of many different types of cells and proteins. These cells and proteins have a common purpose: to rid the body of foreign invaders. The most prominent type of protein found in the immune system is the antibody. Antibodies, also called immunoglobulins (Ig) are proteins produced by B-cells that function to neutralize targets in the body, such as toxins and bacteria, and then recruit other cells to help eliminate the threat. Antibodies are Y-shaped proteins that are made up of two identical heavy chains and two identical light chains, as shown in Figure 3.3. Disulfide linkages and noncovalent interactions hold the heavy and light chains together. Each antibody has an antigen-binding region at the tips of the “Y.” Within this region, there are specific polypeptide sequences that will bind one, and only one, specific antigenic sequence. The remaining part of the antibody molecule is known as the constant region, which is involved in recruitment and binding of other cells of the immune system, such as macrophages. Thus, when antibodies bind to their targets, called antigens, they can cause one of three outcomes:
- Neutralizing the antigen, making the pathogen or toxin unable to exert its effect on the body
- Marking the pathogen for destruction by other white blood cells immediately; this marking function is also called opsonization
- Clumping together (agglutinating) the antigen and antibody into large insoluble protein complexes that can be phagocytized and digested by macrophages
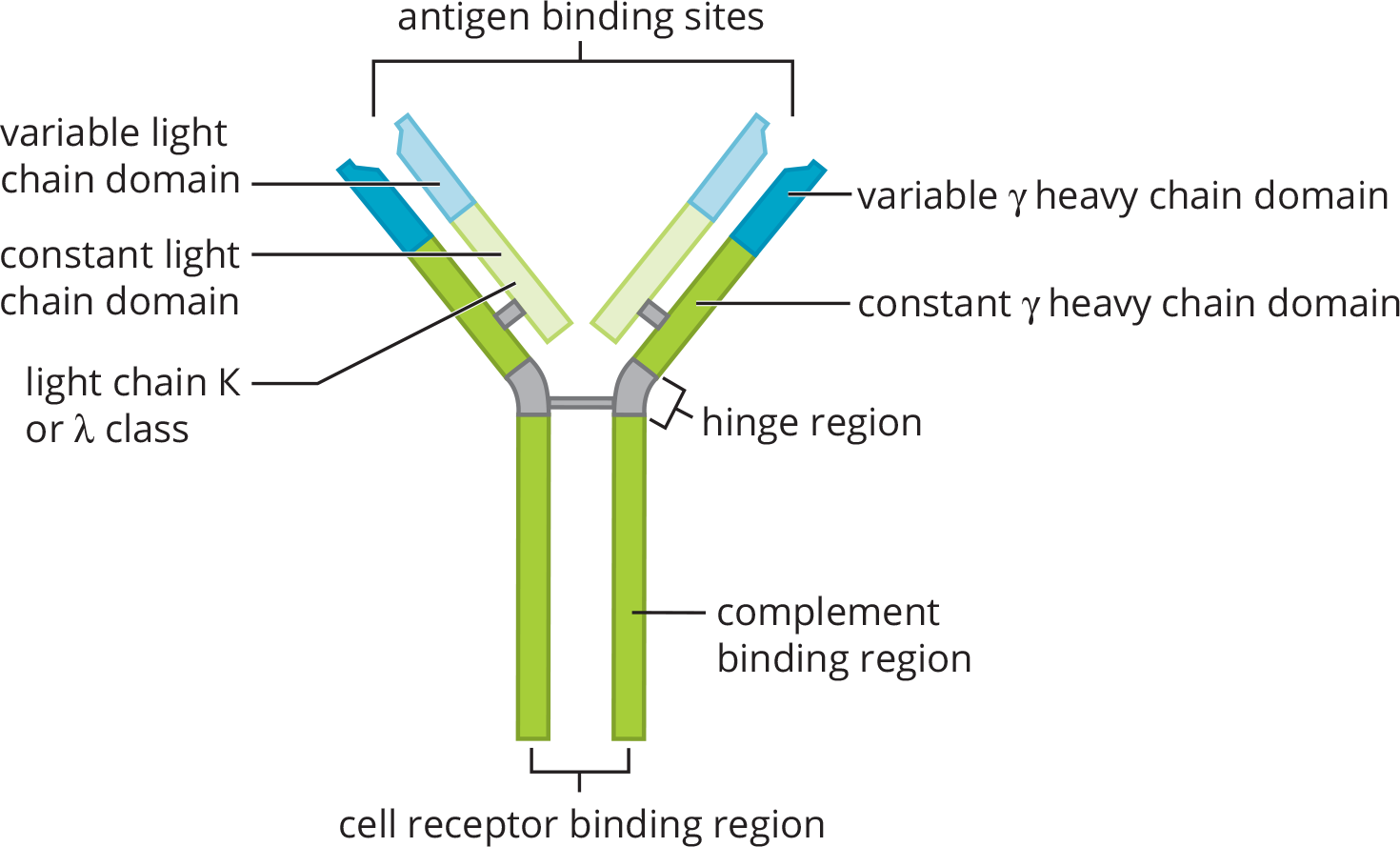
Figure 3.3 Structure of an Antibody Molecule
MCAT CONCEPT CHECK 3.1:
Before you move on, assess your understanding of the material with these questions.
- How do cytoskeletal proteins differ from motor proteins?
________________________________________
- True or False: Motor proteins are not enzymes.
- What could permit a binding protein involved in sequestration to have a low affinity for its substrate and still have a high percentage of substrate bound?
________________________________________
- What are the three main classes of cell adhesion molecules? What type of adhesion does each class form?
Cell Adhesion Molecule Type of Adhesion
- When an antibody binds to its antigen, what are the three possible outcomes of this interaction?
- _______________________________________
- _______________________________________
- _______________________________________
3.2 Biosignaling
LEARNING OBJECTIVES
After Chapter 3.2, you will be able to:
- Contrast enzyme-linked receptors with G protein-coupled receptors
- Distinguish between ungated channels, voltage-gated channels, and ligand-gated channels
- Recognize key features of transport kinetics and biosignaling processes
Biosignaling is a process in which cells receive and act on signals. Proteins participate in biosignaling in different capacities, including acting as extracellular ligands, transporters for facilitated diffusion, receptor proteins, and second messengers. The proteins involved in biosignaling can have functions in substrate binding or enzymatic activity.
Ion Channels
Ion channels are proteins that create specific pathways for charged molecules. They are classified into three main groups that have different mechanisms of opening, but that all permit facilitated diffusion of charged particles. Facilitated diffusion, a type of passive transport, is the diffusion of molecules down a concentration gradient through a pore in the membrane created by this transmembrane protein. It is used for molecules that are impermeable to the membrane (large, polar, or charged). Facilitated diffusion allows integral membrane proteins to serve as channels for these substrates to avoid the hydrophobic fatty acid tails of the phospholipid bilayer. The three main types of ion channels are ungated, voltage-gated, and ligand-gated.
REAL WORLD
Hundreds of ion channels have been identified that function in cell signaling and cell excitability. In addition, ion channels are drug targets in the treatment of everything from heart disease (calcium channel blockers) to seizures (sodium channel blockers) to irritable bowel syndrome (acetylcholine receptor/cation channel blockers).
Ungated Channels
As their name suggests, ungated channels have no gates and are therefore unregulated. For example, all cells possess ungated potassium channels. This means there will be a net efflux of potassium ions through these channels unless potassium is at equilibrium.
Voltage-Gated Channels
In voltage-gated channels, the gate is regulated by the membrane potential change near the channel. For example, many excitable cells such as neurons possess voltage-gated sodium channels. The channels are closed under resting conditions, but membrane depolarization causes a protein conformation change that allows them to quickly open and then quickly close as the voltage increases. Voltage-gated nonspecific sodium–potassium channels are found in cells of the sinoatrial node of the heart. Here, they serve as the pacemaker current; as the voltage drops, these channels open to bring the cell back to threshold and fire another action potential, as shown in Figure 3.4.
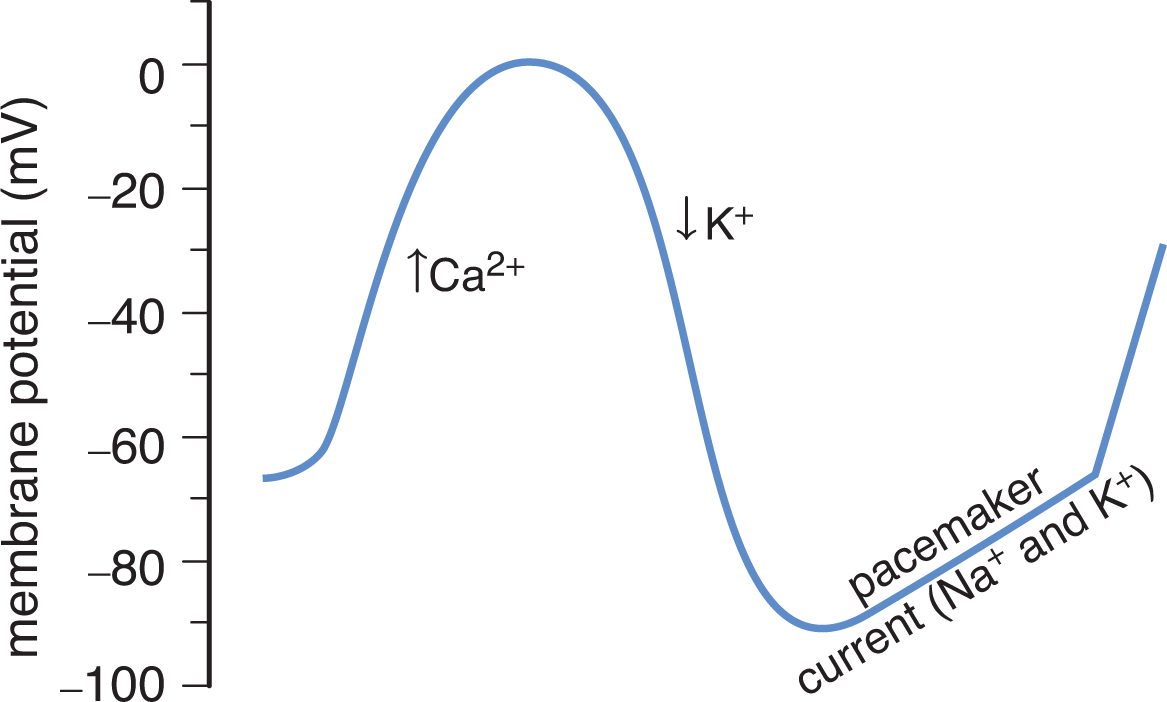
Figure 3.4 Action Potential of the Sinoatrial Node
Ligand-Gated Channels
For ligand-gated channels, the binding of a specific substance or ligand to the channel causes it to open or close. For example, neurotransmitters act at ligand-gated channels at the postsynaptic membrane: the inhibitory neurotransmitter γ-aminobutyric acid (GABA) binds to a chloride channel and opens it.
BRIDGE
The activity at the neuromuscular junction and most chemical synapses relies on ligand-gated ion channels. The nervous system especially makes use of this type of gating, as well as voltage-gated ion channels, as discussed in Chapter 4 of MCAT Biology Review.
The Km and Vmax parameters that apply to enzymes are also applicable to transporters such as ion channels in membranes. The kinetics of transport can be derived from the Michaelis–Menten and Lineweaver–Burk equations, where Km refers to the solute concentration at which the transporter is functioning at half of its maximum activity.
Enzyme-Linked Receptors
Membrane receptors may also display catalytic activity in response to ligand binding. These enzyme-linked receptors have three primary protein domains: a membrane-spanning domain, a ligand-binding domain, and a catalytic domain. The membrane-spanning domain anchors the receptor in the cell membrane. The ligand-binding domain is stimulated by the appropriate ligand and induces a conformational change that activates the catalytic domain. This often results in the initiation of a second messenger cascade. Receptor tyrosine kinases (RTK) are classic examples. RTKs are composed of a monomer that dimerizes upon ligand binding. The dimer is the active form that phosphorylates additional cellular enzymes, including the receptor itself (autophosphorylation). Other classes of enzyme-linked receptors include serine/threonine-specific protein kinases and receptor tyrosine phosphatases.
KEY CONCEPT
Biosignaling can take advantage of either existing gradients (ion channels) or second messenger cascades (enzyme-linked receptors and G protein-coupled receptors).
G Protein-Coupled Receptors
G protein-coupled receptors (GPCR) are a large family of integral membrane proteins involved in signal transduction. They are characterized by their seven membrane-spanning α-helices. The receptors differ in specificity of the ligand-binding area found on the extracellular surface of the cell. In order for GPCRs to transmit signals to an effector in the cell, they utilize a heterotrimeric G protein. G proteins are named for their intracellular link to guanine nucleotides (GDP and GTP). The binding of a ligand increases the affinity of the receptor for the G protein. The binding of the G protein represents a switch to the active state and affects the intracellular signaling pathway. There are several different G proteins that can result in either stimulation or inhibition of the signaling pathway. There are three main types of G proteins:
- Gs stimulates adenylate cyclase, which increases levels of cAMP in the cell.
- Gi inhibits adenylate cyclase, which decreases levels of cAMP in the cell.
- Gq activates phospholipase C, which cleaves a phospholipid from the membrane to form PIP2. PIP2 is then cleaved into DAG and IP3; IP3 can open calcium channels in the endoplasmic reticulum, increasing calcium levels in the cell.
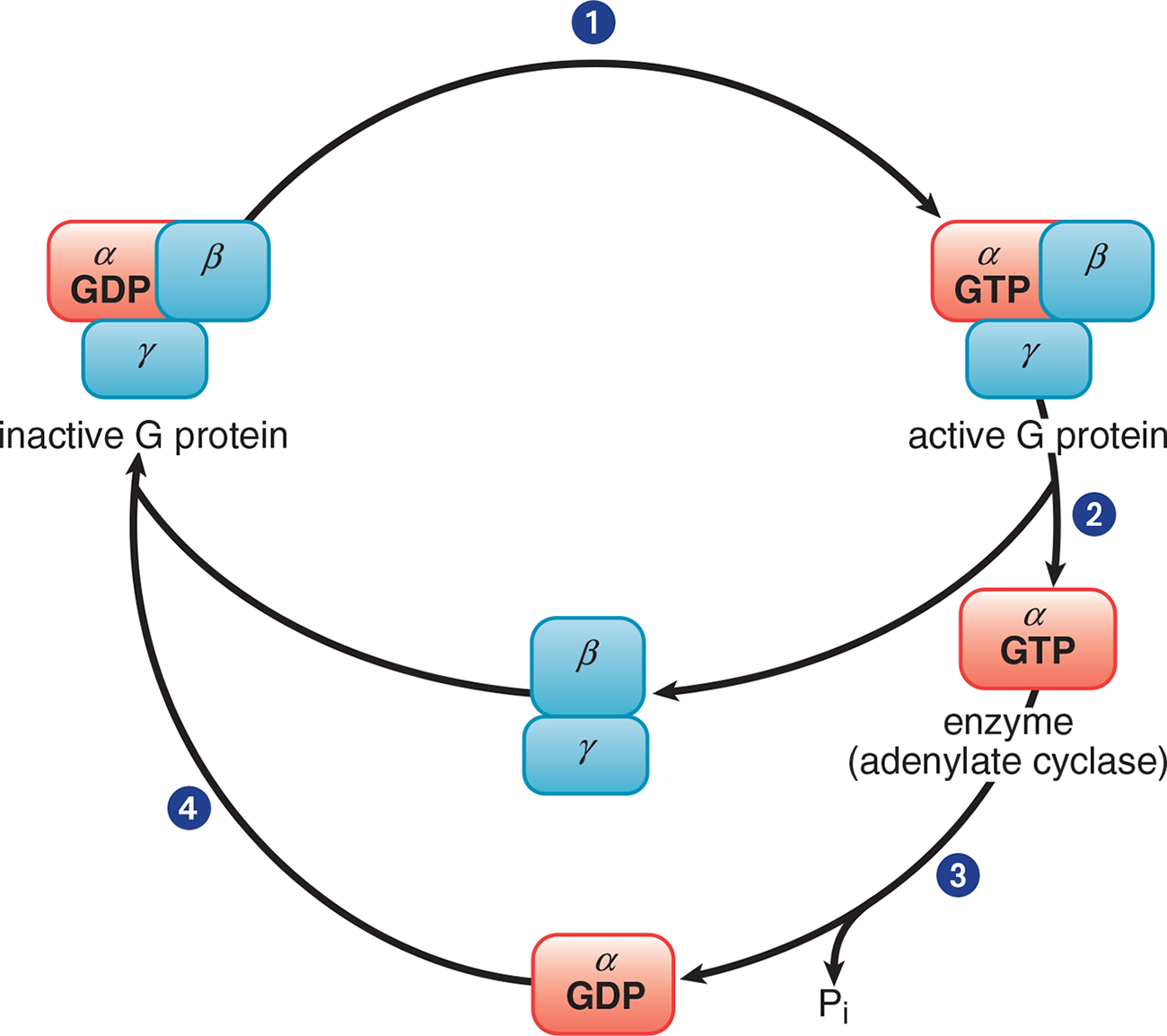
Figure 3.5 Trimeric G Protein Cycle (Gs or Gi)
MNEMONIC
Functions of heterotrimeric G proteins:
- Gs stimulates.
- Gi inhibits.
- “Mind your p’s and q’s”: Gq activates phospholipase C.
The three subunits that comprise the G protein are α, β, and γ. In its inactive form, the ***αsubunitbinds GDP and is in a complex with theβandγ* subunits**. When a ligand binds to the GPCR, the receptor becomes activated and, in turn, engages the corresponding G protein, as shown in Step 1 of Figure 3.5. Once GDP is replaced with GTP, the α subunit is able to dissociate from the β and γ subunits (Step 2). The activated α subunit alters the activity of adenylate cyclase. If the α subunit is αs, then the enzyme is activated; if the α subunit is αi, then the enzyme is inhibited. Once GTP on the activated α subunit is dephosphorylated to GDP (Step 3), the α subunit will rebind to the β and γ subunits (Step 4), rendering the G protein inactive.
MCAT CONCEPT CHECK 3.2:
Before you move on, assess your understanding of the material with these questions.
- Contrast enzyme-linked receptors with G protein-coupled receptors:
- What type of ion channel is active at all times?
________________________________________
- How do transport kinetics differ from enzyme kinetics?
________________________________________
3.3 Protein Isolation
LEARNING OBJECTIVES
After Chapter 3.3, you will be able to:
- Recall the major categories of electrophoresis and chromatography
- Select the appropriate protein isolation method in a given situation
- Identify the mobile and stationary phases when given a separatory apparatus:
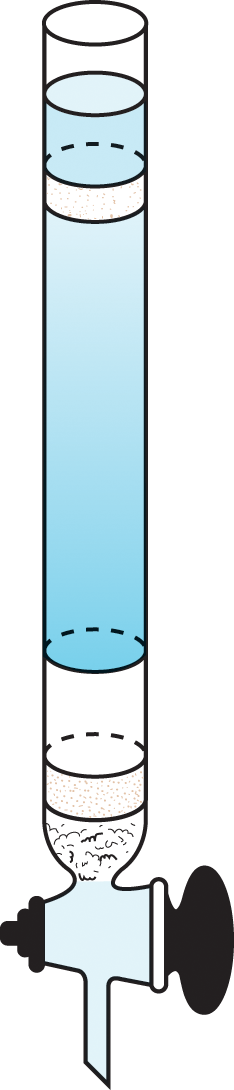
In order to better understand a specific protein, it is important to be able to isolate the protein for study. The purification of proteins can be considered an art form when one considers the difficulty of isolating just one protein from a cell containing hundreds to thousands of proteins. Luckily, as Chapter 1 of MCAT Biochemistry Review highlighted, there is a great amount of variation in the physical and chemical properties of proteins and these differences can be exploited in order to purify the protein of interest. Proteins and other biomolecules are isolated from body tissues or cell cultures by cell lysis and homogenization—crushing, grinding, or blending the tissue of interest into an evenly mixed solution. Centrifugation can then isolate proteins from much smaller molecules before other isolation techniques must be employed. The most common isolation techniques are electrophoresis and chromatography, either of which can be used for native or denatured proteins.
Electrophoresis
One method of separating proteins is with electrophoresis. In molecular biology, this is one of the most important analytical techniques. Electrophoresis works by subjecting compounds to an electric field, which moves them according to their net charge and size. Negatively charged compounds will migrate toward the positively charged anode, and positively charged compounds will migrate toward the negatively charged cathode. The velocity of this migration, known as the migration velocity of a molecule, v, is directly proportional to the electric field strength, E, and to the net charge on the molecule, z, and is inversely proportional to a frictional coefficient, f, which depends on the mass and shape of the migrating molecules:
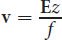
Equation 3.1 Polyacrylamide gel is the standard medium for protein electrophoresis. The gel is a slightly porous matrix mixture, which solidifies at room temperature. Proteins travel through this matrix in relation to their size and charge. The gel acts like a sieve, allowing smaller particles to pass through easily while retaining large particles. Therefore, a molecule will move faster through the medium if it is small, highly charged, or placed in a large electric field. Conversely, molecules will migrate slower (or not at all) when they are bigger and more convoluted, electrically neutral, or placed in a small electric field. The size of a standard polyacrylamide gel allows multiple samples to be run simultaneously, as shown in Figure 3.6.
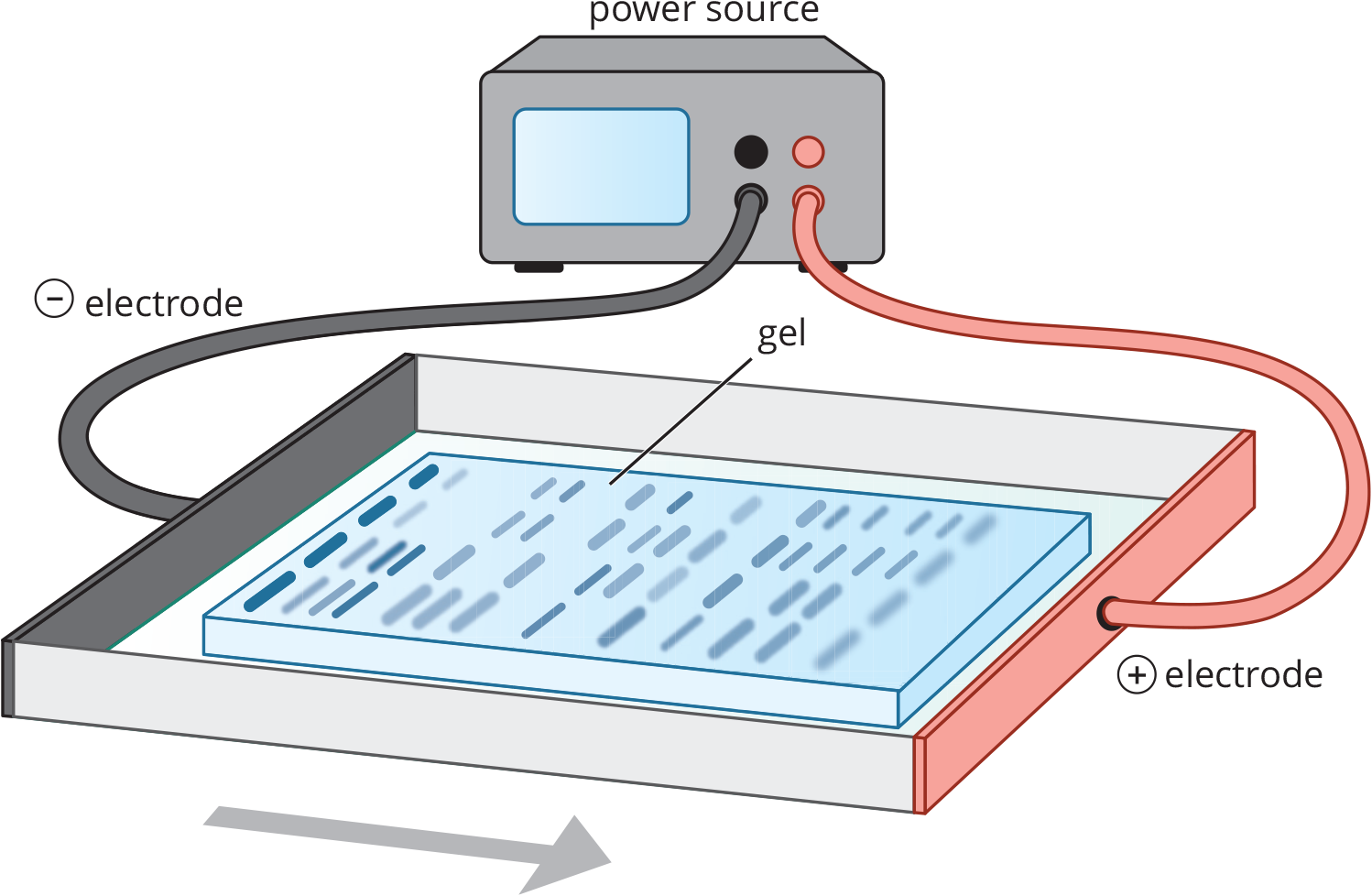
Figure 3.6 Electrophoresis As an electrolytic (nonspontaneous) cell, electrophoresis moves charged particles toward their respective oppositely charged electrodes; the larger the particle, the more slowly it migrates.
BRIDGE
Electrophoresis uses an electrolytic cell (ΔG > 0, Ecell < 0), as described in Chapter 12 of MCAT General Chemistry Review. Remember that anions always move toward the anode and cations always move toward the cathode.
Native PAGE
Polyacrylamide gel electrophoresis (PAGE) is a method for analyzing proteins in their native states. Unfortunately, PAGE is limited by the varying mass-to-charge and mass-to-size ratios of cellular proteins because multiple different proteins may experience the same level of migration. In PAGE, the functional native protein can be recovered from the gel after electrophoresis, but only if the gel has not been stained because most stains denature proteins. PAGE is most useful to compare the molecular size or the charge of proteins known to be similar in size from other analytic methods like SDS-PAGE (described below) or size-exclusion chromatography.
SDS-PAGE
Sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis is a useful tool because it separates proteins on the basis of relative molecular mass alone. The SDS-PAGE technique starts with the premise of PAGE but adds SDS, a detergent that disrupts all noncovalent interactions. It binds to proteins and creates large chains with net negative charges, thereby neutralizing the protein’s original charge and denaturing the protein, as shown in Figure 3.7. As the proteins move through the gel, the only variables affecting their velocity are E, the electric field strength, and f, the frictional coefficient, which depends on mass. After separation, the gel can be stained so the protein bands can be visualized and the results recorded.
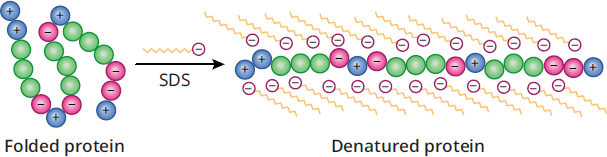
Figure 3.7 Protein Denaturation by SDS
MCAT EXPERTISE
For analytic purposes, protein atomic mass is typically expressed in daltons (Da). A dalton is an alternative term for molar mass
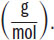
The average molar mass of one amino acid is ~110 daltons, or 110
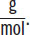
.
Reducing PAGE
Reducing PAGE works on the same general principle as native PAGE in that all non-covalent interactions are preserved, but the reducing agent will break any disulfide links in the tertiary and quaternary structure by reducing them to cysteines. As seen in Figure 3.8, if disulfide bonds link two polypeptide chains together, in native PAGE or even SDS-PAGE, those two proteins would migrate together as one unit. However, in the presence of a reducing agent like dithiothreitol (DTT) or β-mercaptoethanol (βME), the disulfide bond would be broken and the two proteins would migrate separately. Comparing native and reducing PAGE together allows researchers to quickly gain insight into how the quaternary structure of a protein is organized.
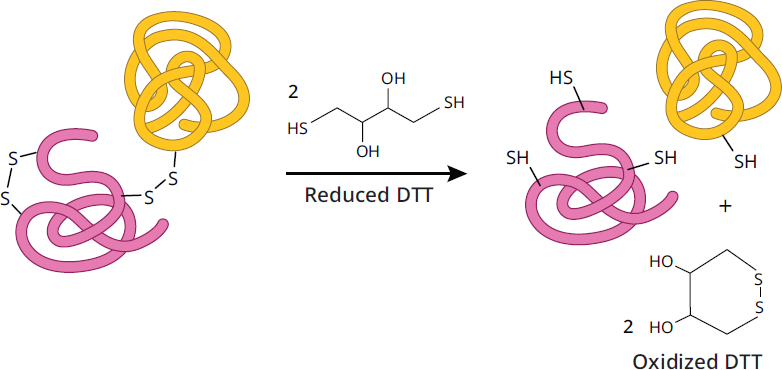
Figure 3.8 Effect of Reducing Agents on Disulfide Bonds
Interpreting PAGE Results
While PAGE can provide valuable information about the molecular weight of proteins, comparing between different conditions in PAGE on the same protein sample allows for researchers to gain insight into the composition of that protein’s quaternary structure. Let’s use Figure 3.9 to walk through an example analysis.
- Examining the top gel labeled “Native,” where no denaturant or reducing agent is added, the molecular weight markers in the first lane indicate the protein has a molecular weight of approximately 250 kDa (in the real world, native PAGE does not always provide an accurate molecular weight; however, the MCAT will always assume it does). A native gel by itself does not allow for any conclusions on quaternary structure, as protein quaternary structures and protein complexes are not affected in native PAGE.
- The second gel labeled “SDS nonreducing” has the detergent SDS added, which denatures proteins by breaking any noncovalent interaction and coating the proteins in a uniform negative charge. SDS, however, cannot break disulfide bonds. In this gel, there are two bands: one band at 200 kDa, and another at 50 kDa. When compared to the native gel, this SDS nonreducing gel indicates the protein in the sample has two polypeptides noncovalently linked together that the SDS interrupted.
- Finally, in the bottom gel labeled SDS reducing, both SDS and a reducing agent were added to the protein sample. As before, SDS breaks any noncovalent interactions, but with the addition of a reducing agent, disulfide bonds will now also be broken. There are two bands again in this gel, but this time at 100 kDa and 50 kDa. Since the 50 kDa band remained the same, it means that the 200 kDa band protein seen on the SDS nonreducing gel is most likely a homodimer (a complex made of two identical proteins) linked together by a disulfide bond. The final conclusion from the analysis of these three different types of PAGE is that the 250 kDa protein is a heterotrimer (a complex made up of three non-identical proteins) that contains a homodimer with two subunits of 100 kDa proteins linked together with a disulfide bond, and a third 50 kDa subunit subunit noncovalently bound to the homodimer.
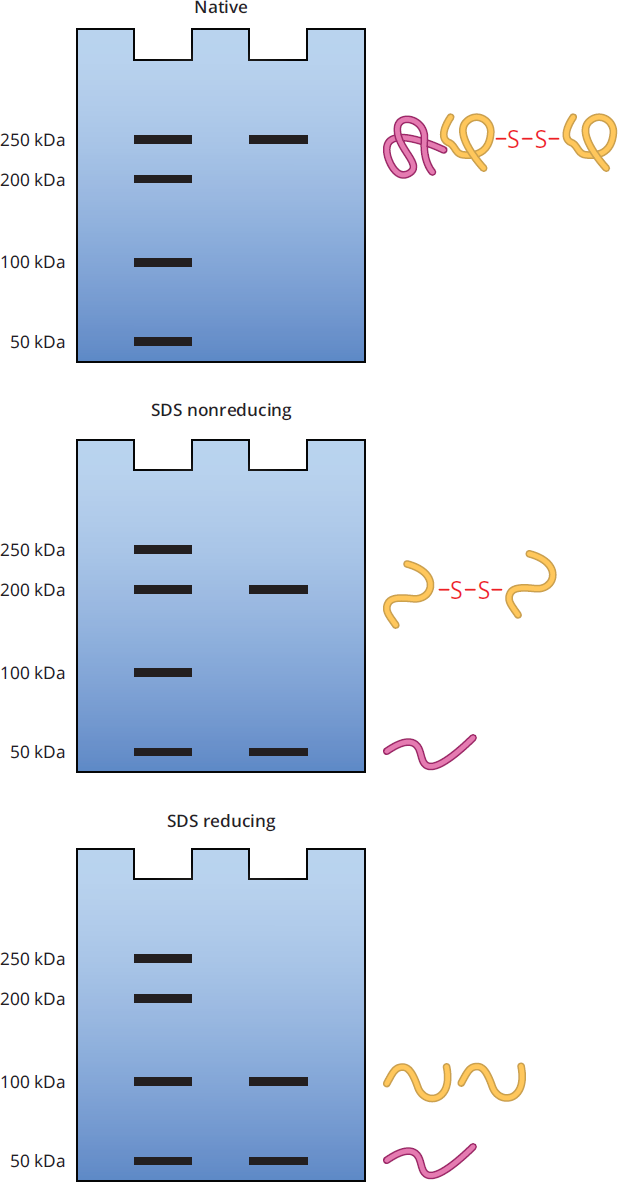
Figure 3.9 Native, SDS Nonreducing, and SDS Reducing PAGE on Example Protein
Isoelectric Focusing
Proteins can be separated on the basis of their isoelectric point (pI). The pI is the pH at which the protein or amino acid is electrically neutral, with an equal number of positive and negative charges. For individual amino acids this electrically neutral form is called a zwitterion.For most amino acids, the zwitterion form occurs when the amino group is protonated, the carboxyl group is deprotonated, and any side chain is electrically neutral. The exceptions to this general rule are arginine and lysine, two amino acids with basic side chains. For these amino acids, whose nitrogen-containing side chains have higher pKa values than their amino groups, the zwitterion form occurs when the amino group is deprotonated (and therefore electrically neutral), while the nitrogenous side chain is protonated and the carboxyl group is deprotonated. The calculation of the pI for an amino acid was discussed in Chapter 1 of MCAT Biochemistry Review. For polypeptides, the isoelectric point is primarily determined by the relative numbers of acidic and basic amino acids.
Isoelectric focusing exploits the acidic and basic properties of amino acids by separating on the basis of pI. As seen in Figure 3.10, the mixture of proteins is placed in a gel with a pH gradient (acidic gel at the positive anode, basic gel at the negative cathode, and neutral in the middle). An electric field is then generated across the gel. Proteins that are positively charged will begin migrating toward the cathode and proteins that are negatively charged will begin migrating toward the anode. As a protein reaches the portion of gel where the pH is equal to the protein’s pI, the protein takes on a neutral charge and will stop moving.
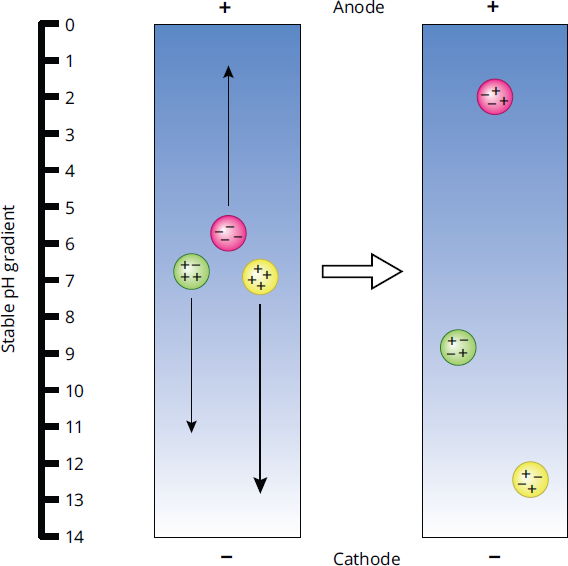
Figure 3.10 Isoelectric Focusing Gel Electrophoresis
KEY CONCEPT
In isoelectric focusing, a protein stops moving when pH = pI.
Let’s take a deeper look. We’ll start with a protein that has a pI of 9. When the protein is in an environment with a pH of 9, it will carry no net charge. If we place this protein onto the gel at a pH of 7, there will be more protons around the protein. These protons will attach to the available basic sites on the protein, creating a net positive charge on the molecule, as occurs with the far left green protein in Figure 3.10. This charge will then cause the protein to be attracted to the negatively charged cathode, which is located on the basic side of the gradient. As the protein moves closer to the cathode, the pH of the gel slowly increases. Eventually, as the protein nears a pH of 9, the protons creating the positive charge will dissociate, and the protein will become neutral again. Remember, cations (positively charged species) will always migrate to the cathode, and anions (negatively charged species) will always migrate to the anode.
MNEMONIC
Cations go to the Cathode Anions go to the Anode
Chromatography
Chromatography is another tool that uses physical and chemical properties to separate and identify compounds from a complex mixture. Chromatography refers to a variety of techniques that require the homogenized protein mixture to be fractionated through a porous matrix. One of the reasons chromatography is a valuable tool is that the isolated proteins are immediately available for identification and quantification. In all forms of chromatography discussed here, the concept is identical: the more similar the compound is to its surroundings (by polarity, charge, and so on), the more it will stick to and move slowly through its surroundings. Chromatography is preferred over electrophoresis when large amounts of protein are being separated.
BRIDGE
Chromatography and other separatory methods are also discussed in Chapter 12 of MCAT Organic Chemistry Review.
The process begins by placing the sample onto a solid medium called the stationary phase or adsorbent. The next step is to run the mobile phase through the stationary phase. This will allow the sample to run through the stationary phase, or elute. Depending on the relative affinity of the sample for the stationary and mobile phases, different substances will migrate through at different speeds. That is, components that have a high affinity for the stationary phase will barely migrate at all; components with a high affinity for the mobile phase will migrate much more quickly. The amount of time a compound spends in the stationary phase is referred to as the retention time. Varying retention times of each compound in the solution results in separation of the components within the stationary phase, or partitioning, as demonstrated in Figure 3.11. Each component can then be isolated individually for study.

Figure 3.11 Partitioning of Black Ink Thin-layer chromatography; original spot placed on the bottom of the card. Components with high retention times remain near the bottom of the card; components with low retention times have migrated toward the top of the card.
KEY CONCEPT
All chromatography is about the affinity of a substance for the mobile and stationary phases, except for size-exclusion chromatography.
We can use myriad different media as our stationary phase, each one exploiting different properties that allow us to separate out our compound. In chromatography for protein separation, common properties include charge, pore size, and specific affinities.
Column Chromatography
In column chromatography, a column is filled with silica or alumina beads as an adsorbent, and gravity moves the solvent and compounds down the column, shown in Figure 3.12. As the solution flows through the column, both size and polarity have a role in determining how quickly a compound moves through the polar silica or alumina beads: the less polar the compound, the faster it can elute through the column (short retention time). In column chromatography, the solvent polarity, pH, or salinity can easily be changed to help elute the protein of interest.
Eventually, the solvent drips out of the end of the column, and different fractions that leave the column are collected over time. Each fraction contains bands that correspond to different compounds. After collection, the solvent can be evaporated and the compounds of interest kept. Column chromatography is particularly useful in biochemistry because it can be used to separate and collect other macromolecules besides proteins, such as nucleic acids.
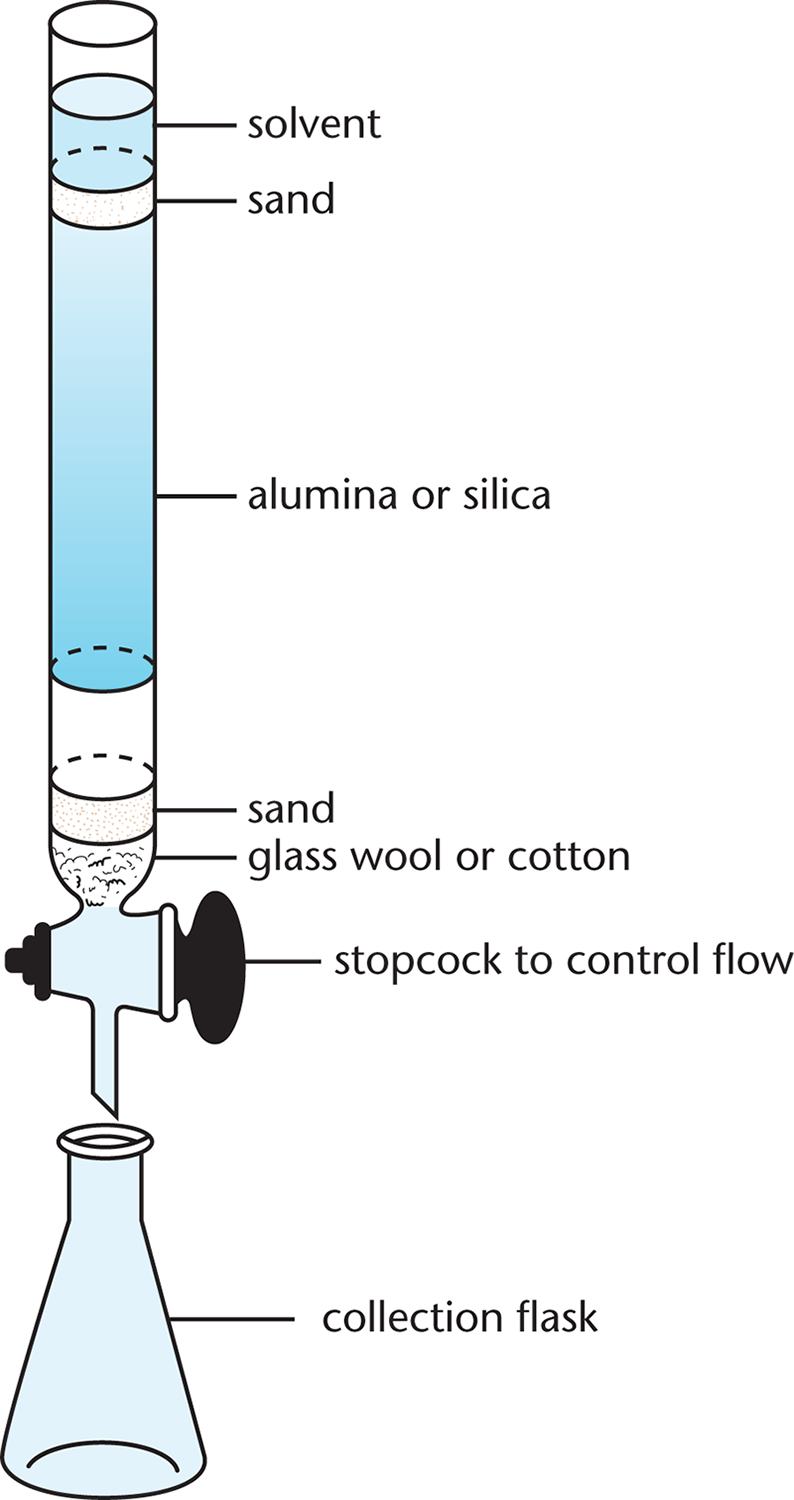
Figure 3.12 Column Chromatography The sample is added at the top of the column and a solvent is poured over it. The more similar the sample is to the solvent (mobile phase), the more quickly it will elute; the more similar it is to the alumina or silica (stationary phase), the more slowly it will elute—if at all.
Ion-Exchange Chromatography
In this method, the beads in the column are coated with charged substances, so they attract or bind compounds that have an opposite charge, as can be seen in Figure 3.13. For instance, a positively charged column will attract and hold a negatively charged protein as it passes through the column, either increasing its retention time or retaining it completely. After all other compounds have moved through the column, a salt gradient is used to elute the charged molecules that have stuck to the column.
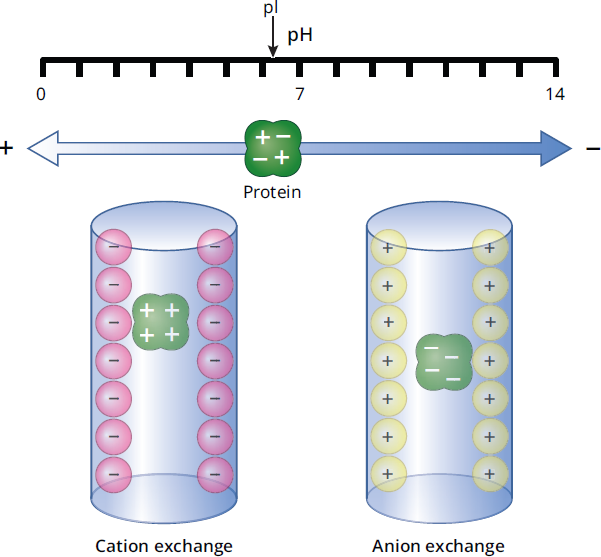
Figure 3.13 Ion-Exchange Chromatography
Size-Exclusion Chromatography
In this method, the beads used in the column contain tiny pores of varying sizes. As demonstrated in Figure 3.14, these tiny pores allow small compounds to enter the beads, thus slowing them down. Large compounds can’t fit into the pores, so they will move around them and travel through the column faster. It is important to remember that in this type of chromatography, the small compounds are slowed down and retained longer—which may be counterintuitive. The size of the pores may be varied so that molecules of different molecular weights can be fractionated. A common approach in protein purification is to use an ion-exchange column followed by a size-exclusion column.
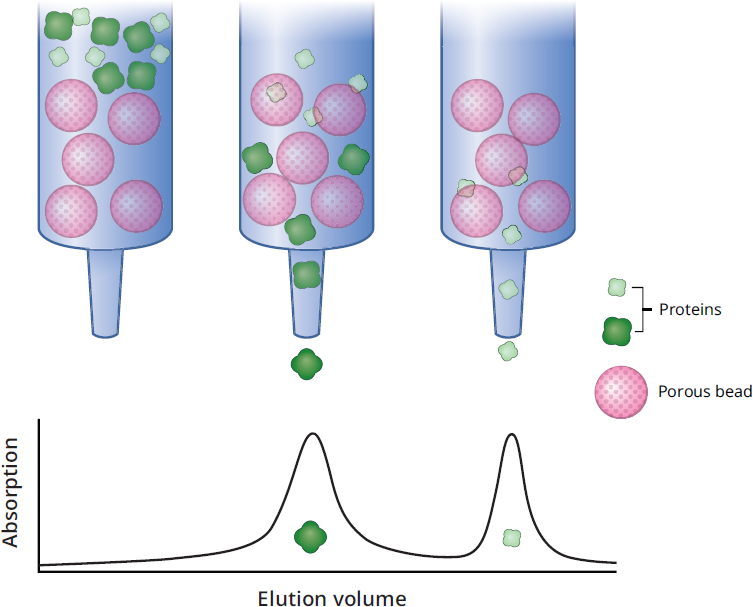
Figure 3.14 Size-Exclusion Chromatography
Affinity Chromatography
We can also customize columns to bind any protein of interest by creating a column with high affinity for that protein. This can be accomplished by coating beads with a receptor that binds the protein or a specific antibody to the protein; in either case, the protein is retained in the column. Nickel is one common choice for stationary phase, as illustrated in Figure 3.15, that is used in the separation of genetically engineered proteins with histidine tags. Other common stationary phase molecules include antibodies (also depicted in Figure 3.15) or antigens, as well as enzyme substrate analogues, which mimic the natural substrate for an enzyme of interest. Once the protein is retained in the column, it can be eluted by washing the column with a free receptor (or target or antibody), which will compete with the bead-bound receptor and ultimately free the protein from the column. Eluents can also be created with a specific pH or salinity level that disrupts the bonds between the ligand and the protein of interest. The only drawback of the elution step is that the recovered substance can be bound to the eluent. If, for example, the eluent was an inhibitor of an enzyme, it could be difficult to remove.
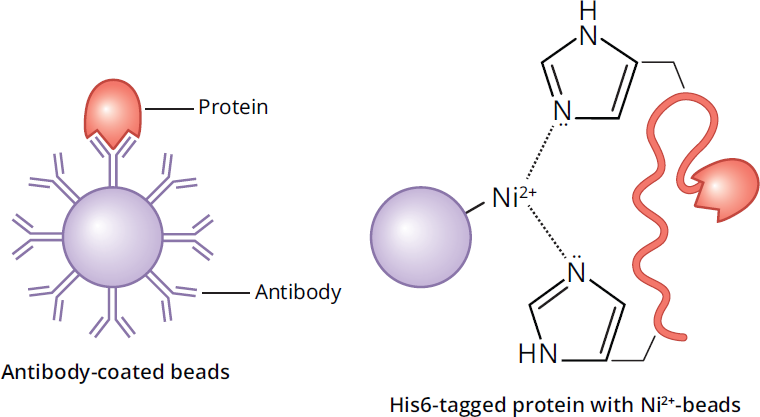
Figure 3.15 Affinity Chromatography
BIOCHEMISTRY GUIDED EXAMPLE WITH EXPERT THINKING
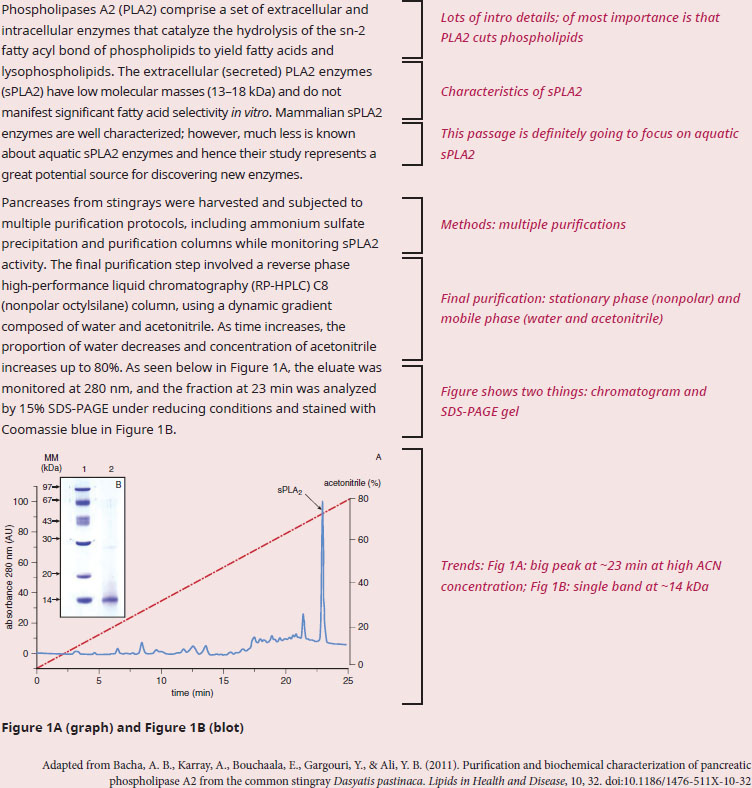
According to the data, what is the approximate monomeric molecular weight of sPLA2? Why is it biologically relevant that sPLA2 required a high acetonitrile concentration to elute off the column?
With "according to the data", this question stem is letting us know we'll need to use one or both parts of the figure to answer. So, let’s start with a quick scan of the passage to see the topic and the kind of information presented to better understand the context of the data. The text is focused on a protein called phospholipase A2 (PLA2), and Figure 1 shows peaks with a gel picture inset. The bulk of the passage seems method-heavy (as indicated by the usage of column names and Figure references). We have two questions to answer: the first is asking for the size of the protein, while the second asks for a connection between the biology behind PLA2 and the way PLA2 interacts with the column.
To determine the monomeric molecular weight of sPLA2, we'll need information from the SDS-PAGE data in Figure 1B. We know from our background content knowledge that an SDS-PAGE gel under reducing conditions will denature proteins and sever any disulfide bonds. Therefore, the bands that appear should only correspond to monomers, and monomers are what this question asked about. The first lane is a series of molecular weight standards measured in kilodaltons (kDa), which are used to estimate the molecular weight of unknown proteins. We’re told that the fraction at 23 minutes, which corresponds to the sPLA2 peak on the chromatogram, is analyzed in lane 2 of the gel. The band in lane 2 corresponding to sPLA2 is right next to the 14 kDa standard in lane 1, implying that the sPLA2 protein has a weight of approximately 14 kDa as well. As an extra measure of verification, paragraph 1 stated that sPLA2s have a molecular weight between 13–18 kDa.
Paragraph 2 discusses the method of the experiment, which will help us answer the second part of the question stem. For all the detail in paragraph 2, RP-HPLC is just a type of a column chromatography. Like all types of column chromatography, RP-HPLC has a stationary phase (the column) and a mobile phase (the solvent). We're told that the column is nonpolar, which means nonpolar molecules will stick inside the column. We're also told that the solvent changes composition over time—starting as mostly water, but ending as mostly acetonitrile. Why is this change in composition significant? We recall that water is a polar solvent, while acetonitrile is closer to nonpolar. Therefore, early on, when the solvent is mostly water, polar molecules will flush out of the column because like dissolves like. But near the end of the experiment, when the solvent is mostly acetonitrile, the now nonpolar solvent will dislodge nonpolar molecules. So, nonpolar molecules will elute off the column near the end of the experiment. Finally, we observe that the sPLA2 protein elutes off the column near the end, when the solvent is mostly acetonitrile. This observation implies that the sPLA2 protein is net nonpolar; the nonpolar protein had initially stuck to the nonpolar column, but the protein eventually eluted out of the column once the solvent became nonpolar enough to solvate the protein.
We can thus conclude that sPLA2 monomers have a molecular weight of around 14 kDa and, from the experimental procedure, we can conclude that sPLA2 proteins have a high affinity for nonpolar substances—first for the walls of the nonpolar column, and later for the nonpolar acetonitrile solvent. This behavior makes biological sense because the biological substrates of the sPLA2 protein are phospholipids!
MCAT CONCEPT CHECK 3.3:
Before you move on, assess your understanding of the material with these questions.
- What separation methods can be used to isolate a protein on the basis of isoelectric point?
________________________________________
- What are the relative benefits of native PAGE compared to SDS-PAGE?
________________________________________
- What are two potential drawbacks of affinity chromatography?
- ___________________________________
- ___________________________________
- True or False: In size-exclusion chromatography, the largest molecules elute first.
3.4 Protein Analysis
LEARNING OBJECTIVES
After Chapter 3.4, you will be able to:
- Recall the traits that are typically analyzed in proteins
- Describe the Edman degradation and the Bradford assay
- Recognize the limitations of protein separation and analysis techniques
Separating proteins from one another is generally only the first step in analysis. The next step is to study the isolated protein. Protein structure, function, or quantity is often of interest for a researcher or a commercial laboratory. Even after protein identification, protein analysis tools may be used. For example, in the case of protein synthesis for commercial use, purity of the product must be periodically assessed. The protein can be studied as a whole or broken down so that its parts can be examined.
Protein Structure
Protein structure can be determined through X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy. Before crystallographic analysis, the protein must be isolated and crystallized. X-ray crystallography is the most reliable and common method; 75 percent of the protein structures known today were analyzed through this method. Crystallography measures electron density on an extremely high-resolution scale and can also be used for nucleic acids. An X-ray diffraction pattern is generated in this method, as shown in Figure 3.16. The small dots in the diffraction pattern can then be interpreted to determine the protein’s structure.
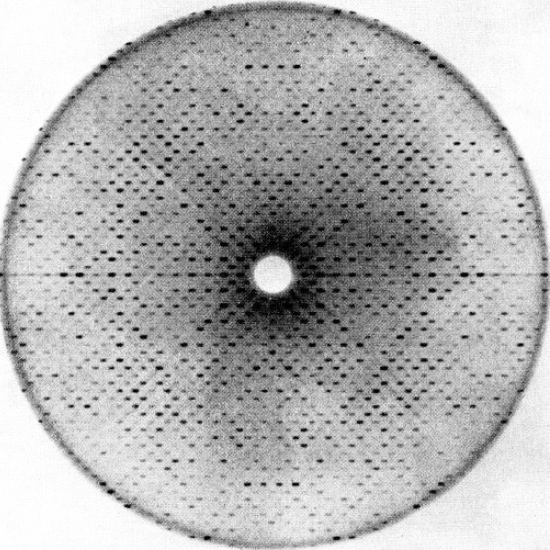

Figure 3.16 Diffraction Pattern in X-Ray Crystallography
BRIDGE
NMR and other forms of spectroscopy are also discussed in Chapter 11 of MCAT Organic Chemistry Review.
The minority of protein structure determination (25 percent) has been accomplished through NMR, as discussed in Chapter 11 of MCAT Organic Chemistry Review.
Amino Acid Composition
The amino acids that compose a protein can be determined by complete protein hydrolysis and subsequent chromatographic analysis. However, the random nature of hydrolysis prevents amino acid sequencing. To determine the primary structure of a protein, sequential digestion of the protein with specific cleavage enzymes is used. Small proteins are best analyzed with the Edman degradation, which uses cleavage to sequence proteins of up to 50 to 70 amino acids. The Edman degradation selectively and sequentially removes the N-terminal amino acid of the protein, which can be analyzed via mass spectroscopy.
For larger proteins, digestion with chymotrypsin, trypsin, and cyanogen bromide, a synthetic reagent, may be used. This digestion selectively cleaves proteins at specific amino acid residues, creating smaller fragments that can then be analyzed by electrophoresis or the Edman degradation. Because disulfide links and salt bridges are broken to reduce the protein to its primary structure, their positions cannot be determined by these methods.
REAL WORLD
Like PCR gene sequencing, protein amino acid sequencing can be automated in a stepwise manner. By combining the information from both techniques, researchers can determine where on a chromosome the gene coding a particular protein resides.
Activity Analysis
Protein activity is generally determined by monitoring a known reaction with a given concentration of substrate and comparing it to a standard. Activity is correlated with substrate concentration, but it is also affected by the methods used to purify the enzyme and the specific conditions of the assay. Reactions that lead to a color change have particular applicability because samples with color changes can be rapidly detected using chromatographic analysis. Enzyme activity is measured as moles of substrate converted by the enzyme into product per unit of time, with an SI unit of katal, though it is much more commonly measured in units of U. Specific activity, however, measures the activity of the enzyme per mass of total protein, and is often used to track the purity of an enzyme within a mixture, like during purification. Comparing the total number of units of activity at the end of the purification to the total units at the beginning of purification can provide an estimate of the purification yield.
Concentration Determination
Concentration is determined almost exclusively through spectroscopy. Because proteins contain aromatic side chains, they can be analyzed with UV spectroscopy without any treatment; however, this type of analysis is particularly sensitive to sample contaminants. Proteins also cause colorimetric changes with specific reactions, particularly the bicinchoninic acid (BCA) assay, Lowry reagent assay, and Bradford protein assay. The Bradford method is most common because of its reliability and simplicity in basic analyses.
Bradford Protein Assay
The Bradford protein assay mixes a protein in solution with Coomassie Brilliant Blue dye. The dye is protonated and green-brown in color prior to mixing with proteins, as depicted in Figure 3.17. The dye gives up protons upon binding to amino acid groups, turning blue in the process. Ionic attractions between the dye and the protein then stabilize this blue form of the dye; thus, increased protein concentrations correspond to a larger concentration of blue dye in solution. Samples of known protein concentrations are reacted with the Bradford reagent and then absorbance is measured to create a standard curve. The unknown sample is then exposed to the same conditions, and the concentration is determined based on the standard curve. This is a very accurate method when only one type of protein is present in solution, but because of variable binding of the Coomassie dye with different amino acids, it is less accurate when more than one protein is present. The Bradford protein assay is limited by the presence of detergent in the sample or by excessive buffer.
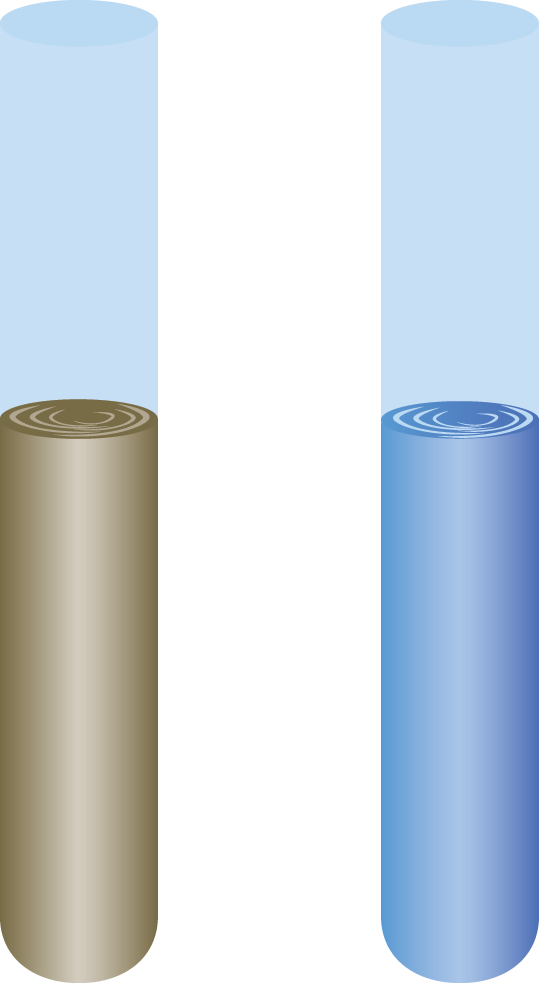

Figure 3.17 Bradford Protein Assay The acidic form (left) has a brown-green hue; the basic form (right), which is created by interactions with proteins in solution, has a brilliant blue hue.
MCAT CONCEPT CHECK 3.4:
Before you move on, assess your understanding of the material with these questions.
- Why are proteins analyzed after isolation?
________________________________________
- What factors would cause an activity assay to display lower activity than expected after concentration determination?
________________________________________
- True or False: The Edman degradation proceeds from the carboxy (C-) terminus.
Conclusion
In this chapter, we have explored the nonenzymatic aspects of proteins as well as the ways proteins can be analyzed. The cellular proteins and their functions that we discussed included structural proteins that play a role in cytoskeletal architecture, motor proteins involved in muscle contraction and movement along the cytoskeleton, and other proteins that play more complex roles such as binding, immunologic function, and biosignaling. The more complex proteins involved in biosignaling highlighted in this chapter included ion channels, enzyme-linked receptors, and G protein-coupled receptors. Finally, we determined how to isolate and identify a protein and its relevant properties.
In the next chapter, we’ll turn our attention to another class of biomolecules: carbohydrates. As we transition from amino acids, peptides, and proteins to monosaccharides, oligosaccharides, and polysaccharides, look for key connections between the different types of macromolecules used by the body for structure and as fuel sources. In the end, all biomolecules are related to each other through metabolism, which we’ll explore in Chapters 9 through 12 of MCAT Biochemistry Review.
GO ONLINE!
You've reviewed the content, now test your knowledge and critical thinking skills by completing a test-like passage set in your online resources!
CONCEPT SUMMARY
Cellular Functions
- Structural proteins compose the cytoskeleton, anchoring proteins, and much of the extracellular matrix.
- The most common structural proteins are collagen, elastin, keratin, actin, and tubulin.
- They are generally fibrous in nature.
- Motor proteins have one or more heads capable of force generation through a conformational change.
- They have catalytic activity, acting as ATPases to power movement.
- Muscle contraction, vesicle movement within cells, and cell motility are the most common applications of motor proteins.
- Common examples include myosin, kinesin, and dynein.
- Binding proteins bind a specific substrate, either to sequester it in the body or hold its concentration at steady state.
- Cell adhesion molecules (CAM) allow cells to bind to other cells or surfaces.
- Cadherins are calcium-dependent glycoproteins that hold similar cells together.
- Integrins have two membrane-spanning chains and permit cells to adhere to proteins in the extracellular matrix. Some also have signaling capabilities.
- Selectins allow cells to adhere to carbohydrates on the surfaces of other cells and are most commonly used in the immune system.
- Antibodies (or immunoglobulins, Ig) are used by the immune system to target a specific antigen, which may be a protein on the surface of a pathogen (invading organism) or a toxin.
- Immunoglobulins contain a constant region and a variable region; the variable region is responsible for antigen binding.
- Two identical heavy chains and two identical light chains form a single antibody; they are held together by disulfide linkages and noncovalent interactions.
Biosignaling
- Ion channels can be used for regulating ion flow into or out of a cell. There are three main types of ion channels.
- Ungated channels are always open.
- Voltage-gated channels are open within a range of membrane potentials.
- Ligand-gated channels open in the presence of a specific binding substance, usually a hormone or neurotransmitter.
- Enzyme-linked receptors participate in cell signaling through extracellular ligand binding and initiation of second messenger cascades.
- G protein-coupled receptors have a membrane-bound protein associated with a trimeric G protein. They also initiate second messenger systems.
- Ligand binding engages the G protein.
- GDP is replaced with GTP; the α subunit dissociates from the β and γ subunits.
- The activated α subunit alters the activity of adenylate cyclase or phospholipase C.
- GTP is dephosphorylated to GDP; the α subunit rebinds to the β and γ subunits.
Protein Isolation
- Electrophoresis uses a gel matrix to observe the migration of proteins in response to an electric field.
- Native PAGE maintains the protein’s shape, but results are difficult to compare because the mass-to-charge ratio differs for each protein.
- SDS-PAGE denatures the proteins and masks the native charge so that comparison of size is more accurate, but the functional protein cannot be recaptured from the gel.
- Isoelectric focusing separates proteins by their isoelectric point (pI); the protein migrates toward an electrode until it reaches a region of the gel where pH = pI of the protein.
- Chromatography separates protein mixtures on the basis of their affinity for a stationary phase or a mobile phase.
- Column chromatography uses beads of a polar compound, like silica or alumina (stationary phase), with a nonpolar solvent (mobile phase).
- Ion-exchange chromatography uses a charged column and a variably saline eluent.
- Size-exclusion chromatography relies on porous beads. Larger molecules elute first because they are not trapped in the small pores.
- Affinity chromatography uses a bound receptor or ligand and an eluent with free ligand or a receptor for the protein of interest.
Protein Analysis
- Protein structure is primarily determined through X-ray crystallography after the protein is isolated, although NMR can also be used.
- Amino acid composition can be determined by simple hydrolysis, but amino acid sequencing requires sequential degradation, such as the Edman degradation.
- Activity levels for enzymatic samples are determined by following the process of a known reaction, often accompanied by a color change.
- Protein concentration is also determined colorimetrically, either by UV spectroscopy or through a color change reaction.
- BCA assay, Lowry reagent assay, and Bradford protein assay each test for protein and have different advantages and disadvantages.
- The Bradford protein assay, which uses a color change from brown-green to blue, is most common.
ANSWERS TO CONCEPT CHECKS
3.1
- Cytoskeletal proteins tend to be fibrous with repeating domains, while motor proteins tend to have ATPase activity and binding heads. Both types of protein function in cellular motility.
- False. An enzyme is a protein or RNA molecule with catalytic activity, which motor proteins do have. Motor function is generally considered nonenzymatic, but the ATPase functionality of motor proteins indicates that these molecules do have catalytic activity.
- If the binding protein is present in sufficiently high quantities relative to the substrate, nearly all substrate will be bound despite a low affinity.
-
Cell Adhesion Molecule Type of Adhesion Cadherin Two cells of the same or similar type using calcium
Integrin One cell to proteins in the extracellular matrix
Selectin One cell to carbohydrates, usually on the surface of other cells
- Antigen–antibody interactions can result in neutralization of the pathogen or toxin, opsonization (marking) of the antigen for destruction, or creation of insoluble antigen–antibody complexes that can be phagocytized and digested by macrophages (agglutination).
**3.2**
-
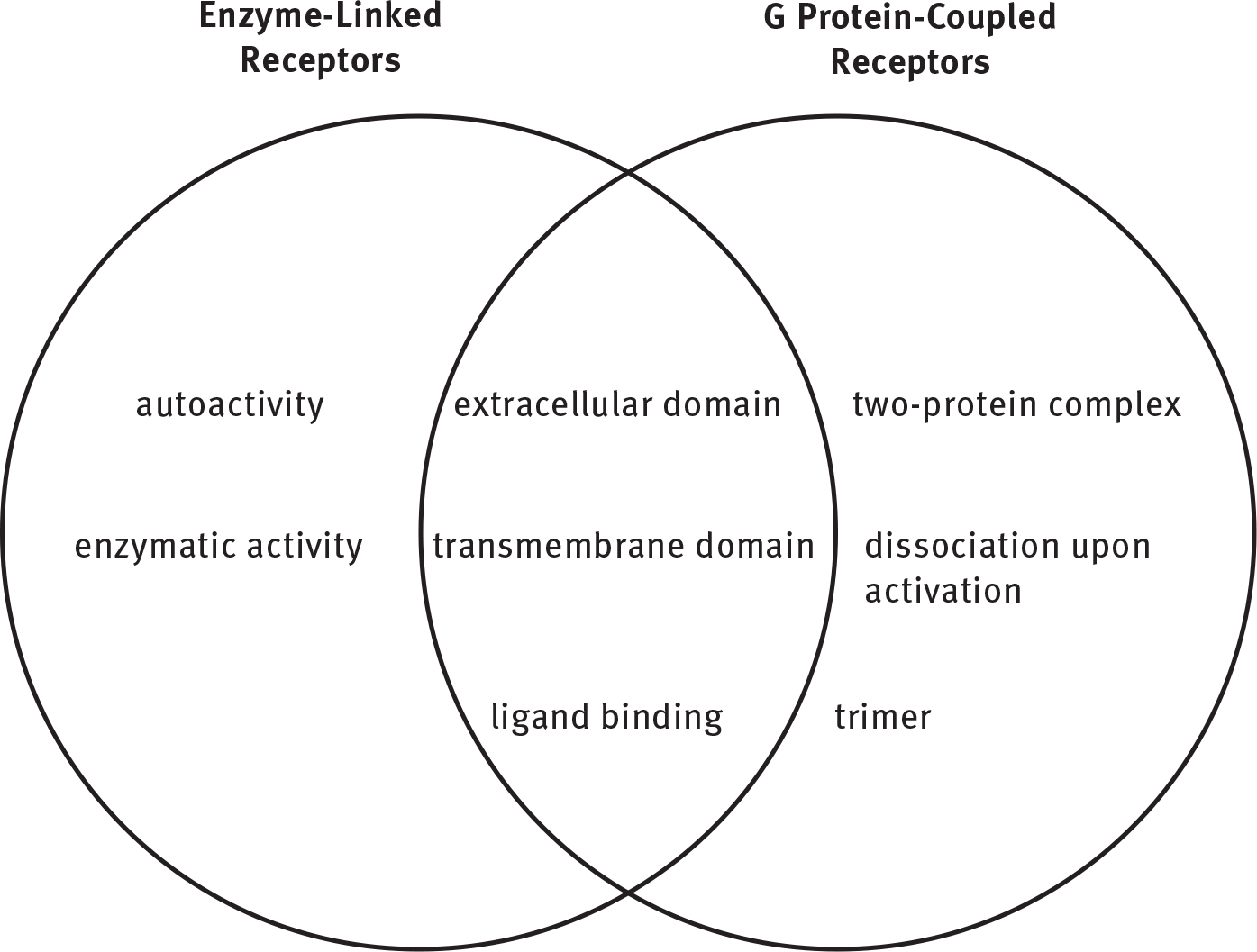
- Ungated channels are always open.
- Transport kinetics display both Km and Vmax values. They also can be cooperative, like some binding proteins. However, transporters do not have analogous Keq values for reactions because there is no catalysis.
**3.3**
- Isoelectric focusing and ion-exchange chromatography both separate proteins based on charge; the charge of a protein in any given environment is determined by its isoelectric point (pI).
- Native PAGE allows a complete protein to be recovered after analysis; it also more accurately determines the relative globular size of proteins. SDS-PAGE can be used to eliminate conflation from mass-to-charge ratios.
- The protein of interest may not elute from the column because its affinity is too high or it may be permanently bound to the free receptor in the eluent.
- True. The small pores in size-exclusion chromatography trap smaller particles, retaining them in the column.
**3.4**
- Protein isolation is generally only the first step in an analysis. The protein identity must be confirmed by amino acid analysis or activity. With unknown proteins, classification of their features is generally desired.
- Contamination of the sample with detergent or SDS could yield an artificially increased protein level, leading to lower activity than expected (because the protein concentration was calculated as higher than its actual value). Alternatively, the enzyme could have been denatured during isolation and analysis.
- False. The Edman degradation proceeds from the amino (N-) terminus.
SCIENCE MASTERY ASSESSMENT EXPLANATIONS
1. D
In most electrophoresis experiments, we attempt to separate out one component from the others. Because we are attempting to isolate protein A only, a pH that causes protein A to be negative while proteins B and C are neutral or positive will be best. pH 5.5 accomplishes this goal; proteins B and C will be positively charged. A pH of 4.5, (C), would make protein A neutral, and it would thus not migrate across the gel. Any neutral impurities would also remain in the well with protein A, making this pH not the best choice.
2. B
Sodium dodecyl sulfate is a detergent and will digest proteins to form micelles with uniform negative charges. Because the protein is sequestered within the micelle, other factors such as charge of the protein and shape have minimal roles during separation. In essence, the protein micelles can be modeled as being spheres, dependent only on size.
3. D
From the given choices, all of them are involved in cell movement with the exception of (D). Centrioles are composed of microtubules, but are involved in mitosis, not cell migration.
4. C
The most prevalent extracellular proteins are keratin, elastin, and collagen. Tubulin and actin are the primary cytoskeletal proteins, while myosin is a motor protein.
5. C
For a ligand present in low quantities to have a strong action, we expect it to initiate a second messenger cascade system. Second messenger systems amplify signals because enzymes can catalyze a reaction more than once while they are active, and often activate other enzymes. Both enzyme-linked receptors and G protein-coupled receptors use second messenger systems, while ion channels do not.
6. D
Ions that are not readily accessible in the cytoplasm or extracellular space are likely to be bound to a binding protein. Classically, calcium and magnesium are protein-bound. Without this background knowledge, the question can still be answered. Sodium, (A), and potassium, (B), must exist in their free states to participate in action potentials. Chloride, (C), is readily excreted by the kidney, which would not be true if it were protein-bound. Calcium must be sequestered in both the bloodstream and intracellularly because calcium is used for muscle contraction, exocytosis (of neurotransmitters and other signals), and many other cellular processes that must be tightly regulated.
7. A
Antibodies are specific to a single antigen. Each B-cell produces a single type of antibody with a constant region that is specific to the host and a variable region that is specific to an antigen.
8. A
The resting membrane potential is displayed by cells that are not actively involved in signal transduction. Ungated or “leak” channels permit limited free flow of ions, while the sodium–potassium pump is also active and corrects for this leakage. Ligand-gated and voltage-gated channels are involved in cell signaling and in the pacemaker potentials of certain cells, but cause deviation from—not maintenance of—the resting membrane potential.
9. D
All trimeric G proteins have α, β, and γsubunits—(A), (B), and (C), respectively. Gs, Gi, and Gq are subtypes of the Gαsubunit of the trimeric G protein and differ depending on the G protein-coupled receptor’s function.
10. B
The proteins described in the question differ primarily in their molecular weights. Their pI values are very close, so ion-exchange chromatography, (A), is not a good choice. The question specifies a large quantity, which is better processed through chromatography than through electrophoresis—(C) and (D)—because the gel can only handle a small volume of protein.
11. C
The overall pI of a protein is determined by the relative number of acidic and basic amino acids. The basic amino acids are arginine, lysine, and histidine, and the acidic amino acids are aspartic acid and glutamic acid. Glycine’s side chain is a hydrogen atom, so it will have the least contribution of all the amino acids.
12. C
The gel in isoelectric focusing uses a pH gradient. When a protein is in a region with a pH above its pI, it is negatively charged and moves toward the anode. When it is in a pH region below its pI, it is positively charged and moves toward the cathode. When the pH equals the pI, the migration of the protein is halted.
13. C
UV spectroscopy is best used with conjugated systems of double bonds. While the double bond in the peptide bond does display resonance, this is not adequate for UV absorption. However, aromatic systems are conjugated, and phenylalanine, tyrosine, and tryptophan all contain aromatic ring structures.
14. B
Protein activity and concentration are generally correlated. Because we have a high concentration of protein, we expect a high activity unless the protein has been damaged or inactivated in some way. The protein could have been inactivated by experimental conditions like detergents, heat, or pH; however, these are not answer choices. Rather, we must consider how the experimental procedure works. Protein elutes off of an affinity column by binding free ligand. In this situation, the binding may not have been reversed and thus the free ligand competes for the active site of the enzyme, lowering its activity.
15. A
The selective cleavage of proteins by digestive enzymes allows fragments of different lengths with known amino acid endpoints to be created. By cleaving the protein with several different enzymes, a basic outline of the amino acid sequence can be created.
GO ONLINE
Consult your online resources for additional practice.
EQUATIONS TO REMEMBER
(3.1) Migration velocity:
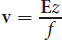
SHARED CONCEPTS
- Biochemistry Chapter 1
- Amino Acids, Peptides, and Proteins
- Biology Chapter 1
- The Cell
- Biology Chapter 8
- The Immune System
- Biology Chapter 11
- The Musculoskeletal System
- General Chemistry Chapter 12
- Electrochemistry
- Organic Chemistry Chapter 12
- Separations and Purifications