Chapter 6: DNA and Biotechnology

Chapter 6: DNA and Biotechnology
SCIENCE MASTERY ASSESSMENT
Every pre-med knows this feeling: there is so much content I have to know for the MCAT! How do I know what to do first or what's important?
While the high-yield badges throughout this book will help you identify the most important topics, this Science Mastery Assessment is another tool in your MCAT prep arsenal. This quiz (which can also be taken in your online resources) and the guidance below will help ensure that you are spending the appropriate amount of time on this chapter based on your personal strengths and weaknesses. Don't worry though— skipping something now does not mean you'll never study it. Later on in your prep, as you complete full-length tests, you'll uncover specific pieces of content that you need to review and can come back to these chapters as appropriate.
How to Use This Assessment
If you answer 0–7 questions correctly:
Spend about 1 hour to read this chapter in full and take limited notes throughout. Follow up by reviewing all quiz questions to ensure that you now understand how to solve each one.
If you answer 8–11 questions correctly:
Spend 20–40 minutes reviewing the quiz questions. Beginning with the questions you missed, read and take notes on the corresponding subchapters. For questions you answered correctly, ensure your thinking matches that of the explanation and you understand why each choice was correct or incorrect.
If you answer 12–15 questions correctly:
Spend less than 20 minutes reviewing all questions from the quiz. If you missed any, then include a quick read-through of the corresponding subchapters, or even just the relevant content within a subchapter, as part of your question review. For questions you got correct, ensure your thinking matches that of the explanation and review the Concept Summary at the end of the chapter.
- In a single strand of a nucleic acid, nucleotides are linked by:
- hydrogen bonds.
- phosphodiester bonds.
- ionic bonds.
- van der Waals forces.
- Which of the following statements regarding differences between DNA and RNA is FALSE?
- In cells, DNA is double-stranded, whereas RNA is single-stranded.
- DNA uses the nitrogenous base thymine; RNA uses uracil.
- The sugar in DNA is deoxyribose; the sugar in RNA is ribose.
- DNA strands replicate in a 5′ to 3′ direction, whereas RNA is synthesized in a 3′ to 5′ direction.
- Which of the following DNA sequences would have the highest melting temperature?
- CGCAACCATCCG
- CGCAATAATACA
- CGTAATAATACA
- CATAACAAATCA
- Which of the following biomolecules is LEAST likely to contain an aromatic ring?
- Proteins
- Purines
- Carbohydrates
- Pyrimidines
- For a compound to be aromatic, all of the following must be true EXCEPT:
- the molecule is cyclic.
- the molecule contains 4n+ 2 πelectrons.
- the molecule contains alternating single and double bonds.
- the molecule is planar.
- Which of the following enzymes is NOT involved in DNA replication?
- Primase
- DNA ligase
- RNA polymerase I
- Telomerase
- How is cDNA best characterized?
- cDNA results from a DNA transcript with noncoding regions removed.
- cDNA results from the reverse transcription of processed mRNA.
- cDNA is the abbreviation for deoxycytosine.
- cDNA is the circular DNA molecule that forms the bacterial genome.
- Which of the following statements regarding polymerase chain reaction is FALSE?
- Human DNA polymerase is used because it is the most accurate.
- A primer must be prepared with a complementary sequence to part of the DNA of interest.
- Repeated heating and cooling cycles allow the enzymes to act specifically and replaces helicase.
- Each cycle of the polymerase chain reaction doubles the amount of DNA of interest.
- Endonucleases are used for which of the following?
- Gene therapy
- Southern blotting
- DNA repair
- I only
- II only
- II and III only
- I, II, and III
- How does prokaryotic DNA differ from eukaryotic DNA?
- Prokaryotic DNA lacks nucleosomes.
- Eukaryotic DNA has telomeres.
- Prokaryotic DNA is replicated by a different DNA polymerase.
- Eukaryotic DNA is circular when not restricted by centromeres.
- I only
- IV only
- II and III only
- I, II, and III only
- Why might uracil be excluded from DNA but NOT RNA?
- Uracil is much more difficult to synthesize than thymine.
- Uracil binds adenine too strongly for replication.
- Cytosine degradation results in uracil.
- Uracil is used as a DNA synthesis activator.
- Tumor suppressor genes are most likely to result in cancer through:
- loss of function mutations.
- gain of function mutations.
- overexpression.
- proto-oncogene formation.
- Which of the following is an ethical concern of gene sequencing?
- Gene sequencing is invasive, thus the potential health risks must be thoroughly explained.
- Gene sequencing impacts relatives, thus privacy concerns may be raised.
- Gene sequencing is very inaccurate, which increases anxiety related to findings.
- Gene sequencing can provide false-negative results, giving a false sense of security.
- Which of the following is NOT a difference between heterochromatin and euchromatin?
- Euchromatin has areas that can be transcribed, whereas heterochromatin is silent.
- Heterochromatin is tightly packed, whereas euchromatin is less dense.
- Heterochromatin stains darkly, whereas euchromatin stains lightly.
- Heterochromatin is found in the nucleus, whereas euchromatin is in the cytoplasm.
- During which phase of the cell cycle are DNA repair mechanisms least active?
- G1
- S
- G2
- M
Answer Key
- B
- D
- A
- C
- C
- C
- B
- A
- D
- D
- C
- A
- B
- D
- D
Chapter 6: DNA and Biotechnology
CHAPTER 6
DNA AND BIOTECHNOLOGY
In This Chapter
6.1 DNA Structure
Nucleosides and Nucleotides
Sugar–Phosphate Backbone
Purines and Pyrimidines
Watson–Crick Model
Denaturation and Reannealing
6.2 Eukaryotic Chromosome Organization
Histones
Heterochromatin and Euchromatin
Telomeres and Centromeres
6.3 DNA Replication
Strand Separation and Origins of Replication
Synthesis of Daughter Strands
Replicating the Ends of Chromosomes
6.4 DNA Repair
Oncogenes and Tumor Suppressor Genes
Proofreading and Mismatch Repair
Nucleotide and Base Excision Repair
6.5 Recombinant Biotechnology
Molecular Cloning and Restriction Enzymes
DNA Libraries and cDNA
Hybridization
Blotting and Molecular Labeling
DNA Sequencing
Applications of DNA Technology
Safety and Ethics
Concept Summary
CHAPTER PROFILE
The content in this chapter should be relevant to about 14% of all questions about biochemistry on the MCAT.
This chapter covers material from the following AAMC content categories:
1B: Transmission of genetic information from the gene to the protein
5D: Structure, function, and reactivity of biologically-relevant molecules
Introduction
How do all of the cells of our body know what job to do? The cells that make up the human body have nucleic acids that instruct the cell on how to function. The nucleic acid *deoxyribonucleic** acid* (DNA) stores the information in our cells and selectively shares that information when appropriate. DNA is a molecule that can be passed from generation to generation. DNA can be replicated in a carefully regulated process designed to keep the genome safe from degradation and free from errors. Seemingly small changes in the genetic code can result in life-threatening and even life-incompatible alterations to protein structure and function. In this chapter, the unique structure of DNA will be discussed, along with replication and repair processes. The primary focus will be on eukaryotic processes, but there will be some review of prokaryotic genetics to help us better understand the molecular basis of life.
Much of the advancement of medicine in the past two decades has been due to our increased understanding of molecular genetics, which has led to the creation of an entire biotechnology industry centered around genomics and the utilization of nucleic acids for various diagnostic tests and therapeutic interventions. We will also take a look at some of these important principles in this chapter.
6.1 DNA Structure
LEARNING OBJECTIVES
After Chapter 6.1, you will be able to:
- Identify the structures of, and distinguish the differences between, nucleotides and nucleosides
- Recognize the key features and rules of purine and pyrimidine structure and pairing
- Recall the structural differences between DNA and RNA molecules
There are two chemically distinct forms of nucleic acids within eukaryotic cells. Deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) are polymers, each with distinct roles, that together create the molecules integral to life in all living organisms. DNA is the focus of this chapter and RNA will be discussed in more detail in Chapter 7 of MCAT Biochemistry Review. The bulk of DNA is found in chromosomes in the nucleus of eukaryotic cells, although some is also present in mitochondria and chloroplasts.
Nucleosides and Nucleotides
DNA is a macromolecule and it is essential to understand how this molecule is constructed. DNA is a polydeoxyribonucleotide that is composed of many monodeoxyribonucleotides linked together. The nomenclature of nucleic acids can be complicated, so the terms have been defined here:
- Nucleosides are composed of a five-carbon sugar (pentose) bonded to a nitrogenous base and are formed by covalently linking the base to C-1′ of the sugar, as shown in Figure 6.1. Note that the carbon atoms in the sugar are labeled with a prime symbol to distinguish them from the carbon atoms in the nitrogenous base.
- Nucleotides are formed when one or more phosphate groups are attached to C-5′ of a nucleoside. Often these molecules are named according to the number of phosphates present. Adenosine di- and triphosphate (ADP and ATP), for example, gain their names from the number of phosphate groups attached to the nucleoside adenosine. These are high-energy compounds because of the energy associated with the repulsion between closely associated negative charges on the phosphate groups, as shown in Figure 6.2. Nucleotides are the building blocks of DNA.
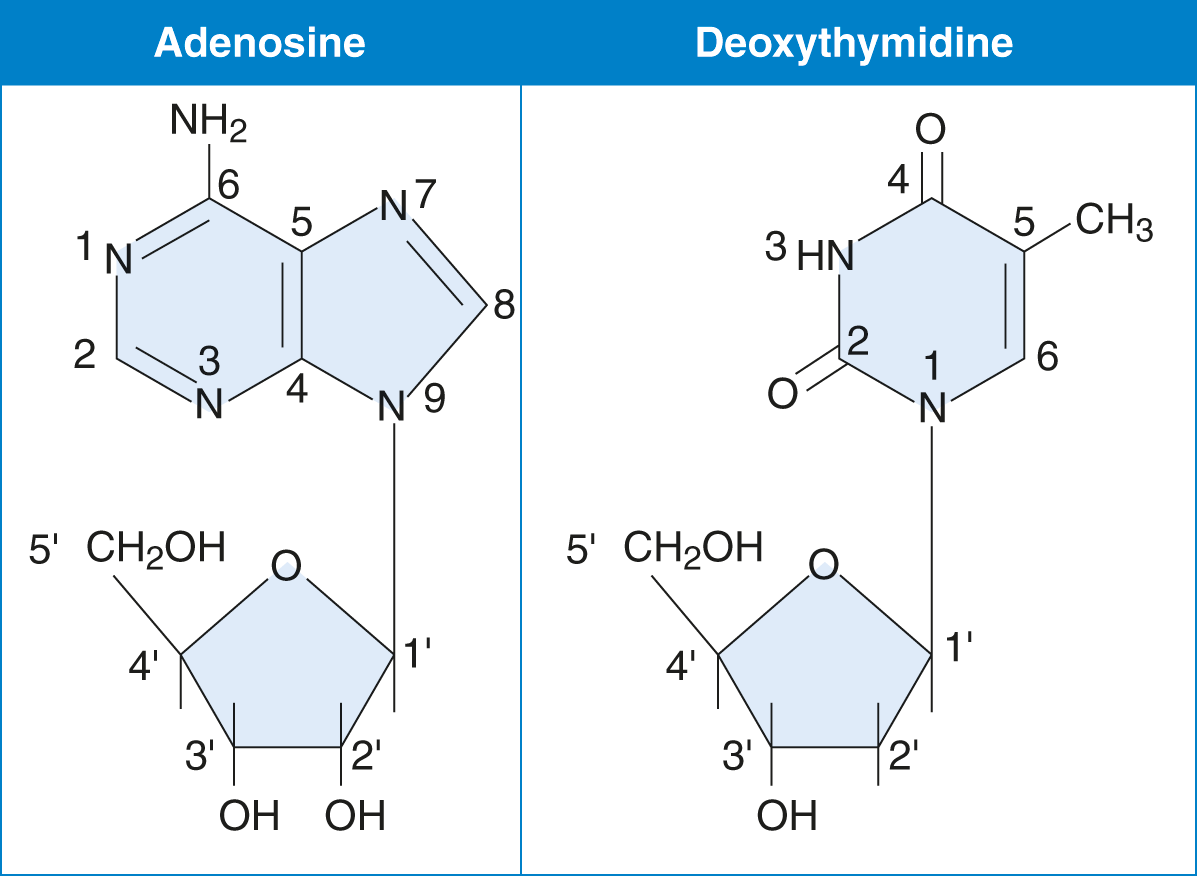
Figure 6.1 Examples of Nucleosides
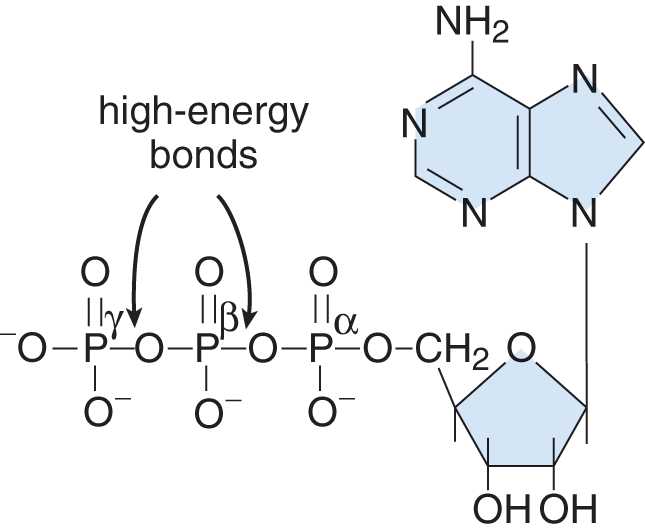
Figure 6.2 High-Energy Bonds in Adenosine Triphosphate, a Nucleotide
BRIDGE
In Chapter 3 of MCAT General Chemistry Review, we learned that bond breaking is usually endothermic and bond making is usually exothermic. ATP offers a biologically relevant—and MCAT tested—exception to this rule. Due to all the negative charges in close proximity, removing the terminal phosphate from ATP actually releases energy, which powers our cells.
Nucleic acids are classified according to the pentose they contain, as shown in Figure 6.3. If the pentose is ribose, the nucleic acid is RNA; if the pentose is deoxyribose (ribose with the 2′ –OH group replaced by –H), then it is DNA.
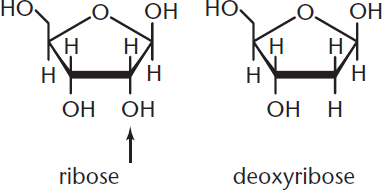
Figure 6.3 Ribose and Deoxyribose Ribose has an –OH group at C-2; deoxyribose has an –H.
The nomenclature for the common bases, nucleosides, and nucleotides is shown in Table 6.1.
Table 6.1. Nomenclature of Important Bases, Nucleosides, and Nucleotides
BASE NUCLEOSIDE NUCLEOTIDES
Adenine Adenosine (Deoxyadenosine) AMP (dAMP) ADP (dADP) ATP (dATP)
Guanine Guanosine (Deoxyguanosine) GMP (dGMP) GDP (dGDP) GTP (dGTP)
Cytosine Cytidine (Deoxycytidine) CMP (dCMP) CDP (dCDP) CTP (dCTP)
Uracil Uridine (Deoxyuridine) UMP (dUMP) UDP (dUDP) UTP (dUTP)
Thymine (Deoxythymidine) (dTMP) (dTDP) (dTTP)
Names of nucleosides and nucleotides attached to deoxyribose are shown in parentheses.
Note that there is no thymidine listed (only deoxythymidine) because thymine appears almost exclusively in DNA. A structural comparison of nucleosides and nucleotides is illustrated in Figure 6.4.
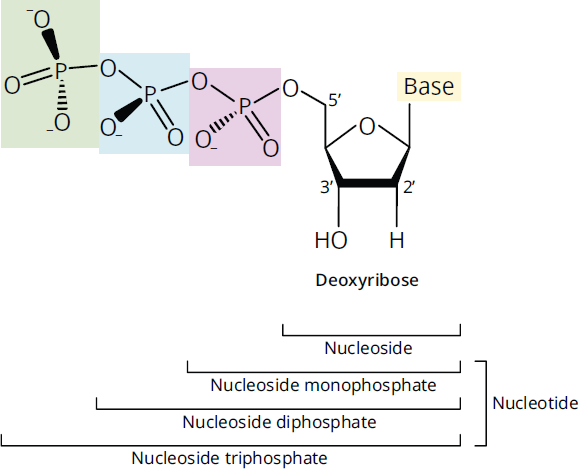
Figure 6.4 Structure of Nucleosides and Nucleotides
Sugar–Phosphate Backbone
The backbone of DNA is composed of alternating sugar and phosphate groups; it determines the directionality of the DNA and is always read from 5′ to 3′. It is formed as nucleotides are joined by 3′–5′ phosphodiester bonds. That is, a phosphate group links the 3′ carbon of one sugar to the 5′ phosphate group of the next incoming sugar in the chain. Phosphates carry a negative charge; thus, DNA and RNA strands have an overall negative charge.
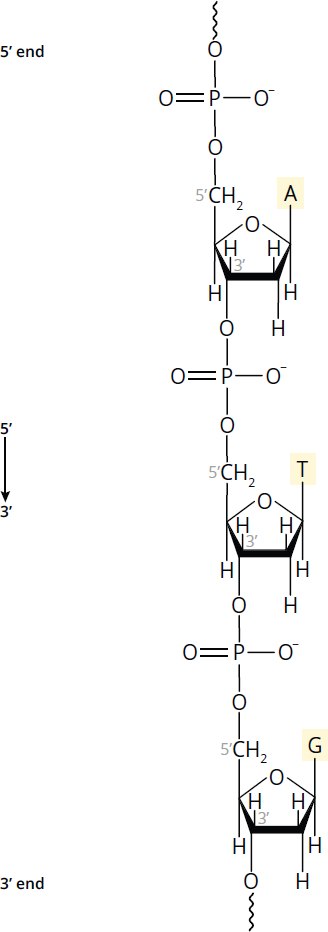
Figure 6.5 DNA Strand Polarity DNA strands run antiparallel to one another; enzymes that replicate and transcribe DNA only work in the 5′ to 3′ direction.
Each strand of DNA has distinct 5′ and 3′ ends, creating polarity within the backbone, as shown in Figure 6.5. The 5′ end of DNA, for instance, will have an –OH or phosphate group bonded to C-5′ of the sugar, while the 3′ end has a free –OH on C-3′ of the sugar. The base sequence of a nucleic acid strand is both written and read in the 5′ to 3′ direction. Thus, the DNA strand in Figure 6.5 must be written: 5′—ATG—3′ (or simply ATG). DNA sequences can also be written in slightly different ways:
- If written backwards, the ends must be labeled: 3′—GTA—5′
- The position of phosphates may be shown: pApTpG
- “d” may be used as shorthand for deoxyribose: dAdTdG
DNA is generally double-stranded (dsDNA) and RNA is generally single-stranded (ssRNA). Exceptions to this rule may be seen, especially in viruses, as described in Chapter 1 of MCAT Biology Review.
MCAT EXPERTISE
On the MCAT, always check nucleic acids for polarity. One of the easiest ways to generate incorrect answers is to simply reverse the reading frame: 3′—GATTACA—5′ is not the same as 3′—ACATTAG—5′.
Purines and Pyrimidines
There are two families of nitrogen-containing bases found in nucleotides: purines and pyrimidines. The bases described below, and shown in Figure 6.6, represent the common bases in eukaryotes; however, it should be noted that exceptions may be seen in tRNA and in some prokaryotes and viruses. Purines contain two rings in their structure. The two purines found in nucleic acids are adenine (A) and guanine (G); both are found in DNA and RNA. Pyrimidines contain only one ring in their structure. The three pyrimidines are cytosine (C), thymine (T), and uracil (U); while cytosine is found in both DNA and RNA, thymine is only found in DNA and uracil is only found in RNA.
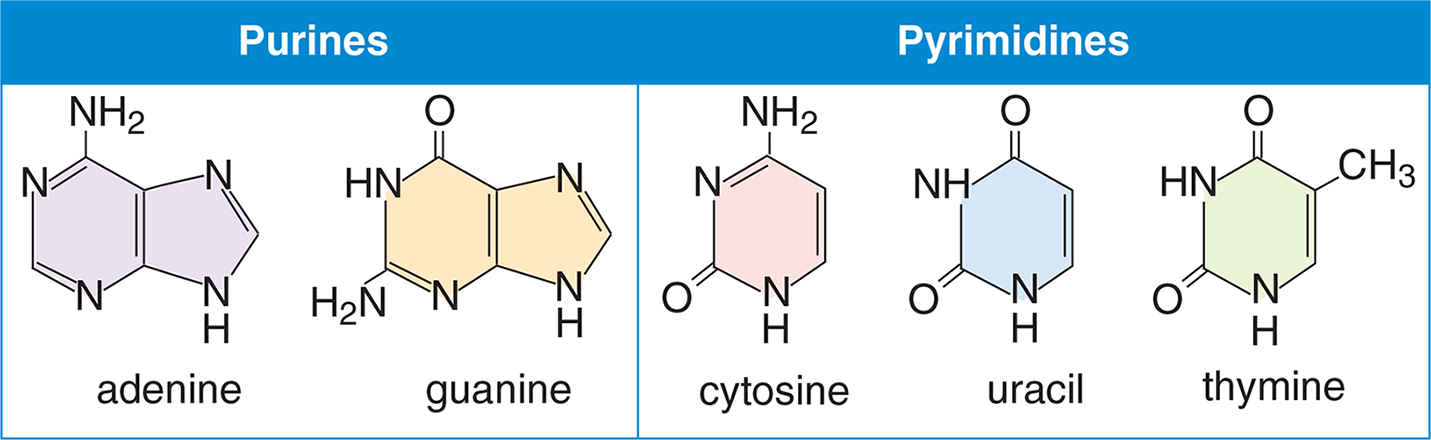
Figure 6.6 Bases Commonly Found in Nucleic Acids
MNEMONIC
To remember the types and structures of these two classes of nitrogenous bases, remember to CUT the PYe (as C, U, and T are pyrimidines). You can also note that pie has one ring of crust, and pyrimidines have only one ring in their structure. You can also remember PURe As Gold (as A and G are purines); think of gold wedding rings. It takes two gold rings at a wedding, just like purines have two rings in their structure.
Purines and pyrimidines are examples of biological aromatic heterocycles. In chemistry, the term aromatic describes any unusually stable ring system that adheres to the following four specific rules:
- The compound is cyclic.
- The compound is planar.
- The compound is conjugated (has alternating single and multiple bonds, or lone pairs, creating at least one unhybridized p-orbital for each atom in the ring).
- The compound has 4n + 2 (where n is any integer) π electrons. This is called Hückel’s rule.
The most common example of an aromatic compound is benzene, but many different structures obey these rules. In an aromatic compound, the extra stability is due to the delocalized π electrons, which can travel throughout the entire compound using available molecular orbitals. All six of the carbon atoms in benzene are sp2-hybridized, and each of the six orbitals overlaps equally with its two neighbors. As a result, the delocalized electrons form two π electron clouds (one above and one below the plane of the ring), as shown in Figure 6.7. This delocalization is characteristic of all aromatic molecules, and because of this, aromatic molecules are fairly unreactive.
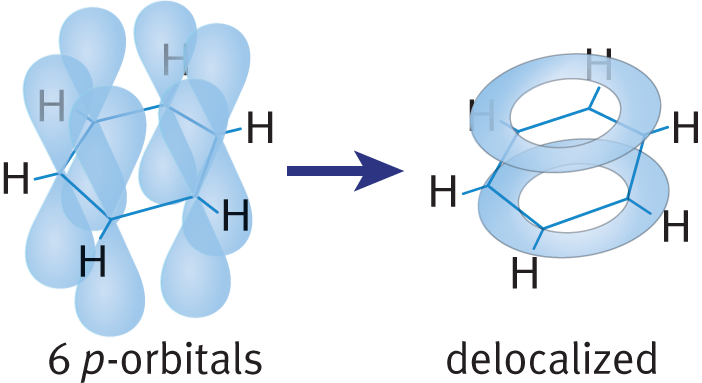
Figure 6.7 Delocalization of π Electrons in Benzene
Heterocycles are ring structures that contain at least two different elements in the ring. As shown in Figure 6.6 earlier, both purines and pyrimidines contain nitrogen in their aromatic rings. Nucleic acids are thus imbued with exceptional stability. This helps to explain the utility of nucleotides as the molecule for storing genetic information. A furan is a heterocyclic compound that is a five-membered aromatic ring containing one oxygen atom and four carbon atoms. Compounds that contain these rings are referred to as furans. Pyridine is another type of heterocyclic compound, but has the chemical formula C5H5N. The molecule is similar in structure to benzene, with one of the carbons in the ring substituted with a nitrogen atom instead.
Nucleotides can also undergo pi stacking, which is a type of non-covalent attractive interaction between two aromatic rings. The most commonly observed types of pi stacking, which are shown in Figure 6.8, are staggered stacking (also called parallel displaced) and pi-teeing (also called T-shaped). In staggered stacking, the aromatic rings are parallel to one another but are not directly overlapping, with one aromatic ring slightly offset from the other. In this formation, carbon atoms that have a partial negative charge are placed above hydrogen atoms with a partial positive charge, leading to an attractive electrostatic interaction. In pi-teeing, one aromatic ring is perpendicular to the other aromatic ring to form a T shape, which similarly aligns partially-negative carbon atoms with partially-positive hydrogen atoms.
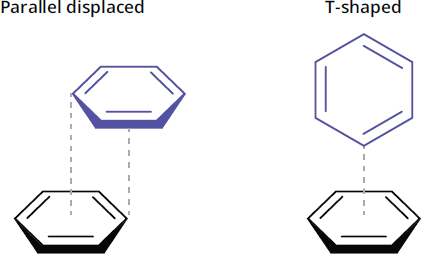
Figure 6.8 Two Common Types of Pi Stacking Between Aromatic Rings
The staggered formation is an important contributor of nucleobase stacking within DNA and RNA polynucleotide strands, which increases structural stability. Staggered stacking and pi-teeing formations are also commonly found between amino acids with aromatic side chains, which, along with other types of molecular interactions, can help drive protein folding.
Watson–Crick Model
Putting this information together, we can start looking at the Watson–Crick model of DNA structure. In 1953, James Watson and Francis Crick presented one of the landmark findings of modern biology and medicine: the three-dimensional structure of DNA. With X-ray patterns of DNA produced by Rosalind Franklin laying the necessary foundation for their work, Watson and Crick were able to deduce the double-helical nature of DNA and propose specific base-pairing that would be the basis of a copying mechanism. In the double helix, two linear polynucleotide chains of DNA are wound together in a spiral orientation along a common axis. The key features of the model—some of which have already been mentioned—are:
- The two strands of DNA are antiparallel; that is, the strands are oriented in opposite directions. When one strand has polarity 5′ to 3′ down the page, the other strand has 5′ to 3′ polarity up the page.
- The sugar–phosphate backbone is on the outside of the helix with the nitrogenous bases on the inside.
- There are specific base-pairing rules, often referred to as complementary base-pairing, as shown in Figure 6.9. An adenine (A) is always base-paired with a thymine (T) via two hydrogen bonds. A guanine (G) always pairs with cytosine (C) via three hydrogen bonds. The three hydrogen bonds make the G–C base pair interaction stronger. These hydrogen bonds, and the hydrophobic interactions between bases, provide stability to the double helix structure. Thus, the base sequence on one strand defines the base sequence on the other strand.
- Because of the specific base-pairing, the amount of A equals the amount of T, and the amount of G equals the amount of C. Thus, total purines will be equal to total pyrimidines overall. These properties are known as Chargaff’s rules.
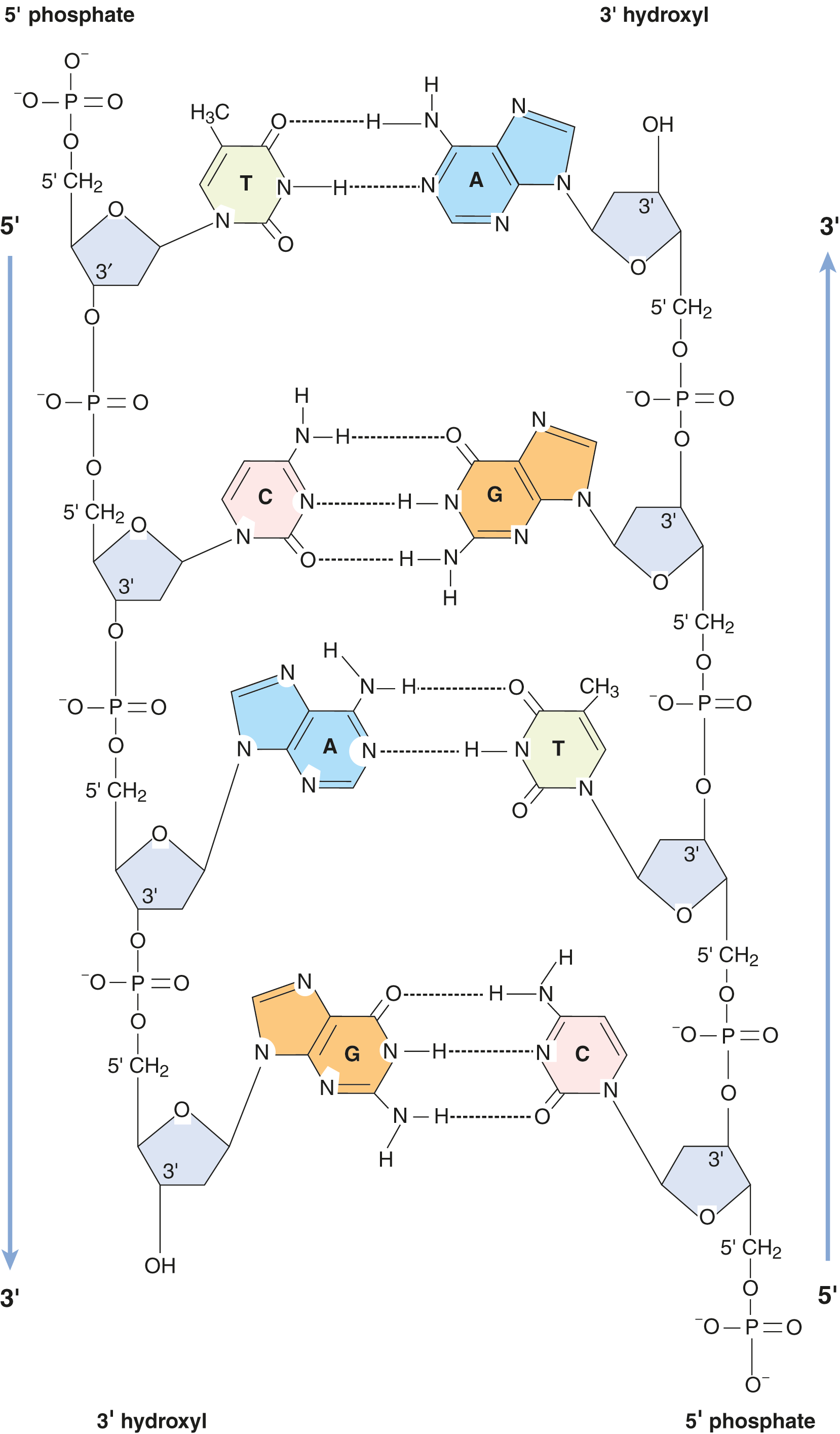
Figure 6.9 Base-Pairing in DNA
KEY CONCEPT
When writing a complementary strand of DNA, it is important to not only remember the base-pairing rules but to also keep track of the 5′ and 3′ ends. Remember that the sequences need to be both complementary and antiparallel. For example, 5′—ATCG—3′ will be complementary to 5′—CGAT—3′.
MCAT EXPERTISE
Using Chargaff’s rules:
In double-stranded DNA, purines = pyrimidines:
- %A = %T
- %G = %C
If a sample of DNA has 10% G, what is the % of T?
10% G = 10% C, thus %G + %C = 20% %A + %T = 80%, thus %T = 40%.
The double helix of most DNA is a right-handed helix, forming what is called B-DNA, as shown in Figure 6.10. The helix in B-DNA makes a turn every 3.4 nm and contains about 10 bases within that span. Major and minor grooves can be identified between the interlocking strands and are often the site of protein binding. Another form of DNA is called Z-DNA for its zigzag appearance; it is a left-handed helix that has a turn every 4.6 nm and contains 12 bases within each turn. A high GC-content or a high salt concentration may contribute to the formation of this form of DNA. No biological activity has been attributed to Z-DNA partly because it is unstable and difficult to research.
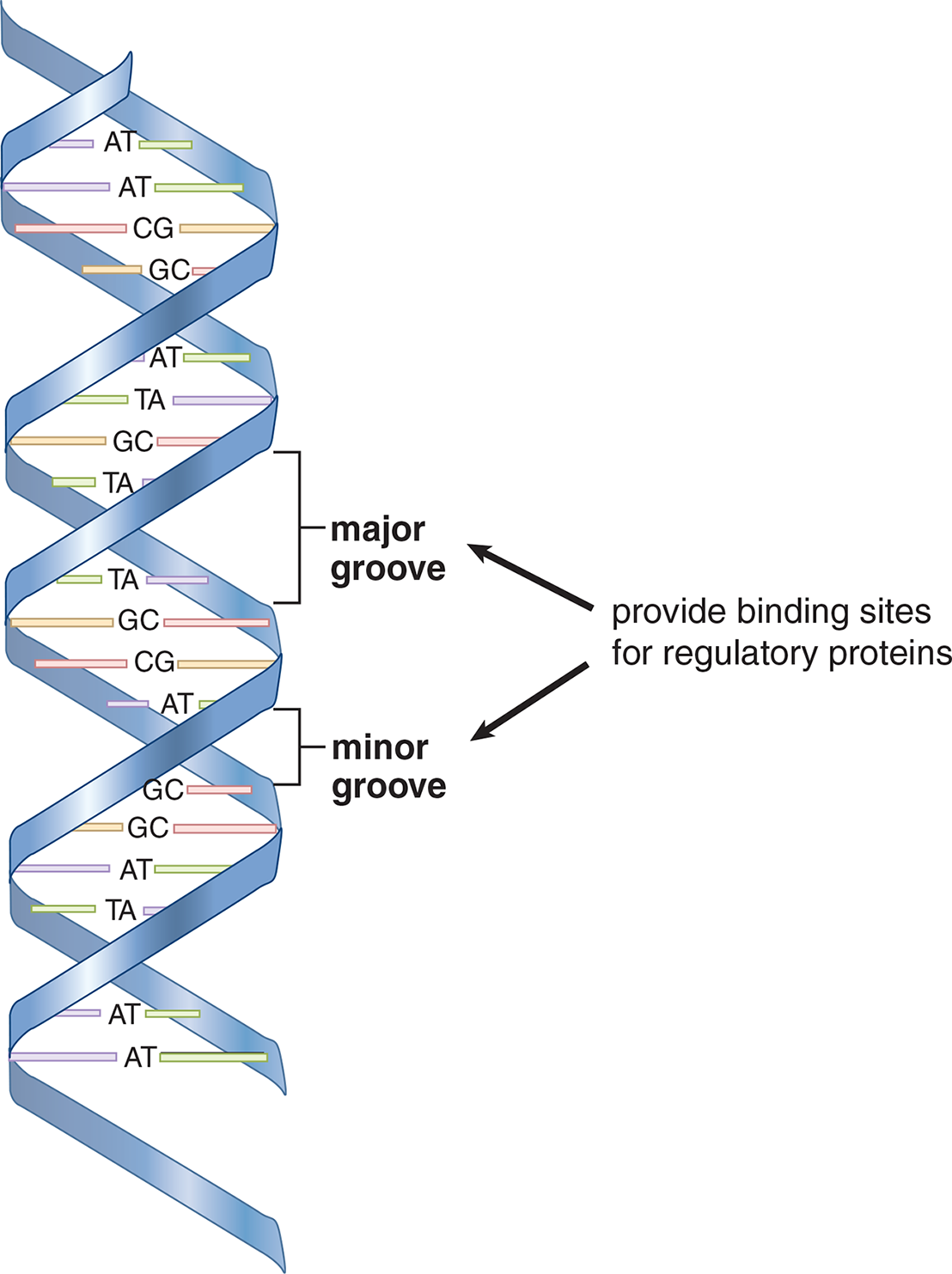
Figure 6.10 The B-DNA Double Helix
Denaturation and Reannealing
During processes such as replication and transcription, it is necessary to gain access to the DNA. The double helical nature of DNA can be denatured by conditions that disrupt hydrogen bonding and base-pairing, resulting in the “melting” of the double helix into two single strands that have separated from each other. Notably, none of the covalent links between the nucleotides in the backbone of the DNA break during this process. Heat, alkaline pH, and chemicals like formaldehyde and urea are commonly used to denature DNA.
Denatured, single-stranded DNA can be reannealed (brought back together) if the denaturing condition is slowly removed. If a solution of heat-denatured DNA is slowly cooled, for example, then the two complementary strands can become paired again, as shown in Figure 6.11.
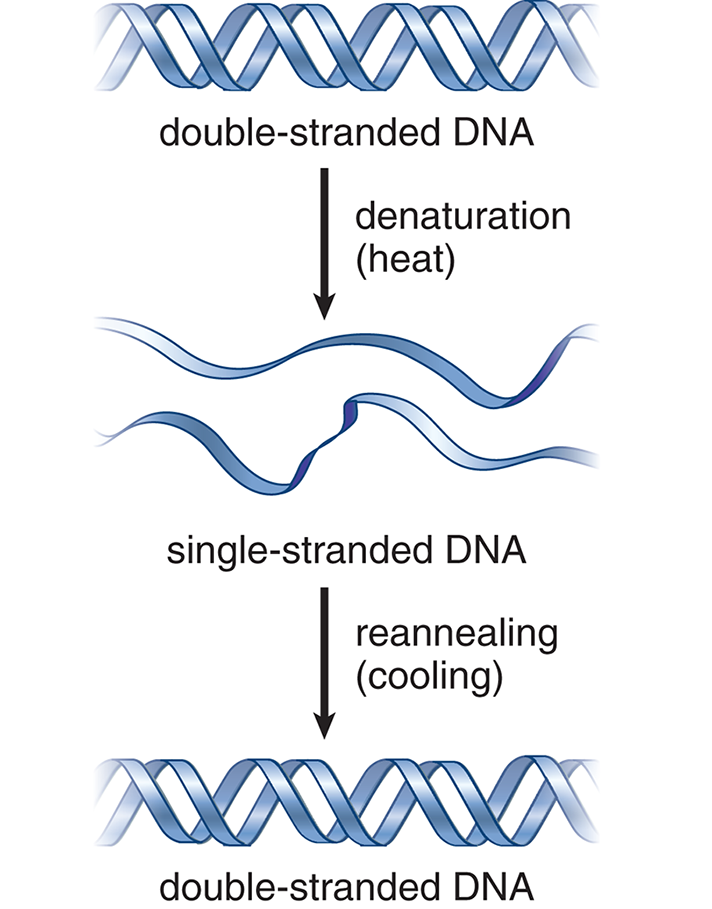
Figure 6.11 Denaturation and Reannealing of DNA
Such annealing of complementary DNA strands is also an important step in many laboratory processes, such as polymerase chain reactions (PCR) and in the detection of specific DNA sequences. In these techniques, a well-characterized probe DNA (DNA with known sequence) is added to a mixture of target DNA sequences. When probe DNA binds to target DNA sequences, this may provide evidence of the presence of a gene of interest. This binding process is called hybridization and is described in further detail later in this chapter.
MCAT CONCEPT CHECK 6.1:
Before you move on, assess your understanding of the material with these questions.
- What is the difference between a nucleoside and a nucleotide?
_________________________________________
- What are the base-pairing rules according to the Watson–Crick model?
_________________________________________
- What are the three major structural differences between DNA and RNA?
- ____________________________________
- _____________________________________
- _____________________________________
- How does the aromaticity of purines and pyrimidines underscore their genetic function?
_________________________________________
-
If a strand of RNA contained 15% cytosine, 15% adenine, 35% guanine, and 35% uracil, would this violate Chargaff’s rules? Why or why not?
________________________________________
6.2 Eukaryotic Chromosome Organization
LEARNING OBJECTIVES
After Chapter 6.2, you will be able to:
- Recall the names and the role of the five histone proteins
- Differentiate between the major characteristics of heterochromatin and euchromatin
- Describe the traits of telomeres and centromeres
There are over 6 billion bases of DNA in each human cell. It is important for the cell to organize these bases effectively. These bases must be replicated during the cell cycle and also utilized in gene expression for normal cellular functions. In humans, DNA is divided up among the 46 chromosomes found in the nucleus of the cell. The supercoiling of the DNA double helix does provide some compaction, but much more is necessary.
Histones
The DNA that makes up a chromosome is wound around a group of small lysine- and arginine-rich proteins called histones, forming chromatin. The positive charge on the sidechain of lysine and arginine residues interact favorably with the negatively charged phosphate backbone of DNA. There are five histone proteins found in eukaryotic cells. Two copies each of the histone proteins H2A, H2B, H3, and H4 form a histone core and about 200 base pairs of DNA are wrapped around this protein complex, forming a nucleosome, as shown in Figure 6.12. Under an electron microscope, the nucleosomes look like beads on a string. The last histone, H1, seals off the DNA as it enters and leaves the nucleosome, adding stability to the structure. Together, the nucleosomes create a much more organized and compacted DNA.
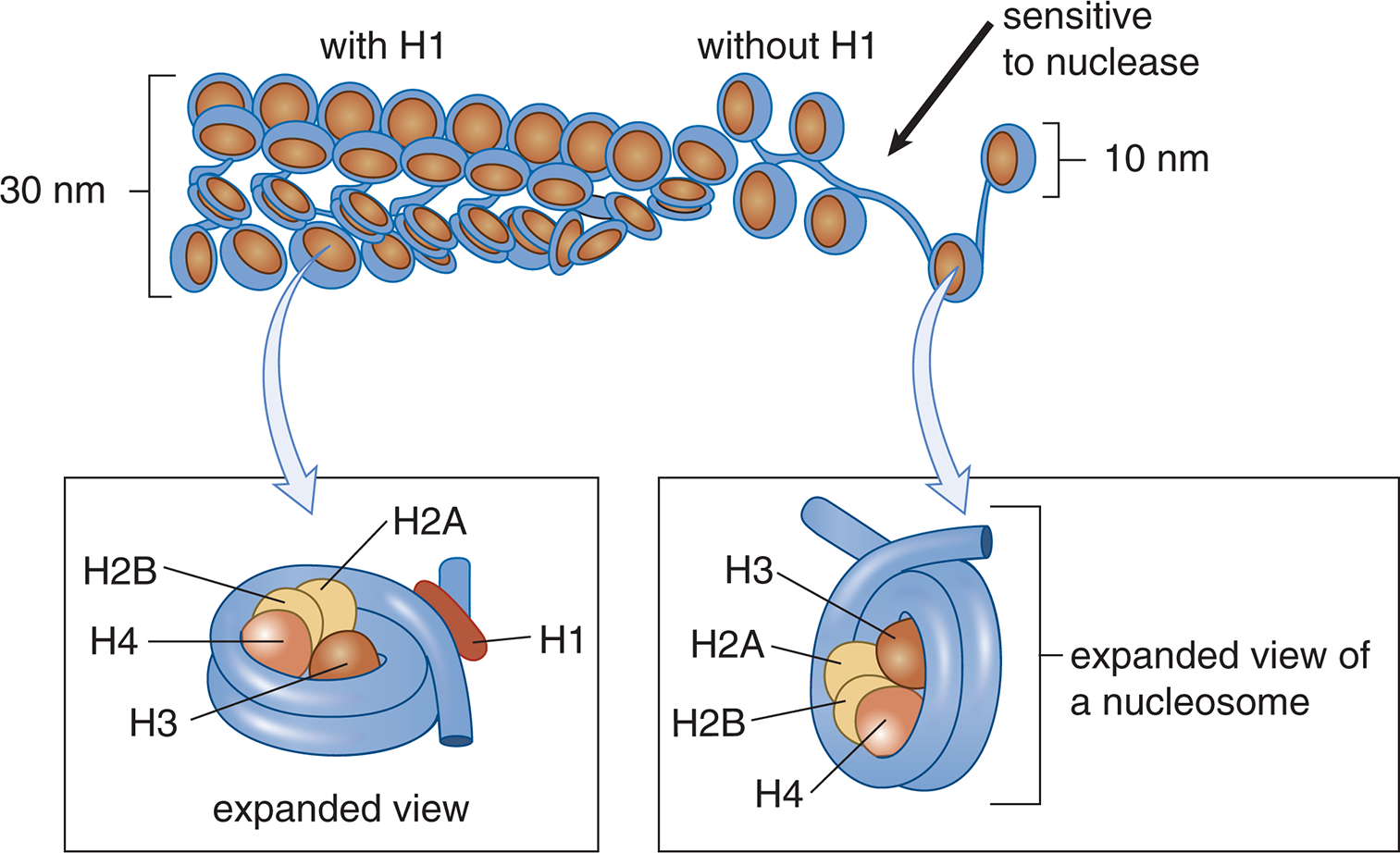
Figure 6.12 Nucleosome Structure Nucleosomes are composed of DNA wrapped around histone proteins.
Histones are one example of nucleoproteins (proteins that associate with DNA). Most other nucleoproteins are acid-soluble and tend to stimulate processes such as transcription. Histone acetyltransferases are enzymes that can modify lysine residues on histones by masking the positive charge on the sidechain of lysine residues through the addition of an acetyl group. As illustrated in Figure 6.13, this loss of the positive charge on the lysine side chain, which typically interacts with the negatively-charged DNA wound around the histones, leads to the release of the DNA due to the weakened interaction. Post-translational modifications by histone acetyltransferases are often associated with transcriptional activation, whereas histone deacetylases (HDACs) usually repress the transcription of specific genes. Similarly, histone methyltransferases are enzymes that add methyl groups to lysine and arginine residues. Because methylation preserves the positive charges on lysine and arginine, histone methylation tends to be associated with decreased transcription.
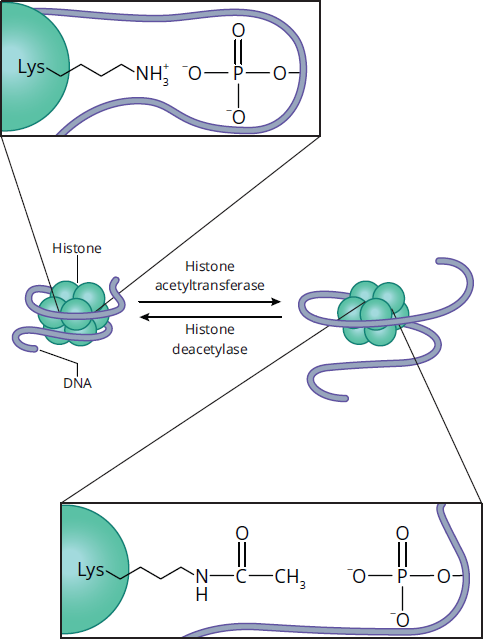
Figure 6.13 Histone Acetylation and Deacetylation Acetylation removes the positive charge on the lysine side chain, loosening the interaction between histones and DNA.
Heterochromatin and Euchromatin
The chromosomes have a diffuse configuration during interphase of the cell cycle. The cell will undergo DNA replication during the S phase of interphase and having the DNA uncondensed and accessible makes the process more efficient. A small percentage of the chromatin remains compacted during interphase and is referred to as heterochromatin. Heterochromatin appears dark under light microscopy and is transcriptionally silent. Heterochromatin often consists of DNA with highly repetitive sequences. In contrast, the dispersed chromatin is called euchromatin, which appears light under light microscopy. Euchromatin contains genetically active DNA. Both heterochromatin and euchromatin can be seen in the nucleus in Figure 6.14.

Figure 6.14 Euchromatin and Heterochromatin in an Interphase Nucleus
KEY CONCEPT
Heterochromatin is dark, dense, and silent. Euchromatin is light, uncondensed, and expressed.
Telomeres and Centromeres
As described later in this chapter, DNA replication cannot extend all the way to the end of a chromosome. This will result in losing sequences and information with each round of replication. The solution for our cells is a simple repeating unit (TTAGGG) at the end of the DNA, forming a telomere. Some of the sequence is lost in each round of replication and can be replaced by the enzyme telomerase. Telomerase is more highly expressed in rapidly dividing cells. Animal studies indicate that there are a set number of replications possible, and that the progressive shortening of telomeres contributes to aging. Telomeres also serve a second function: their high GC-content creates exceptionally strong strand attractions at the end of chromosomes to prevent unraveling; think of telomeres as “knotting off” the end of the chromosome.
Centromeres, as their name suggests, are a region of DNA found in the center of chromosomes. They are often referred to as sites of constriction because they form noticeable indentations. This part of the chromosome is composed of heterochromatin, which is in turn composed of tandem repeat sequences that also contain high GC-content. During cell division, the two sister chromatids can therefore remain connected at the centromere until microtubules separate the chromatids during anaphase.
BIOCHEMISTRY GUIDED EXAMPLE WITH EXPERT THINKING
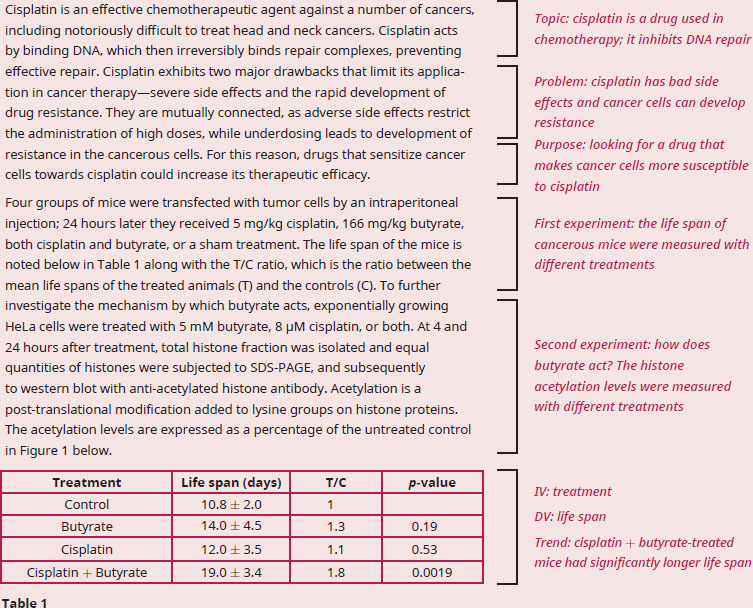
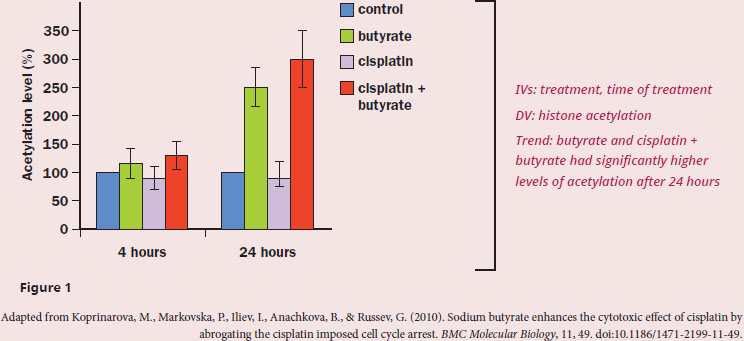
Given the fact that histones are enriched with surface-exposed basic amino acids, how does butyrate enhance the cytotoxic effect of cisplatin on tumor cells?
The question stem asks us to explain the action of butyrate in conjunction with cisplatin, which means we need to dig into the experiment and results of the passage. The first paragraph tells us that cisplatin blocks DNA repair, and when targeting cancerous cells, cisplatin would likely cause cell death. Table 1 clearly shows that cisplatin alone doesn’t increase the life span of mice with cancer, but coupling cisplatin with butyrate does. However, this result doesn’t tell us how this paired treatment works (just that it does work), which means that we need to find this information in Figure 1. The treatments do not differ from the control after 4 hours, but butyrate and cisplatin + butyrate do increase the levels of histone acetylation. The passage tells us that histone acetylation is a posttranslational addition of an acetyl group on to lysines. We know from the structure of amino acids that lysines are positively charged, so adding an acetyl group would mask the positive charge. We’re also told that histones have many surface-exposed basic amino acids, and basic amino acids (like lysine and arginine) are positively charged at physiological pH. Let’s also recall that when DNA is wrapped tightly around histone proteins, it is referred to as heterochromatin, which means the DNA is transcriptionally silent. Since DNA has a negative charge due to the phosphodiester backbone, acetylating the lysines on histones will remove some of the positive charge. Without that positive charge binding the negatively charged DNA, the histones will no longer be held tightly to the DNA, meaning they will dissociate from the DNA. DNA that is not tightly bound to histones forms euchromatin, which is genetically active: those genes can now be transcribed and translated. The DNA will then also be accessible to cisplatin, which will bind and stop DNA repair complexes. A build up of unrepaired DNA often causes cells to die, usually through apoptotic mechanisms.
In summary, butyrate causes DNA to separate from histones, allowing for cisplatin to bind and stop DNA repair complexes from fixing errors in the DNA, leading to eventual cell death.
MCAT CONCEPT CHECK 6.2:
Before you move on, assess your understanding of the material with these questions.
- What are the five histone proteins in eukaryotic cells? Which one is not part of the histone core around which DNA wraps to form chromatin?
________________________________________
- Compare and contrast heterochromatin and euchromatin based on the following characteristics:
Characteristic Heterochromatin Euchromatin Density of chromatin packing Appearance under light microscopy Transcriptional activity
- What property of telomeres and centromeres allows them to stay tightly raveled, even when the rest of DNA is uncondensed?
_________________________________________
6.3 DNA Replication
LEARNING OBJECTIVES
After Chapter 6.3, you will be able to:
- List the names and functions of the major enzymes of DNA synthesis in prokaryotes and eukaryotes
- Differentiate between synthesis of the leading and lagging strands
- Explain the role of telomerase and the function of the telomere
The DNA is an organism’s “blueprint” that provides not only the ability to sustain activities of life but also insight into our evolutionary past. The process of DNA replication is highly regulated to ensure as close to 100 percent perfection when making a copy of our genome as possible. DNA replication is necessary for the reproduction of a species and for any dividing cell.
Strand Separation and Origins of Replication
The human genome has about 3 billion base pairs packed into multiple chromosomes. The replisome or replication complex is a set of specialized proteins that assist the DNA polymerases. To begin the process of replication, DNA unwinds at points called origins of replication. The generation of new DNA proceeds in both directions, creating replication forks on both sides of the origin, as shown in Figure 6.15.
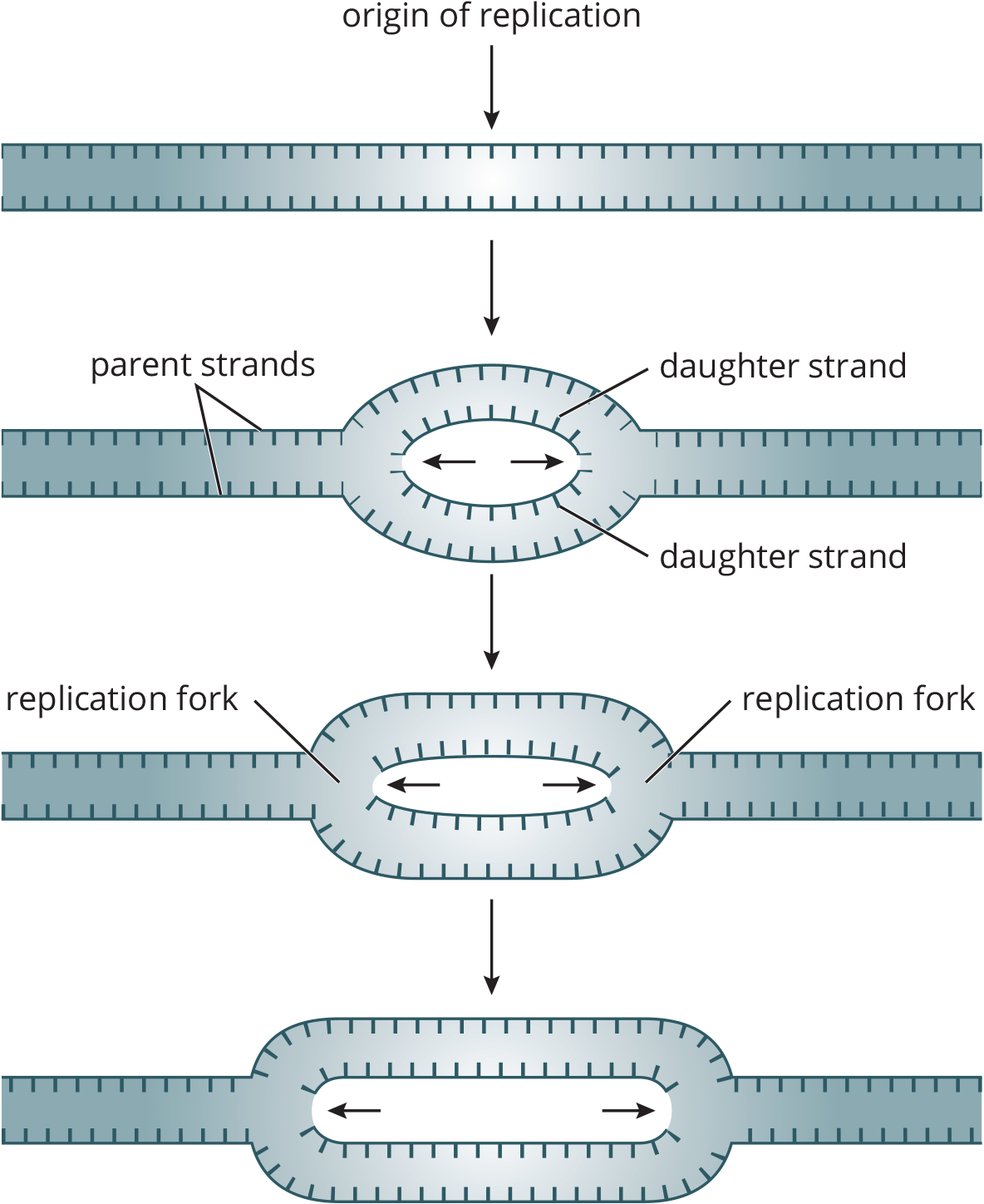
Figure 6.15 Origins of Replication Replication forks form on both sides of the origin, increasing the efficiency of replication.
The bacterial chromosome is a closed, double-stranded circular DNA molecule with a single origin of replication. Thus, there are two replication forks that move away from each other in opposite directions around the circle. The two replication forks eventually meet, resulting in the production of two identical circular molecules of DNA.
Eukaryotic replication must copy many more bases compared to prokaryotic and is a slower process. In order to duplicate all of the chromosomes efficiently, each eukaryotic chromosome contains one linear molecule of double-stranded DNA having multiple origins of replication. As the replication forks move toward each other and sister chromatids are created, the chromatids will remain connected at the centromere. The differences between prokaryotic and eukaryotic replication patterns are shown in Figure 6.16.
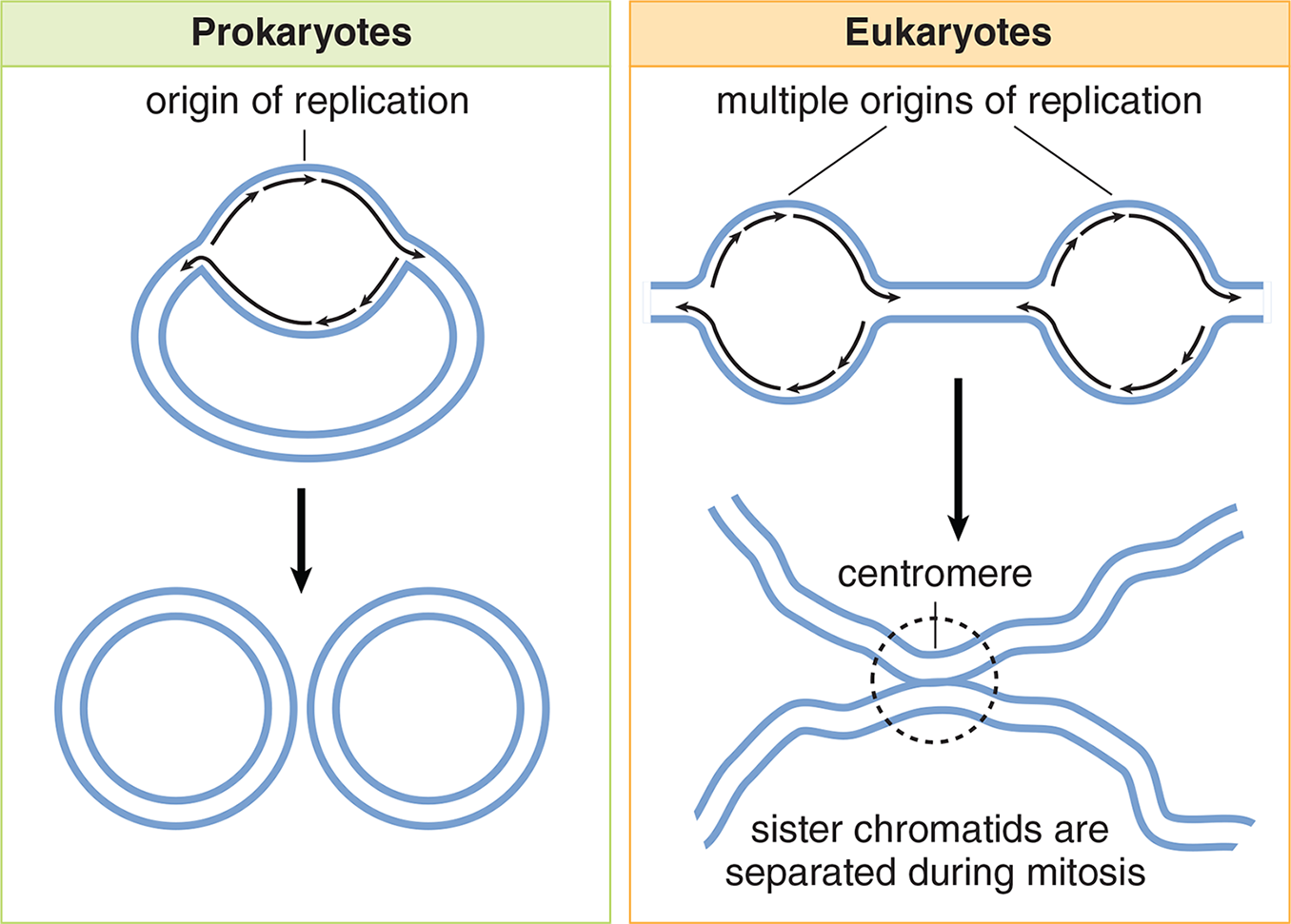
Figure 6.16 DNA Replication in Prokaryotes and Eukaryotes
Helicase is the enzyme responsible for unwinding the DNA, generating two single-stranded template strands ahead of the polymerase. Once opened, the unpaired strands of DNA are very sticky, in a molecular sense. The free purines and pyrimidines seek out other molecules with which to hydrogen bond. Proteins are therefore required to hold the strands apart: single-stranded DNA-binding proteins will bind to the unraveled strand, preventing both the reassociation of the DNA strands and the degradation of DNA by nucleases. As the helicase unwinds the DNA, it will cause positive supercoiling that strains the DNA helix. Supercoiling is a wrapping of DNA on itself as its helical structure is pushed ever further toward the telomeres during replication; picture an old-fashioned telephone cord that’s become tangled on itself. To alleviate this torsional stress and reduce the risk of strand breakage, DNA topoisomerases introduce negative supercoils. They do so by working ahead of helicase, nicking one or both strands, allowing relaxation of the torsional pressure, and then resealing the cut strands.
REAL WORLD
Any antibiotic ending with –floxacin (ciprofloxacin, moxifloxacin, and so on) is called a fluoroquinolone and targets the prokaryotic topoisomerase DNA gyrase. This stops bacterial replication and slows the infection. Fluoroquinolones are routinely used for pneumonia and certain infections of the genitourinary system.
During replication, these parental strands will serve as templates for the generation of new daughter strands. The replication process is termed semiconservative because one parental strand is retained in each of the two resulting identical double-stranded DNA molecules, as shown in Figure 6.17.
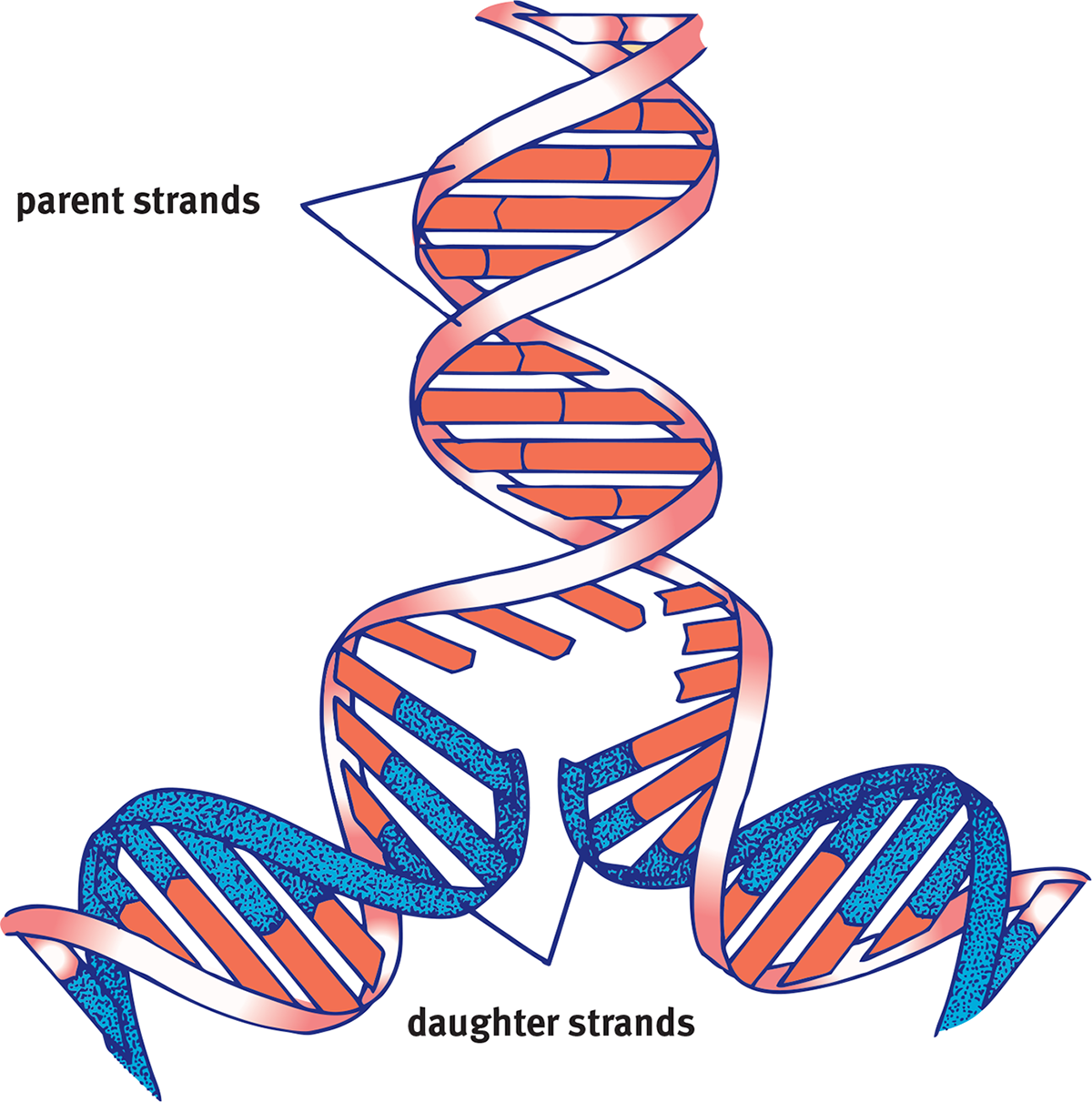
Figure 6.17 Semiconservative Replication A new double helix is made of one old parent strand and one new daughter strand.
Synthesis of Daughter Strands
DNA polymerases are responsible for reading the DNA template, or parental strand, and synthesizing the new daughter strand. The DNA polymerase can read the template strand in a 3′ to 5′ direction while synthesizing the complementary strand in the 5′ to 3′ direction. This will result in a new double helix of DNA that has the required antiparallel orientation. Due to this directionality of the DNA polymerase, certain constraints arise. Remember that the two separated parental strands of the helix are also antiparallel to each other. Thus, at each replication fork, one strand is oriented in the correct direction for DNA polymerase; the other strand is antiparallel.
KEY CONCEPT
With the exception of DNA polymerase’s reading direction (and a few untested endonucleases), everything in molecular biology is 5′ to 3′. DNA polymerase reads 3′ to 5′, but the following processes occur 5′ to 3′:
- DNA synthesis
- DNA repair
- RNA transcription
- RNA translation (reading of codons)
The leading strand in each replication fork is the strand that is copied in a continuous fashion, in the same direction as the advancing replication fork. This parental strand will be read 3′ to 5′ and its complement will be synthesized in a 5′ to 3′ manner, as discussed above.
The lagging strand is the strand that is copied in a direction opposite the direction of the replication fork. On this side of the replication fork, the parental strand has 5′ to 3′ polarity. DNA polymerase cannot simply read and synthesize on this strand. How does it solve this problem? Because DNA polymerase can only synthesize in the 5′ to 3′ direction from a 3′ to 5′ template, small strands called Okazaki fragments are produced. As the replication fork continues to move forward, it clears additional space that DNA polymerase must fill in. Each time DNA polymerase completes an Okazaki fragment, it turns around to find another gap that needs to be filled in.
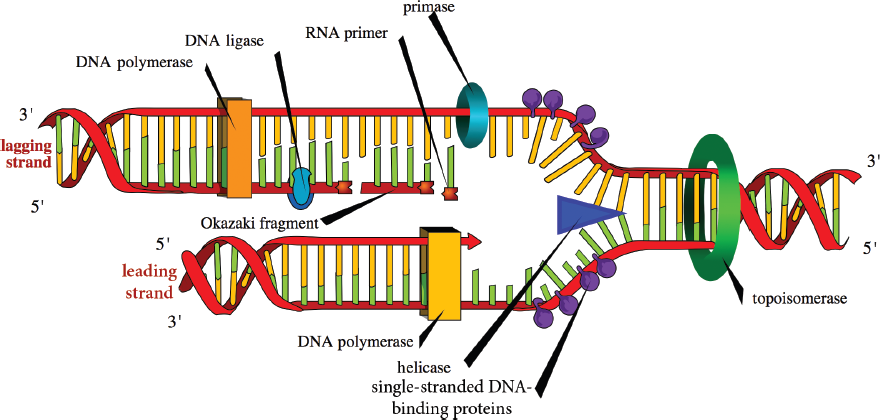
Figure 6.18 Enzymes of DNA Replication This process involves the action of DNA helicase, gyrase, polymerase, and ligase to create two identical molecules of DNA.
The process of DNA replication is shown in Figure 6.18. The first step in the replication of DNA is actually to lay down an RNA primer. DNA cannot be synthesized de novo; that is, it needs another molecule to “hook on” to. RNA, on the other hand, can be directly paired with the parent strand. Thus, primase synthesizes a short primer (roughly 10 nucleotides) in the 5′ to 3′ direction to start replication on each strand. These short RNA sequences are constantly being added to the lagging strand because each Okazaki fragment must start with a new primer. In contrast, the leading strand requires only one, in theory (in reality, there are usually a few primers on the leading strand). DNA polymerase III (prokaryotes) or DNA polymerases α, δ, and ε (eukaryotes) will then begin synthesizing the daughter strands of DNA in the 5′ to 3′ manner. The incoming nucleotides are 5′ deoxyribonucleotide triphosphates: dATP, dCTP, dGTP, and dTTP. As the new phosphodiester bond is made, a free pyrophosphate (PPi) is released.
The RNA must eventually be removed to maintain integrity of the genome. This is accomplished by the enzyme DNA polymerase I (prokaryotes) or RNase H (eukaryotes). Then, DNA polymerase I (prokaryotes) or DNA polymerase δ (eukaryotes) adds DNA nucleotides where the RNA primer had been. DNA ligase seals the ends of the DNA molecules together, creating one continuous strand of DNA.
While prokaryotic DNA synthesis has been worked out in great detail, and eukaryotic synthesis is considered to be very similar, there are a few differences in the enzymes involved. There are five “classic” DNA polymerases in eukaryotic cells, which are designated with the Greek letters α, β, γ, δ, and ε. Further research has revealed more polymerases named ζ through μ, but these are outside the scope of the MCAT. Table 6.2 highlights differences in the names of the enzymes associated with DNA replication in prokaryotic vs. eukaryotic cells; special attention should be paid to the eukaryotic DNA polymerases:
- DNA polymerases α, δ, and ε work together to synthesize both the leading and lagging strands; DNA polymerase δ also fills in the gaps left behind when RNA primers are removed
- DNA polymerase γ replicates mitochondrial DNA
- DNA polymerases β and ε are important to the process of DNA repair
- DNA polymerases δ and ε are assisted by the PCNAprotein, which assembles into a trimer to form the sliding clamp. The clamp helps to strengthen the interaction between these DNA polymerases and the template strand
Table 6.2. Steps and Proteins Involved in DNA Replication
STEP IN REPLICATION PROKARYOTIC CELLS EUKARYOTIC CELLS (NUCLEI)
Origin of replication One per chromosome Multiple per chromosome
Unwinding of DNA double helix Helicase Helicase
Stabilization of unwound template strands Single-stranded DNA-binding protein Single-stranded DNA-binding protein
Synthesis of RNA primers Primase Primase
Synthesis of DNA DNA polymerase III DNA polymerases **α,δ, andε Removal of RNA primers** DNA polymerase I (5′ → 3′ exonuclease) RNase H (5′ → 3′ exonuclease)
Replacement of RNA with DNA DNA polymerase I DNA polymerase δ
Joining of Okazaki fragments DNA ligase DNA ligase
Removal of positive supercoils ahead of advancing replication forks DNA topoisomerases (DNA gyrase) DNA topoisomerases
Synthesis of telomeres Not applicable Telomerase
Replicating the Ends of Chromosomes
While DNA polymerase does an excellent job of synthesizing DNA, it unfortunately cannot complete synthesis of the 5′ end of the strand. Thus, each time DNA synthesis is carried out, the chromosome becomes a little shorter. To lengthen the time that cells can replicate and synthesize DNA before necessary genes are damaged, chromosomes contain telomeres. As described earlier, telomeres are located at the very tips of the chromosome and consist of repetitive sequences with a high GC-content. This repetition means that telomeres can be slightly degraded between replication cycles without loss of function.
MCAT CONCEPT CHECK 6.3:
Before you move on, assess your understanding of the material with these questions.
- For each of the enzymes listed below, list the function of the enzyme and if it is found in prokaryotes, eukaryotes, or both.
**Enzyme Prokaryotes/ Eukaryotes/Both Function Helicase Single-stranded DNA-binding protein Primase DNA polymerase III DNA polymerase α DNA polymerase I RNase H DNA ligase DNA topoisomerases**
- Between the leading strand and lagging strand, which is more prone to mutations? Why?
_________________________________________
- What is the function of a telomere?
______________________________
6.4 DNA Repair
LEARNING OBJECTIVES
After Chapter 6.4, you will be able to:
- Describe the major DNA repair processes, including proofreading, mismatch repair, and excision repair
- Recognize the key components and locations of each DNA repair process, including both mismatch and excision processes
- Identify major traits of oncogenes and tumor suppressor genes
The structure of DNA can be damaged in a number of ways such as exposure to chemicals or radiation. DNA is very susceptible to damage and if the damage is not corrected, it will subsequently be copied and passed on to daughter cells. Damage can include breaking of the DNA backbone, structural or spontaneous alterations of bases, or incorporation of the incorrect base during replication. Any defect in the genetic code can cause an increased risk of cancer, so the cell has multiple processes in place to catch and correct genetic errors. This helps maintain the integrity and stability of the genome from cell to cell, and from generation to generation.
Oncogenes and Tumor Suppressor Genes
Certain genes, when mutated, can lead to cancer. Cancer cells proliferate excessively because they are able to divide without stimulation from other cells and are no longer subject to the normal controls on cell proliferation. By definition, cancer cells are able to migrate by local invasion or metastasis, a migration to distant tissues by the bloodstream or lymphatic system. Over time, cancer cells tend to accumulate mutations.
Mutated genes that cause cancer are termed oncogenes. Oncogenes primarily encode cell cycle–related proteins. Before these genes are mutated, they are often referred to as proto-oncogenes. The first gene in this category to be discovered was src (named after sarcoma, a category of connective tissue cancers). The abnormal alleles encode proteins that are more active than normal proteins, promoting rapid cell cycle advancement. Typically, a mutation in only one copy is sufficient to promote tumor growth and is therefore considered dominant.
KEY CONCEPT
While the outcome of oncogenes and mutated tumor suppressor genes is the same (cancer), the actual cause is different. Oncogenes promote the cell cycle while mutated tumor suppressors can no longer slow the cell cycle. Oncogenes are like stepping on the gas pedal; mutated tumor suppressors are like cutting the brakes.
Tumor suppressor genes, like p53 or Rb (retinoblastoma), encode proteins that inhibit the cell cycle or participate in DNA repair processes. They normally function to stop tumor progression, and are sometimes called antioncogenes. Mutations of these genes result in the loss of tumor suppression activity, and therefore promote cancer. Inactivation of both alleles is necessary for the loss of function because, in most cases, even one copy of the normal protein can function to inhibit tumor formation. In this example, multiple mutations or “hits” are required.
Proofreading and Mismatch Repair
DNA polymerase moves along a single strand of DNA, building the complementary strand as it goes. While DNA polymerase is almost 100 percent accurate, it does occasionally make errors.
Proofreading
During synthesis, the two double-stranded DNA molecules will pass through a part of the DNA polymerase enzyme for proofreading. When the complementary strands have incorrectly paired bases, the hydrogen bonds between the strands can be unstable, and this lack of stability is detected as the DNA passes through this part of the polymerase. The incorrect base is excised and can be replaced with the correct one, as shown in Figure 6.19. If both the parent and daughter strands are simply DNA, how does the enzyme discriminate which is the template strand, and which is the incorrectly paired daughter strand? It looks at the level of methylation: the template strand has existed in the cell for a longer period of time, and therefore is more heavily methylated. Methylation also plays a role in the transcriptional activity of DNA, as described in Chapter 7 of MCAT Biochemistry Review. This system is very efficient, correcting most of the errors put into the sequence during replication. DNA ligase, which closes the gaps between Okazaki fragments, lacks proofreading ability. Thus, the likelihood of mutations in the lagging strand is considerably higher than the leading strand.
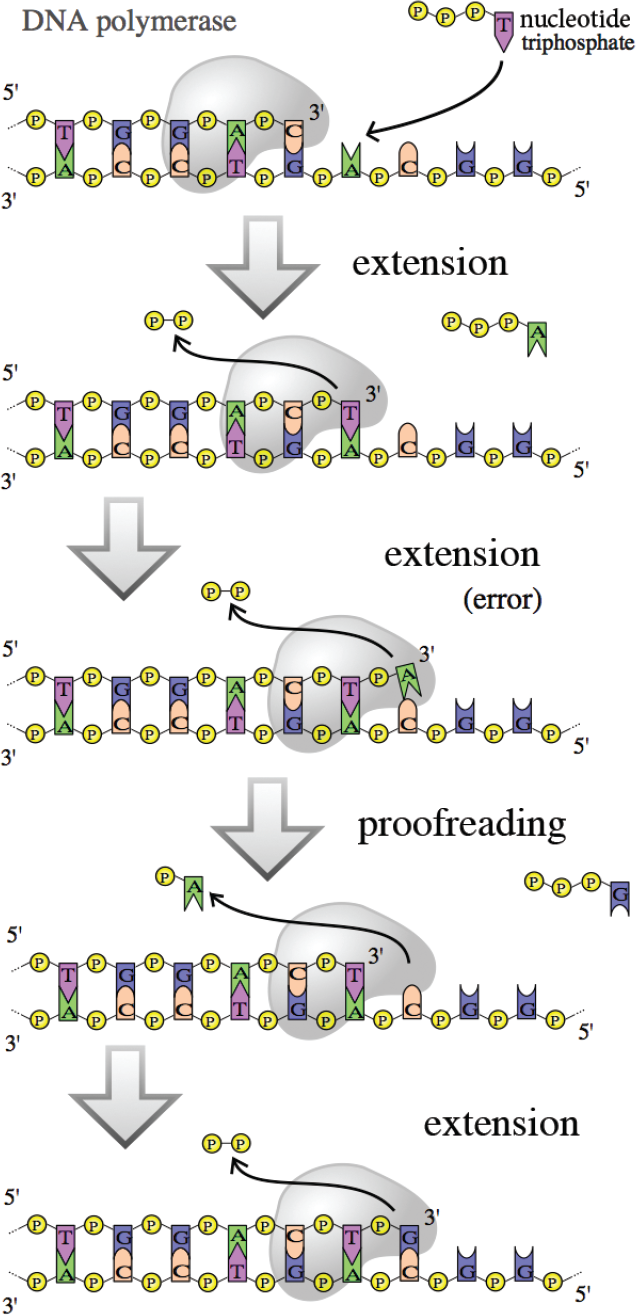
Figure 6.19 Proofreading by DNA Polymerase
Mismatch Repair
Cells also have machinery in the G2 phase of the cell cycle for mismatch repair; these enzymes are encoded by genes MSH2 and MLH1, which detect and remove errors introduced in replication that were missed during the S phase of the cell cycle. These enzymes are homologues of MutS and MutL in prokaryotes, which serve a similar function.
Nucleotide and Base Excision Repair
Most of the repair mechanisms involve proteins that recognize damage or a lesion, remove the damage, and then use the complementary strand as a template to fill in the gap. Our cell machinery recognizes two specific types of DNA damage in the G1 and G2 cell cycle phases and fixes them through nucleotide excision repair or base excision repair.
Nucleotide Excision Repair
Ultraviolet light induces the formation of dimers between adjacent thymine residues in DNA. The formation of thymine dimers interferes with DNA replication and normal gene expression, and distorts the shape of the double helix. Thymine dimers are eliminated from DNA by a nucleotide excision repair (NER) mechanism, which is a cut-and-patch process, as shown in Figure 6.20. First, specific proteins scan the DNA molecule and recognize the lesion because of a bulge in the strand. An excision endonuclease then makes nicks in the phosphodiester backbone of the damaged strand on both sides of the thymine dimer and removes the defective oligonucleotide. DNA polymerase can then fill in the gap by synthesizing DNA in the 5′ to 3′ direction, using the undamaged strand as a template. Finally, the nick in the strand is sealed by DNA ligase.
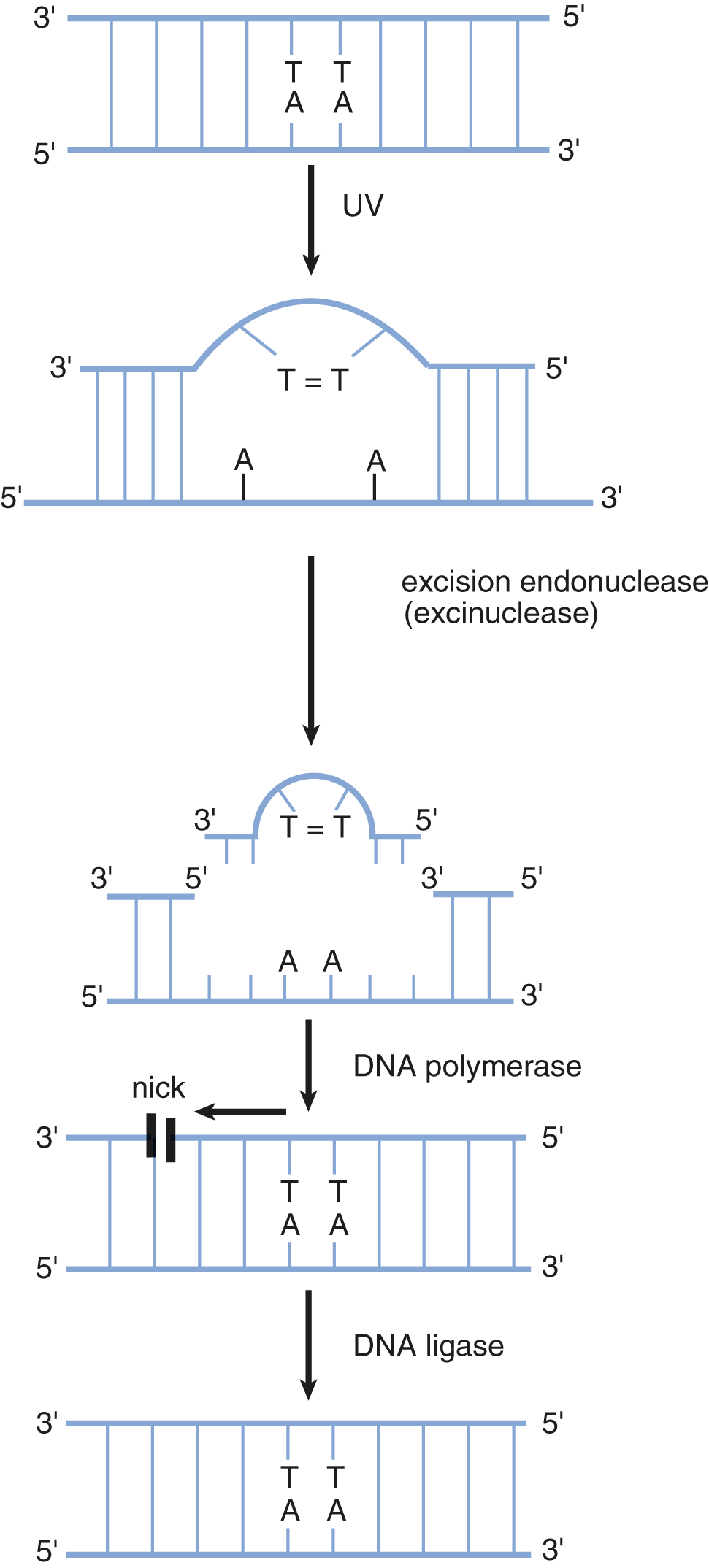
Figure 6.20 Thymine Dimer Formation and Nucleotide Excision Repair
Base Excision Repair
Alterations to bases can occur with other cellular insults. For example, thermal energy can be absorbed by DNA and may lead to cytosine deamination. This is the loss of an amino group from cytosine and results in the conversion of cytosine to uracil. Uracil should not be found in a DNA molecule and is thus easily detected as an error; however, detection systems exist for small, non-helix-distorting mutations in other bases as well. These are repaired by base excision repair. First, the affected base is recognized and removed by a glycosylase enzyme, leaving behind an apurinic/apyrimidinic (AP) site, also called an abasic site. The AP site is recognized by an AP endonuclease that removes the damaged sequence from the DNA. DNA polymerase and DNA ligase can then fill in the gap and seal the strand, as described above.
MCAT CONCEPT CHECK 6.4:
Before you move on, assess your understanding of the material with these questions.
- What is the difference between an oncogene and a tumor suppressor gene?
_________________________________
- How does DNA polymerase recognize which strand is the template strand once the daughter strand is synthesized?
_________________________________
- For each of the repair mechanisms below, in which phase of the cell cycle does the repair mechanism function? What are the key enzymes or genes specifically associated with each mechanism?
Repair Mechanism Phase of Cell Cycle Key Enzymes/Genes DNA polymerase (proofreading) Mismatch repair Nucleotide excision repair Base excision repair
- What is the key structural difference in the types of lesions corrected by nucleotide excision repair vs. those corrected by base excision repair?
_______________________________________
6.5 Recombinant Biotechnology
LEARNING OBJECTIVES
After Chapter 6.5, you will be able to:
- Predict the most effective DNA library technique for a given laboratory application
- Recall the inputs and outputs of biotechnology techniques, including PCR, Southern blotting, and sequencing
- Describe the differences between transgenic mice and knockout mice
Now that we have reviewed the basics of DNA structure and function, we can discuss how this knowledge has been harnessed for a variety of research and treatment innovations. Recombinant DNA technology allows for the creation of recombinant DNA, which is DNA that is created from combining pieces of DNA from different sources together to make one new molecule, and then multiplication of the recombinant sequence through either gene cloning or polymerase chain reaction (PCR). This provides a means of analyzing and altering genes and proteins. It also provides the reagents necessary for genetic testing, such as carrier detection (detecting heterozygote status for a particular disease) and prenatal diagnosis of genetic diseases; it is also useful for gene therapy. Additionally, this technology can provide a source of a specific protein, such as recombinant human insulin, in almost unlimited quantities.
Molecular Cloning and Restriction Enzymes
Molecular cloning is a technique that is used to create recombinant DNA and produce large amounts of it using host organisms. Cloning conventionally starts with the desired sequence of DNA being inserted into a vector backbone to create a recombinant vector. The recombinant vector is then transferred to a host bacterium that is then allowed to grow into colonies. Colonies that contain the recombinant vector are isolated and grown in large quantities. Depending on the investigator’s goal and experimental setup, the bacteria can then be lysed to purify the recombinant vectors or any proteins expressed from the recombinant sequence.
DNA vector backbones, often referred to as simply vectors, are particles used to carry foreign DNA sequences. The most commonly used (and most likely to be tested) type of vector on the MCAT are plasmids, which are small circular DNA molecules. Plasmids that are used as vectors tend to have specific sequences that make them ideal for molecular cloning, including a promoter, origin of replication, multiple cloning site (MCS) (also known as a polylinker), and selectable marker, which are described in further detail below.
Restriction enzymes (restriction endonucleases) are used in molecular cloning that recognize specific double-stranded DNA sequences called restriction sites. These sequences are palindromic, meaning that the 5′ to 3′ sequence of one strand is identical to the 5′ to 3′ sequence of the other strand (in antiparallel orientation). Restriction enzymes are isolated from bacteria, which are their natural source. In bacteria, they act as part of the restriction modification system that protects the bacteria from infection by DNA viruses. Once the specific restriction site has been identified, the restriction enzyme can cut through the backbones of the double helix. Thousands of restriction enzymes have been studied and many are commercially available to laboratories, allowing scientists to process DNA in very specific ways. Some restriction enzymes produce offset cuts, yielding single-stranded “sticky ends” on the DNA fragments, as shown in Figure 6.21.
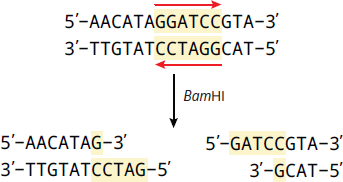
Figure 6.21 A Restriction Enzyme (Bam HI) Creating Sticky Ends Restriction enzymes cut at palindromic sequences, such as BamHI, which cuts the sequence GGATCC.
Sticky ends generated from restriction enzymes are advantageous in facilitating the insertion of DNA into a vector to create recombinant DNA. First, the DNA fragment to be inserted is created with the chosen restriction site flanking both ends of the DNA. The DNA fragment is then treated with the restriction enzyme. The vector of choice, which contains the same restriction site, is also cut with the same restriction enzyme, which allows the DNA fragment to be inserted directly into the vector and then ligated together. The polylinker, or MCS, is a common site of DNA fragment insertion within a vector. The polylinker is a short segment of DNA that contains the sequences of multiple different restriction sites that are not found in any other parts of the plasmid to allow for easy insertion of the DNA fragment into the vector after restriction enzyme digestion. Just 5′ of the polylinker is a promoter sequence that will initiate transcription of the inserted DNA fragment after the final recombinant plasmid is inserted into a host cell. The formation of a recombinant plasmid using restriction enzymes is shown in Figure 6.22.
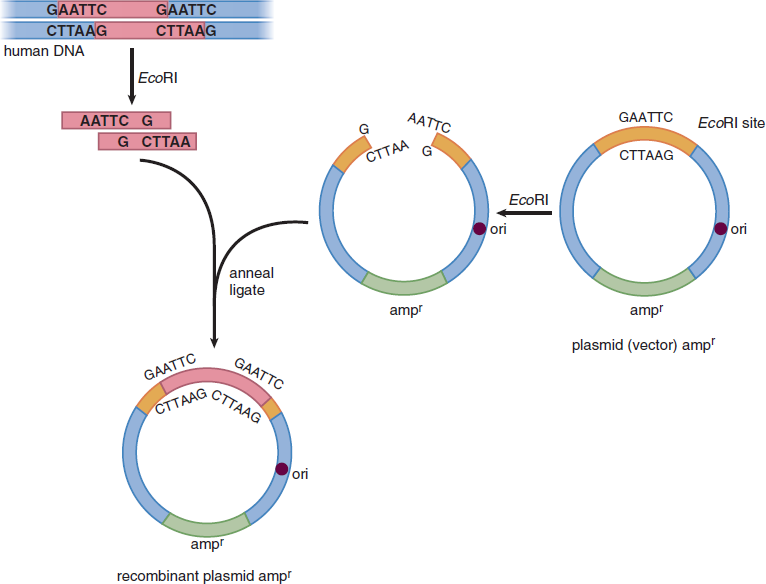
Figure 6.22 Formation of a Recombinant Plasmid Vector ori: origin of replication; ampr: gene for resistance to ampicillin (an antibiotic)
Once the DNA sequence of interest is inserted into the vector backbone, the recombinant plasmid can be inserted into a host, such as certain strains of the bacterium E. coli. The recombinant plasmid is inserted into the bacteria through transformation (a process described in further detail in Chapter 1 of MCAT Biology Review). Once inside the host bacteria, the origin of replication (ori) within the recombinant plasmid allows for its duplication. The bacteria are then spread onto plates containing a solid growth medium and allowed to form colonies. The selection marker of the plasmid aids in allowing for the isolation of bacteria that contain the recombinant plasmid. For example, plasmids that contain antibiotic resistance genes confer antibiotic resistance to the host bacteria, so when the transformed bacteria are spread onto plates whose growth media contain that antibiotic, only the bacteria that carry the plasmid with the resistance gene can grow. A summary of common key sequences found in vectors used in molecular cloning are summarized below in Table 6.3.
Table 6.3. Summary of Common Elements in Plasmid Vectors
PLASMID ELEMENT DESCRIPTION
Origin of Replication (Ori) An origin of replication (ori) is a DNA sequence that is the site on the plasmid where replication of the plasmid is initiated.
Multiple Cloning Site (MCS) or Polylinker These are short segments of DNA in vectors that contain multiple different unique restriction enzyme sites to allow for easy insertion of foreign DNA into the plasmid.
Selectable Marker Selectable markers are genes that allow for the marking or differentiation of cells that contain the plasmid through expression of the gene. Markers can include antibiotic resistance genes (see below), fluorescent proteins, or proteins that turn cells into different colors.
Antibiotic Resistance Gene Commonly, plasmids contain genes that confer resistance to certain antibiotics. This allows cells that contain the plasmid to have a survival advantage over other cells, and can thus be used to select only for cells that contain the desired plasmid. Antibiotic resistance genes are a type of selectable marker.
Once the bacterial colonies carrying the recombinant plasmid are isolated, they can be grown in large quantities to isolate large amounts of the now-amplified recombinant DNA for further use or analysis. Bacteria transformed with recombinant plasmids that carry protein-coding genes can also be used to express the protein, which can be purified from the cells for further use. The process of creating recombinant DNA by gene cloning and some of its applications are summarized in Figure 6.23.
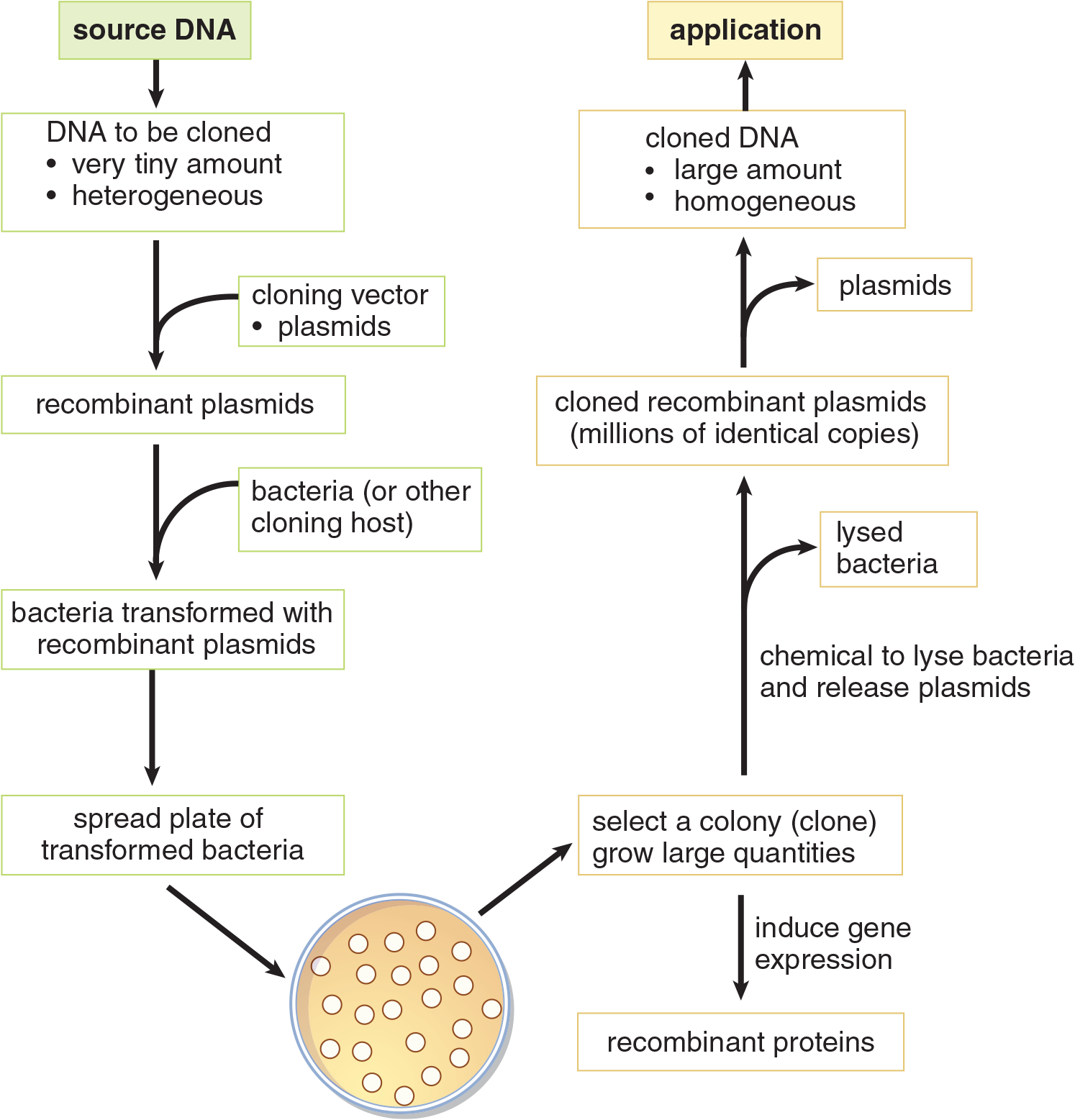
Figure 6.23 Cloning Recombinant DNA Cloning allows production of recombinant proteins, or identification and characterization of DNA by increasing its volume and purity.
DNA Libraries and cDNA
DNA cloning can be used to produce DNA libraries. DNA libraries are large collections of known DNA sequences; in sum, these sequences could equate to the genome of an organism. To make a DNA library, DNA fragments, often digested randomly, are cloned into vectors and can be utilized for further study. Libraries can consist of either genomic DNA or cDNA. Genomic libraries contain large fragments of DNA, and include both coding (exon) and noncoding (intron) regions of the genome. cDNA (complementary DNA) libraries are constructed by reverse-transcribing processed mRNA, as shown in Figure 6.24. As such, cDNA lacks noncoding regions, such as introns, and only includes the genes that are expressed in the tissue from which the mRNA was isolated. For that reason, these libraries are sometimes called expression libraries. While genomic libraries contain the entire genome of an organism, genes may by chance be split into multiple vectors. Therefore, only cDNA libraries can be used to reliably sequence specific genes and identify disease-causing mutations, produce recombinant proteins (such as insulin, clotting factors, or vaccines), or produce transgenic animals. Several of these applications are discussed in more detail in subsequent sections of this chapter. Table 6.4 contrasts some of the characteristics of genomic and cDNA libraries.
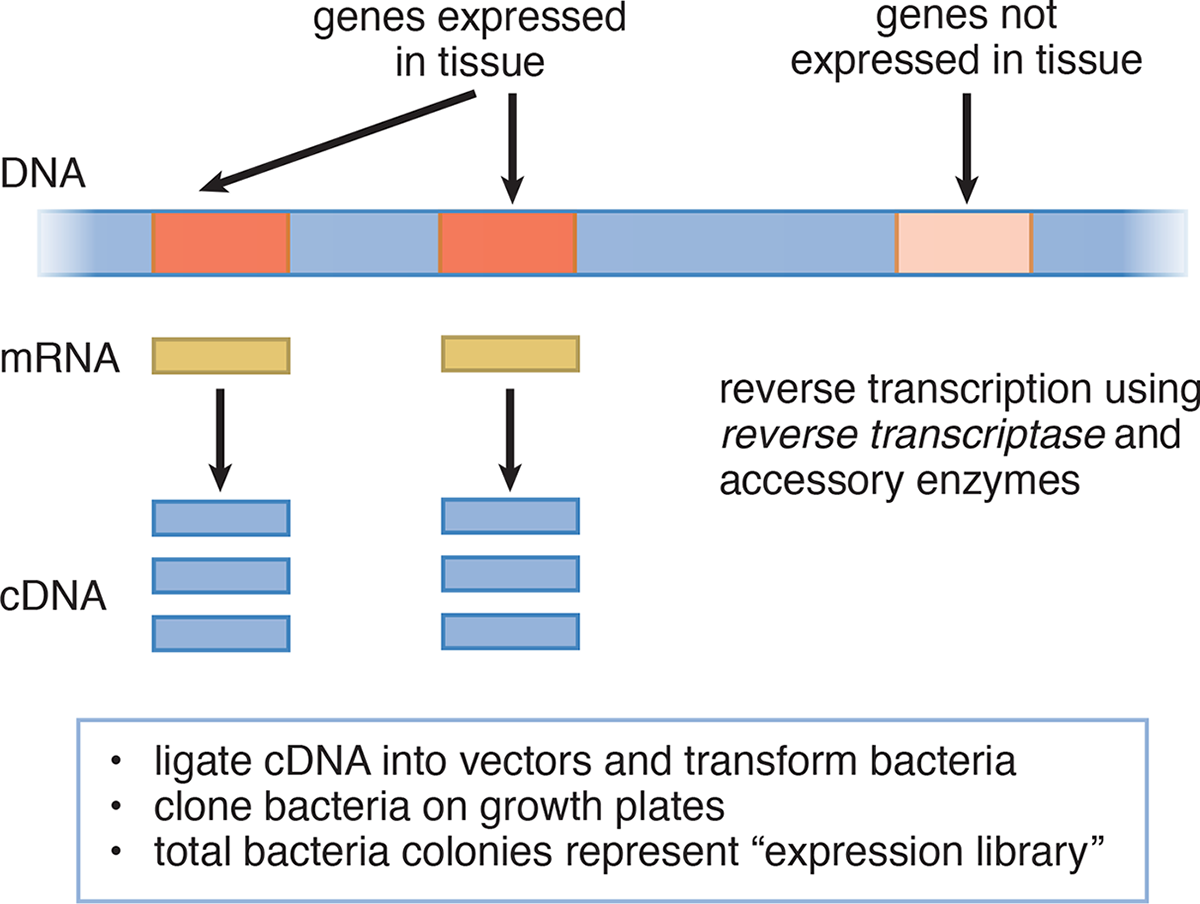
Figure 6.24 Cloning Expressed Genes by Producing cDNA
Table 6.4. Comparison of Genomic and cDNA (Expression) Libraries
GENOMIC LIBRARIES cDNA (EXPRESSION) LIBRARY
Source of DNA Chromosomal DNA mRNA (cDNA)
Enzymes to make library Restriction endonuclease DNA ligase Reverse transcriptase DNA ligase
Contains nonexpressed sequences of chromosomes Yes No
Cloned genes are complete sequences Not necessarily Yes
Cloned genes contain introns Yes No
Promoter and enhancer sequences present Yes, but not necessarily in same clone No
Gene can be expressed in cloning host (recombinant proteins) No Yes
Can be used for gene therapy or constructing transgenic animals No Yes
Hybridization
Another tool often used by researchers is called hybridization. Hybridization is the joining of complementary base pair sequences. This can be DNA–DNA recognition or DNA–RNA recognition. This technique uses two single-stranded sequences and is a vital part of polymerase chain reaction and Southern blotting.
Polymerase Chain Reaction
Polymerase chain reaction (PCR) is a powerful and widely-used method of DNA amplification, developed in 1983 by the biochemist Kary Mullis, that can produce millions of copies of a DNA sequence. PCR is fundamental in many research-based assays, and is commonly used as a way to identify criminal suspects, assess familial relationships, and look for the presence of disease-causing organisms in medical samples.
BRIDGE
PCR provides a great example of the temperature dependence of enzymes. While human DNA polymerase denatures at the high temperatures required in PCR, the DNA polymerase from T. aquaticus functions optimally at these temperatures. Refer to Chapter 2 of MCAT Biochemistry Review for more on the link between temperature and enzyme activity.
To amplify a desired region of DNA, a PCR reaction requires the DNA that contains the region of interest, primers that are complementary to the DNA that flanks the region of interest, nucleotides (dATP, dTTP, dCTP, and dGTP), and DNA polymerase. Once all combined, the reaction mixture will then undergo a thermal cycle to double the amount of initial DNA. A thermal cycle in PCR involves 3 major steps:
- Denaturation: In this step, the PCR reaction mixture is heated to nearly boiling temperatures (around 95 °C) to separate the two DNA strands.
- Primer annealing: The solution is cooled enough that the primers are able to bind to the complementary regions that flank the region of interest, while staying warm enough to avoid the two DNA strands from reannealing.
- Elongation: DNA polymerase will synthesize, starting at the primers, a new complementary DNA strand, leading to doubling the amount of DNA that was initially in the mixture.
This process of thermal cycling, with each cycle involving the steps of denaturation, primer annealing, and elongation, is repeated up to 20-40 times, or until there are enough copies of the DNA sequence for further use. Figure 6.25 illustrates an example of this. Unfortunately, the DNA polymerase found in the human body does not work at high temperatures: Thus, heat-resistant DNA polymerases, such as the DNA polymerase Taq Polymerase from Thermus aquaticus, a species of bacteria that thrives in the hot springs of Yellowstone National Park at 70 °C, are typically used instead.
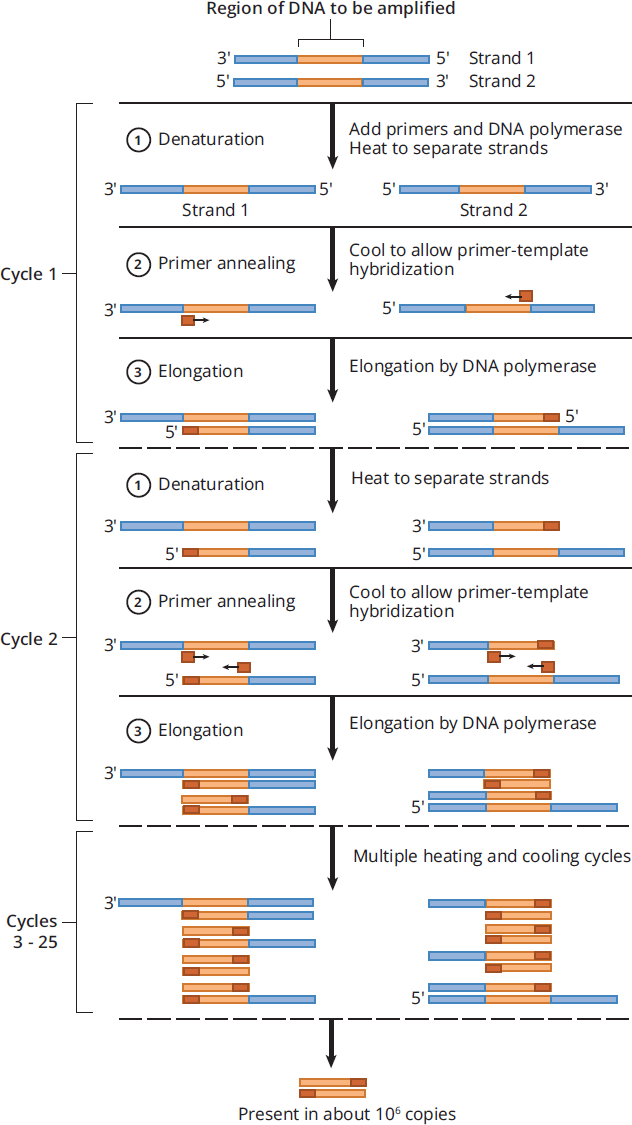
Figure 6.25 Denaturation, Primer Annealing, and Elongation in Each Thermal Cycle in PCR.
Reverse Transcription Polymerase Chain Reaction
Reverse transcription polymerase chain reaction (RT-PCR) is a technique used to convert a specific RNA sequence into DNA and then amplify the DNA. This technique can be used for cloning purposes but more commonly is used to measure the transcriptional activity of a gene of interest. In fact, RT-PCR is one of the most commonly used methods for detecting the COVID-19 virus. As seen in Figure 6.26, the RNA from a tissue sample is first extracted. Then, reverse transcriptase, an RNA-dependent DNA polymerase derived from retroviruses, is added, along with a primer complementary to the target mRNA, nucleotides, and DNA polymerase. Reverse transcriptase then creates a complementary DNA strand (cDNA) to the RNA, which is followed by amplification of the DNA by PCR with DNA polymerase. This technique allows for the quantification of RNA, even at low levels of expression, as the final amount of DNA produced correlates to how much RNA was in the original sample.
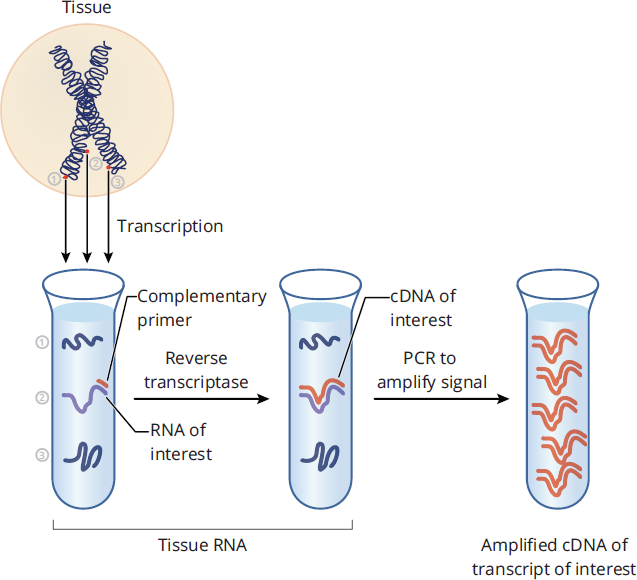
Figure 6.26 Reverse Transcription Polymerase Chain Reaction
Small Interfering RNA
Small interfering RNA (siRNA), also known as silencing RNA, are small non-coding RNA molecules that decrease the amount of a particular protein by interfering with the expression of specific mRNA strands before translation. In an experimental technique called a knockdown, introduction of siRNA into a cell temporarily represses the levels of a particular protein at the level of translation, in comparison to a knockout, which permanently disables gene expression entirely at the level of DNA. Typically, siRNAs are double-stranded RNAs that match the sequence of the mRNA of the target protein. While naturally occurring, they can also be synthetically created to target a specific protein and then transfected into cells, where they are then incorporated into what is known as the RNA-inducing silencing complex (RISC), subsequently binding to the complementary target mRNA sequence. Activated RISC has endonuclease activity, which cleaves the target mRNA, making it unusable for translation and halting new protein creation. Figure 6.27 shows the general process by which siRNAs inhibit translation of proteins.
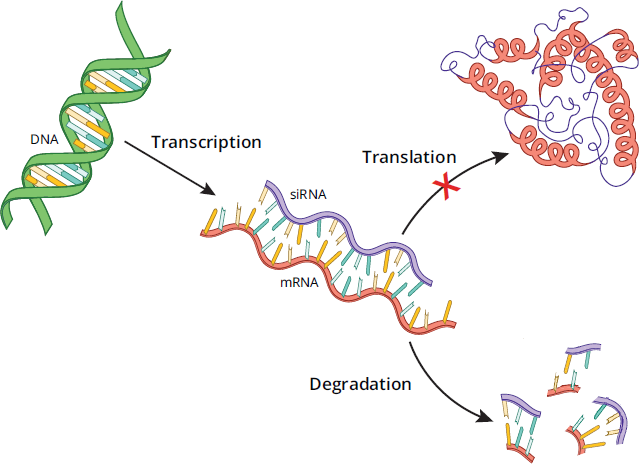
Figure 6.27 Mechanism of Protein Knockdowns Using Small Interfering RNAs (siRNAs)
Blotting and Molecular Labeling
Gel electrophoresis is a technique used to separate macromolecules, such as DNA and proteins, by size and/or charge. Electrophoresis of proteins and the different types of PAGE were discussed in detail in Chapter 3 of MCAT Biochemistry Review, but DNA can also be separated using gel electrophoresis. All molecules of DNA are negatively charged because of the phosphate groups in the backbone of the molecule, so all DNA strands will migrate toward the anode of an electrochemical cell. The preferred gel for DNA electrophoresis is agarose gel, and—just like proteins in polyacrylamide gel—the longer the DNA strand, the slower it will migrate in the gel.
MCAT EXPERTISE
The MCAT of course does not test capitalization conventions, but you may have noticed a difference in how the different types of blotting are spelled out. The term “Southern blot” derives its title from its inventor, Edwin Southern. For this reason, Southern is capitalized. However, western and northern blots, which are a play on words of “Southern” and not the name of any individual, are lower case.
Gel electrophoresis is often used prior to blotting, which is a method of transferring molecules like DNA, RNA, and proteins onto a thin membrane. Once the molecules have transferred onto the membrane, they can then be visualized in a number of different ways, depending on the experimental set-up. On the MCAT, three types of blots are tested on the exam: Southern blots, northern blots, and western blots. In each of these, detection of the molecule of interest creates bands on the membrane. However, it is important to note that blots are semi-quantitative methods of detection. While thicker bands correlate with more of the molecule of interest being present, an exact amount of how much of the molecule of interest is present cannot be determined. Figure 6.28 summarizes Southern, northern, and western blotting, each described in detail below.
Southern Blot
A Southern blot is used after gel electrophoresis to detect the presence of specific DNA sequences in a sample, as seen in Figure 6.28. After purification, DNA is cut by restriction enzymes into fragments before being separated by gel electrophoresis. The DNA fragments are then carefully transferred (or blotted) onto a membrane. The membrane is then probed with many copies of a labeled DNA sequence that is complementary to the DNA sequence of interest, which will bind and create double-stranded DNA. The probes are labeled, usually either with radioisotopes or fluorescent proteins, which are used to visualize the presence of the desired DNA sequence after probe binding.
Northern Blot
A northern blot is similar to a Southern blot, but instead separates and probes for RNA sequences of interest. Figure 6.28 shows the process of generating a northern blot. After purification, RNA is separated by gel electrophoresis and then transferred onto a membrane that is subsequently washed with labeled nucleotide probes that are complementary to the RNA sequence of interest. Note that unlike DNA, which requires fragmentation due to size, RNA molecules are usually short enough to not need fragmentation with restriction enzymes prior to gel electrophoresis.
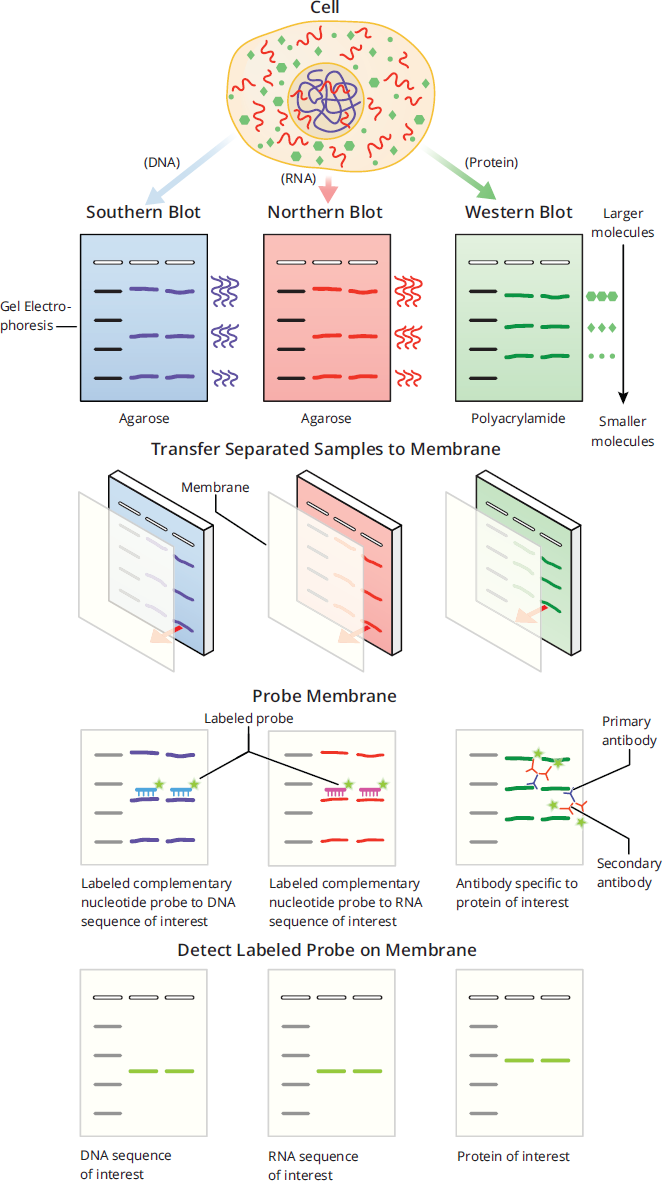
Figure 6.28 Summary of Southern, Western, and Northern Blots
Western Blot
Like northern and Southern blots, western blots follow the pattern of separation by gel electrophoresis, transfer to a membrane, and detection through the use of a labeled probe, as shown in Figure 6.28. However, western blots are used to detect specific proteins rather than nucleotides. As discussed in detail in Chapter 3 of MCAT Biochemistry Review, after purification, proteins are separated on gels made of polyacrylamide rather than agarose. After transfer to a membrane, proteins are then probed using antibodies specific to that protein. The antibodies that bind directly to the protein of interest are called primary antibodies. While primary antibodies can be radiolabeled or bound with a fluorescent protein for detection, it is more common that unlabeled primary antibodies are detected with a labeled secondary antibody, which recognizes and binds to the primary antibodies. The use of a secondary antibody amplifies the signal for better detection of proteins on the membrane: for every primary antibody bound to the protein, multiple secondary antibodies can bind to the primary antibody. On a practical level, the use of labeled secondary antibodies is also more widely applicable than using labeled primary antibodies: Primary antibodies bind to very specific molecules and protein sequences, even down to the level of post-translational modifications, meaning detection can only occur if each different primary antibody is labeled. However, secondary antibodies are created to recognize the Fc portion of other antibodies and can therefore bind to multiple different primary antibodies, making for a more generalized method of detection. Antibodies are covered in more depth in Chapter 8 of MCAT Biology Review.
Western blots also use loading controls to ensure that all the samples loaded onto the gel contain equal amounts of protein. Loading controls in western blots are usually proteins that are known as “house-keeping” genes, which are highly and ubiquitously expressed in cells, and therefore should have consistent levels regardless of experimental conditions. Loading controls are also used in other types of assays, such as Southern and northern blots, but most frequently appear on the MCAT in the context of western blots. The MCAT will not expect memorization or knowledge of specific loading controls, such as β-actin, GAPDH (an enzyme found in glycolysis), or vinculin, but they will expect recognition of loading controls. Without loading controls, it would be impossible to determine if an experimental condition changed the levels of a protein, or if more or less sample was added. Figure 6.29 shows the use of GAPDH as a loading control, where the levels of GAPDH are equal between the three samples, thus ensuring that there was indeed equal loading of all three wells with equal amounts of sample. This means that the levels of BRCA1 and BARD1 truly did decrease in Samples 1 and 2 compared to the control.
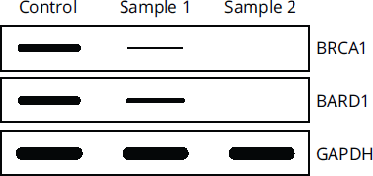
Figure 6.29 GAPDH as a Loading Control GAPDH levels are equal, indicating BRCA1 and BARD1 levels actually decreased.
Molecular Labeling
Many molecules cannot be directly seen with the human eye. Therefore, methods of detection are crucial for the visualization of what is normally invisible. There are numerous different techniques, many of which are beyond the scope of the MCAT, but a few different types that frequently come up in passages and questions include fluorescence and autoradiography.
A fluorescent tag, also called a fluorescent probe, is a fluorescent molecule that is bound or chemically linked to a non-fluorescent molecule, such as a protein, amino acid, or oligonucleotide. The phenomenon of fluorescence is described in detail in Chapter 9 of MCAT Physics and Math Review, but in short, the addition of a fluorescent tag allows for the detection of the labeled molecule using light. Fluorescent proteins, such as green fluorescent protein (GFP), can be added to other non-fluorescent proteins for tracking or detection, such as on antibodies during western blot visualization. Ethidium bromide, an intercalator that inserts itself between the bases of DNA, is a type of fluorescent dye that allows for the visualization of DNA when exposed to UV light as shown in Figure 6.30.
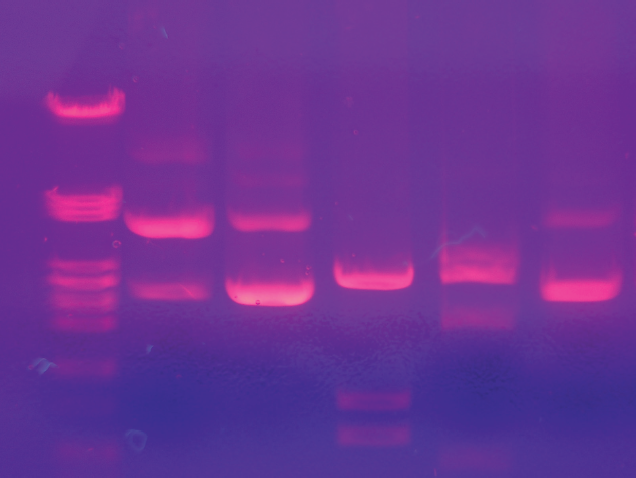
Figure 6.30 DNA Labeled with Ethidium Bromide An agarose gel with DNA lights up under UV light after treatment with ethidium bromide.
Autoradiography is a photographic method of detecting radioactive substances through the exposure of film or a digital detection device to the sample to create an image. Radiolabeling, which is a form of isotopic labeling, is commonly used to label molecules with radioactivity for tracking and detection using autoradiography. This is accomplished by replacing certain atoms with radioactive isotopes. On the MCAT, the most common radioactive isotopes include 14C, 35S, and 32P (note that 18O is not radioactive). The use of radiolabels allows for the tracking and identification of atoms and molecules like proteins and nucleic acids as they go through biochemical reactions. A common method of tagging phosphorylated proteins, as shown in Figure 6.31, is through the use of radiolabeled ATP, where the phosphorous atom in the γ-phosphate group has been tagged with 32P. When the γ-phosphate group is used in protein phosphorylation, the protein gains a radioactive phosphate group, which can then be tracked using autoradiography.
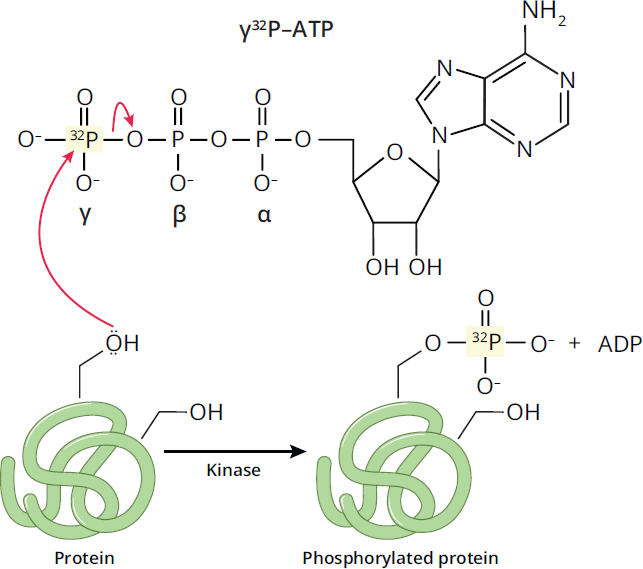
Figure 6.31 Radiolabeling ATP with 32P
Radiolabeling has been used to make many ground-breaking discoveries. In the Hershey-Chase experiments, which helped show that DNA, not protein, was genetic material, bacteriophages were labeled with either radioactive DNA using 32P or with radioactive proteins using 35S. When the bacteriophages were allowed to infect unlabeled bacteria, the viral progenies contained radioactive isotopes only with the bacteriophages labeled with 32P, while the progeny from 35S-labeled bacteriophages remained unlabeled.
DNA Sequencing
DNA sequencing, or the process of determining the nucleic acid sequence of a polynucleotide, has revolutionized the world that we live in. The applications of this technique are far-reaching, from the medical field to the justice system. Since the 1970s when DNA sequencing was first developed, the field has undergone major advancements and is still ever-evolving, with multiple methods of sequencing being used and improved upon in laboratories across the world. As such, the MCAT will not expect detailed knowledge on the many different types of DNA sequencing, but the test will expect a fundamental understanding of the basic principles that underlie many of these techniques.
One method of DNA sequencing that can be tested by the MCAT is Sanger sequencing (the chain termination method), an early first-generation sequencing technique developed in 1977 by Frederick Sanger and colleagues. It was, for nearly four decades, the main method of DNA sequencing, and while next-generation sequencing (NGS) methods have become more popular, Sanger sequencing is still widely used today.
REAL WORLD
The Human Genome Project, initiated in 1991, involved the identification of all 3 billion base pairs of the human DNA sequence. The first draft of this project was completed in 2000. This project demonstrated that although humans appear to be quite different from each other, the sequence of our DNA is, in reality, highly conserved. On average, two unrelated individuals still share over 99.9% of their DNA sequences.
The premise of Sanger sequencing relies on the use of analogues to the four nucleotide bases called dideoxyribonucleotides (ddNTPs), which include ddATP, ddCTP, ddGTP, and ddTTP. Unlike deoxyribonucleotides (dNTPs), which are dATP, dCTP, dGTP, and dTTP, ddNTPs contain a hydrogen at C-3′ of the ribose sugar, rather than a hydroxyl group. The C-3′ hydroxyl group is required for nucleotide chain extension, so when a ddNTP is incorporated into the growing chain, a subsequent nucleotide cannot be added by DNA polymerase, terminating the chain. The difference between dNTPs and ddNTPs is shown in Figure 6.32.
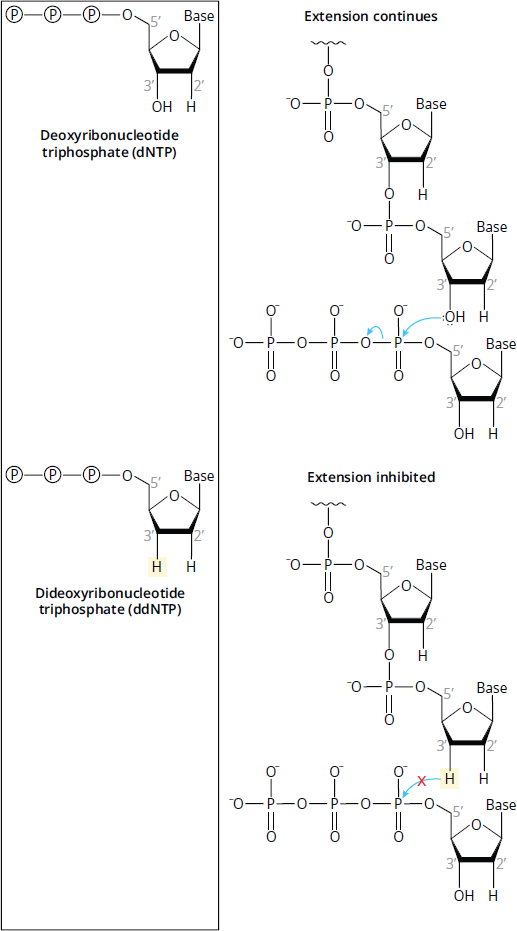
Figure 6.32 Comparison of Deoxynucleotides (dNTPs) and Dideoxynucleotides (ddNTPs) Once a ddNTP has been incorporated, another nucleotide cannot be added.
The basic steps of Sanger sequencing using ddNTPs are outlined in Figure 6.33.
- In the first step, a mixture is created that contains the DNA strand to be sequenced, all four dNTPs, a primer, and DNA polymerase. This mixture is then split into four different tubes, where each tube contains only one of the four ddNTPs, each of which are labeled.
- In the second step, the complementary strand of the template DNA in each of the four tubes is synthesized by DNA polymerase. During elongation, extension of the complementary sequence is terminated when a ddNTP is incorporated instead of a dNTP, leading to the creation of DNA fragments. Because each tube only contains one type of ddNTP, all of the DNA fragments within one specific tube will terminate with that specific ddNTP. For example, in the reaction mixture that contains ddCTP, the template sequence 3′—CAATCGGT—5′ will create two complementary DNA fragments: 5′—GTTAGC—3′ and 5′—GTTAGCC—3′—
- Once the reactions are complete, these fragments are then separated by size using gel electrophoresis for sequencing. To read the final sequence, from the 5′ to the 3′ direction, the bands are read from the bottom of the gel to the top of the gel, where the lane the bands are in corresponds to the incorporated base. Each subsequent band, going from bottom to top, represents a DNA fragment that is one nucleotide longer than the previous DNA fragment, and the added nucleotide is determined by which lane the band is in. The bands visualized within each individual lane correspond to the fragments that terminate with the ddNTP used for that specific reaction mixture.
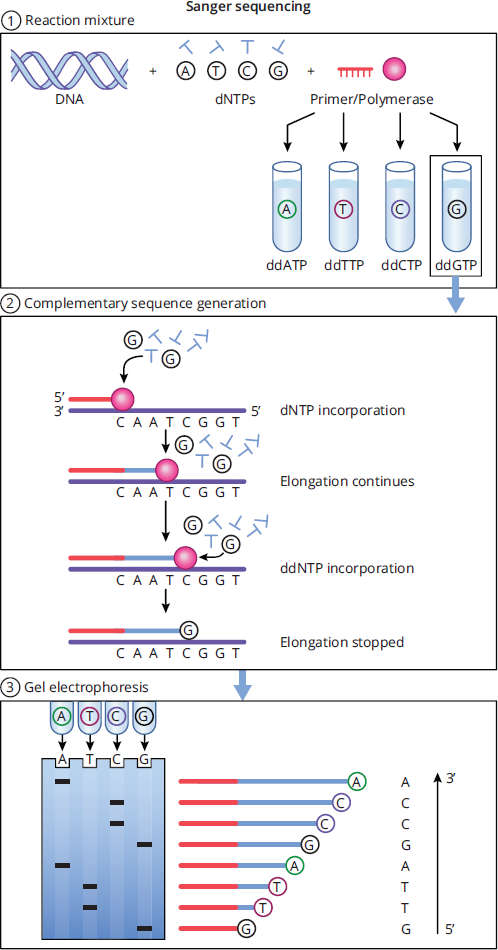
Figure 6.33 The Steps of Sanger Sequencing
Applications of DNA Technology
In addition to its utility as a research tool, DNA biotechnology has led to a number of therapeutic breakthroughs, ranging from gene therapy—described in this section—to development of personalized chemotherapeutic regimens in cancer by genotyping the tumor cells. DNA technology is also used in industry, including the development of genetically modified foods that are enriched with specific nutrients and testing of the environment for risk assessment and cleanup procedures. As mentioned previously, DNA technology also plays a key role in forensic pathology and crime scene investigation. This is likely only the beginning, as biotechnology continues to be an active area of research.
Gene Therapy
Gene therapy now offers potential cures for individuals with inherited diseases. Gene therapy is intended for diseases in which a given gene is mutated or inactive, giving rise to pathology. By transferring a normal copy of the gene into the affected tissues, the pathology should be fixed, essentially curing the individual. For instance, about half of children with severe combined immunodeficiency(SCID) have a mutation in the gene encoding the γ chain common to several of the interleukin receptors. By placing a working copy of the gene for the γ chain into a virus, one can transmit the functional gene into human cells. The first successful case of gene therapy was for SCID (caused by a different mutation) in 1990.
For gene replacement therapy to be a realistic possibility, efficient gene delivery vectors must be used to transfer the cloned gene into the target cells’ DNA. Because viruses naturally infect cells to insert their own genetic material, most gene delivery vectors in use are modified viruses. A portion of the viral genome is replaced with the cloned gene such that the virus can infect but not complete its replication cycle, as shown in Figure 6.34. Randomly integrated DNA poses a risk of integrating near and activating a host oncogene. Among the children treated for SCID, a small number have developed leukemias (cancers of white blood cells).
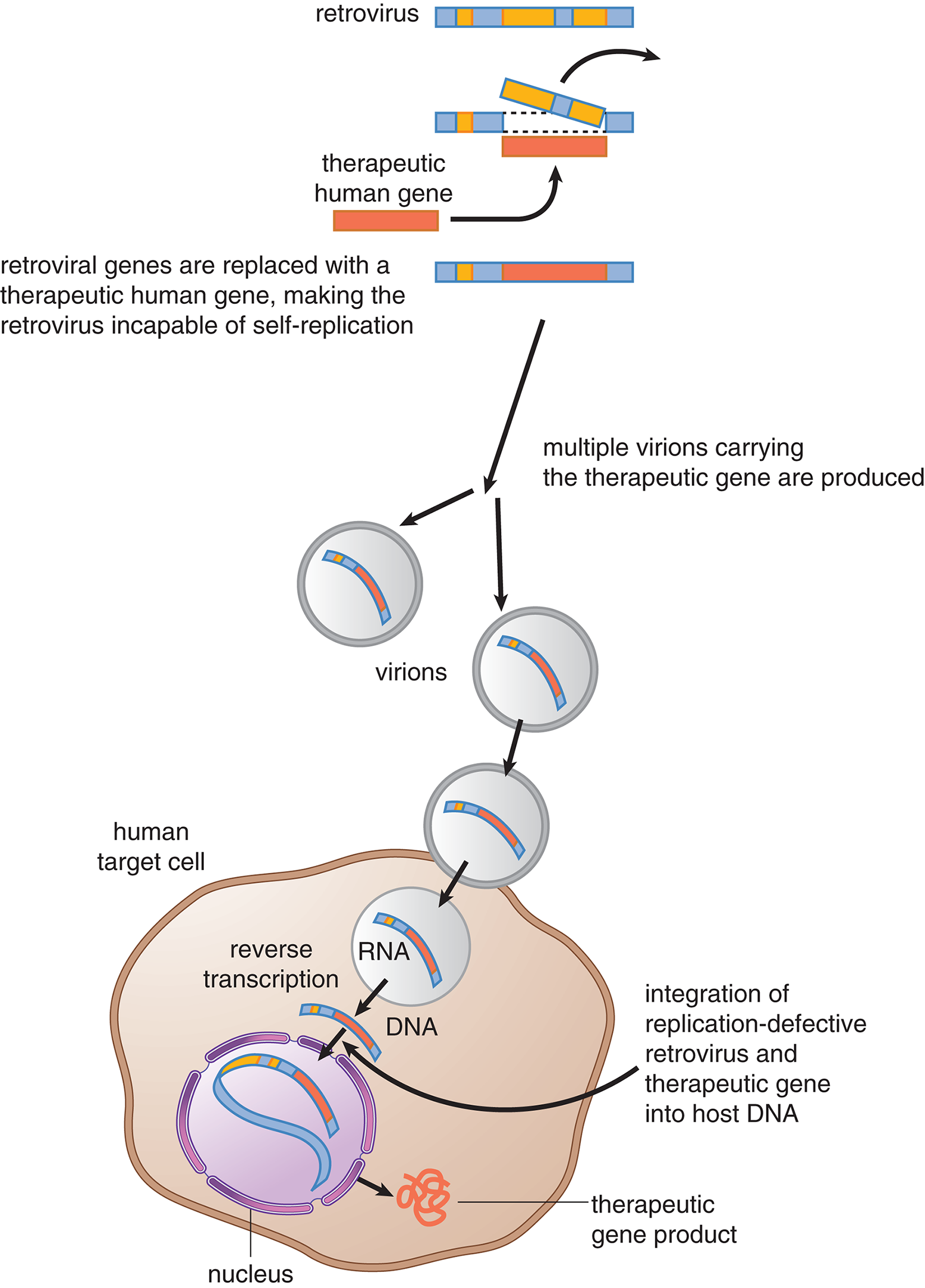
Figure 6.34 Retroviral Gene Therapy The example given here uses a retrovirus, but other viruses may also be used for gene therapy.
Transgenic and Knockout Mice
Once DNA has been isolated, it can be introduced into eukaryotic cells. Transgenic mice are altered at their germ line by introducing a cloned gene into fertilized ova or into embryonic stem cells. The cloned gene that is introduced is referred to as a transgene. If the transgene is a disease-producing allele, the transgenic mice can be used to study the disease process from early embryonic development through adulthood. A similar approach can be used to produce knockout mice, in which a gene has been intentionally deleted (knocked out). These mice provide valuable models in which to study human diseases.
There are different approaches to developing transgenic mice. A cloned gene may be microinjected into the nucleus of a newly fertilized ovum. Rarely, the gene may subsequently incorporate into the nuclear DNA of the zygote. The ovum is implanted into a surrogate, and, if successful, the resulting offspring will contain the transgene in all of their cells, including their germ line cells (gametes). Consequently, the transgene will also be passed to their offspring. The transgene coexists in the animals with their own copies of the gene, which have not been deleted. This approach is useful for studying dominant gene effects but is less useful as a model for recessive disease because the number of copies of the gene that insert into the genome cannot be controlled; the transgenic mice may each contain a different number of copies of the transgene. This method is demonstrated in Figure 6.35.
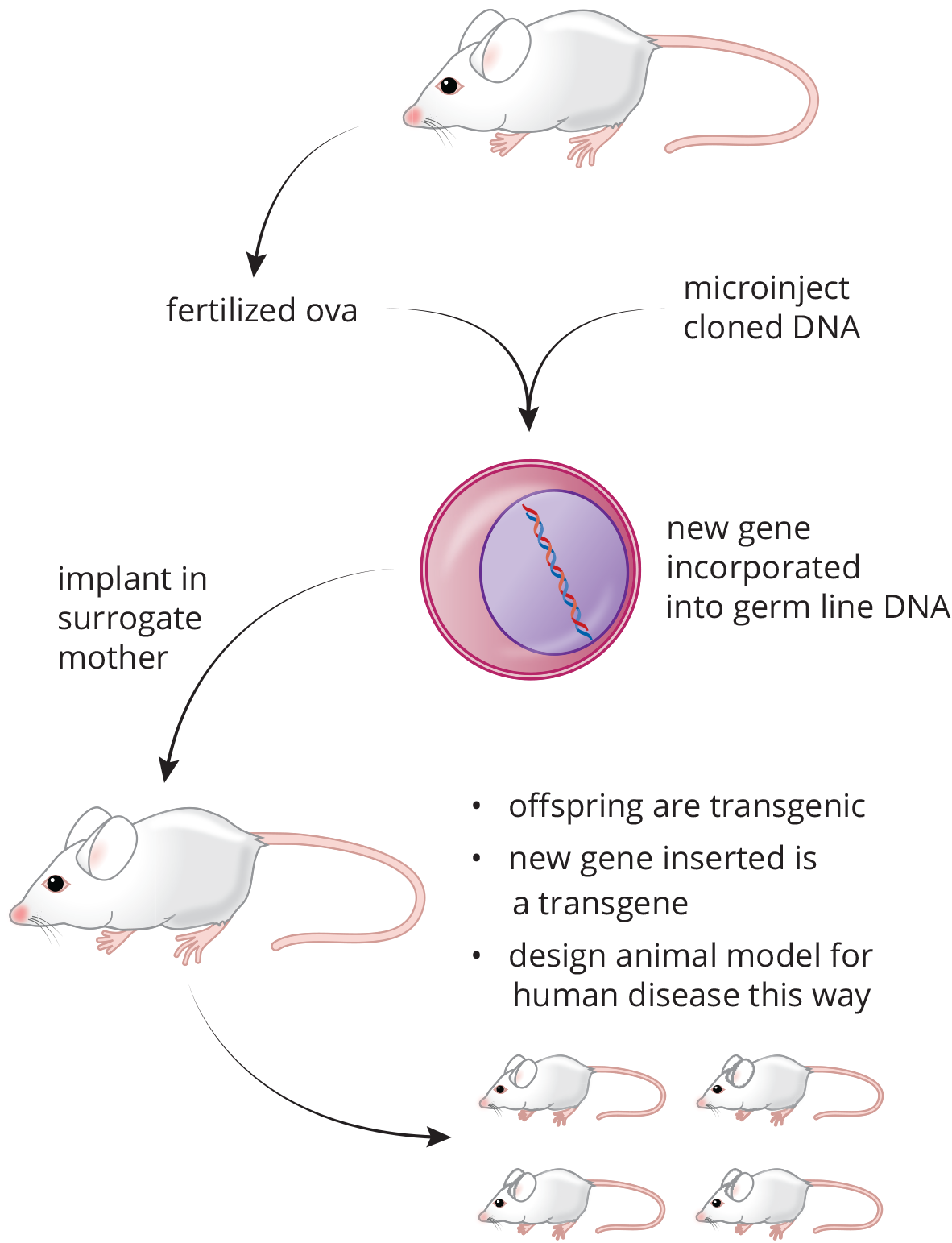
Figure 6.35 Creation of a Transgenic Mouse
Embryonic stem cell lines can also be used for developing transgenic mice. Advantages of using stem cell lines are that the cloned genes can be introduced in cultures, and that one can select for cells with the transgene successfully inserted. The altered stem cells are injected into developing blastocysts and implanted into surrogates. The blastocyst itself is thus composed of two types of stem cells: the ones containing the transgene and the original blastocyst cells that lack the transgene. The resulting offspring is a chimera, meaning that it has patches of cells, including germ cells, derived from each of the two lineages. This is evident if the two cell lineages (transgenic cells and host blastocyst) come from mice with different coat colors. The chimeras will have patchy coats of two colors, allowing them to be easily identified. These chimeras can then be bred to produce mice that are heterozygous for the transgene and mice that are homozygous for the transgene.
Safety and Ethics
The different procedures and techniques that have been reviewed provide great insight for researchers in many different fields of study. However, it is also important to acknowledge the potential risks associated with these technologies. Safety concerns such as increased resistance in viruses and bacteria can impact both humans and the environment in which we live. Ethical dilemmas arise: Is it ethical to test for life-threatening genetic diseases and potentially terminate a pregnancy based on the results? What about testing for eye or hair color? What are the ethical questions around choosing human test subjects? If a disease-causing gene were found in one individual of a family, does this need to be communicated to other relatives at risk, potentially violating principles of privacy? Is it permissible to carry out potentially risky therapies in individuals whose illnesses makes them unable to communicate? The medical community and bioethicists at large continue to wrestle with this question: How much should we meddle with our own genetic makeup?
REAL WORLD
The story of Jesse Gelsinger is one particularly harrowing example of the problems that biotechnology can create. Diagnosed with mild ornithine transcarbamylase (OTC) deficiency, a defect of the urea cycle, Gelsinger volunteered for a clinical trial of gene therapy. During the trial (which was also fraught with questionable consent practices), Gelsinger died from a massive immune response to the viral vector, leading to multiple organ failure and brain death. He was 18 years old.
MCAT CONCEPT CHECK 6.5:
Before you move on, assess your understanding of the material with these questions.
- When creating a DNA library, what are some of the advantages of genomic libraries? What about cDNA libraries?
________________________________________
- Genomic:
________________________________________
- cDNA:
- What does PCR accomplish for a researcher? What about Southern blotting?
________________________________________
- PCR:
________________________________________
- Southern blotting:
________________________________________
- During DNA sequencing, why does the DNA polymer stop growing once a dideoxyribonucleotide is added?
________________________________________
- What is the difference between a transgenic and a knockout mouse?
Conclusion
In this chapter, the DNA molecule was discussed. The importance of this molecule as an archive in the cell was highlighted. The unique structure of the DNA double helix and complementary base-pairing are integral to DNA molecules’ ability to replicate and pass information from cell to cell and from generation to generation. Replication of DNA is a complex process involving many enzymes and proteins that are highly coordinated to ensure efficiency. The discoveries of DNA structure and function aid us in understanding how cellular processes take place but also provide us with many tools to exploit in the laboratory.
DNA is one of the most heavily tested concepts on the MCAT. This makes sense because not only has our understanding of the molecule increased exponentially over the last few decades, but also our ability to manipulate DNA. This has led to the creation of an entire industry of biotechnology that will assuredly grow during your career as a medical student and physician. In the next chapter, we turn to DNA’s counterpart, RNA. We’ll explore how the genes discussed in this chapter can actually turn into functional proteins through the key processes of transcription and translation.
GO ONLINE!
You've reviewed the content, now test your knowledge and critical thinking skills by completing a test-like passage set in your online resources!
CONCEPT SUMMARY
DNA Structure
- Deoxyribonucleic acid (DNA) is a macromolecule that stores genetic information in all living organisms.
- Nucleosides contain a five-carbon sugar bonded to a nitrogenous base; nucleotides are nucleosides with one to three phosphate groups added.
- Nucleotides in DNA contain deoxyribose; in RNA, they contain ribose.
- Nucleotides are abbreviated by letter: adenine (A), cytosine (C), guanine (G), thymine (T), and uracil (U).
- DNA is organized according to the Watson–Crick model.
- The backbone is composed of alternating sugar and phosphate groups, and is always read 5′ to 3′.
- There are two strands with antiparallel polarity, wound into a double helix.
- Purines (A and G) always pair with pyrimidines (C, U, and T). In DNA, A pairs with T (via two hydrogen bonds) and C pairs with G (via three hydrogen bonds). RNA does not contain thymine, but contains uracil instead; thus, in RNA, A pairs with U (via two hydrogen bonds).
- Purines and pyrimidines are biological aromatic heterocycles. Aromatic compounds are cyclic, planar, and conjugated, and contain 4n + 2 π electrons (where n is any integer; Hückel’s rule). Heterocycles are ring structures that contain at least two different elements in the ring.
- Chargaff’s rules state that purines and pyrimidines are equal in number in a DNA molecule, and that because of base-pairing, the amount of adenine equals the amount of thymine, and the amount of cytosine equals the amount of guanine.
- Most DNA is B-DNA, forming a right-handed helix. Low concentrations of Z-DNA, with a zigzag shape, may be seen with high GC-content or high salt concentration.
- DNA strands can be pulled apart (denatured) and brought back together (reannealed). Heat, alkaline pH, and chemicals like formaldehyde and urea can cause denaturation of DNA; removal of these conditions may result in reannealing of the strands.
Eukaryotic Chromosome Organization
- DNA is organized into 46 chromosomes in human cells.
- In eukaryotes, DNA is wound around histone proteins (H2A, H2B, H3, and H4) to form nucleosomes, which may be stabilized by another histone protein (H1). As a whole, DNA and its associated histones make up chromatin in the nucleus.
- Heterochromatin is dense, transcriptionally silent DNA that appears dark under light microscopy.
- Euchromatin is less dense, transcriptionally active DNA that appears light under light microscopy.
- Telomeres are the ends of chromosomes. They contain high GC-content to prevent unraveling of the DNA. During replication, telomeres are slightly shortened, although this can be (partially) reversed by the enzyme telomerase.
- Centromeres are located in the middle of chromosomes and hold sister chromatids together until they are separated during anaphase in mitosis. They also contain a high GC-content to maintain a strong bond between chromatids.
DNA Replication
- The replisome (replication complex) is a set of specialized proteins that assist the DNA polymerases.
- To replicate DNA, it is first unwound at an origin of replication by helicases. This produces two replication forks on either side of the origin.
- Prokaryotes have a circular chromosome that contains only one origin of replication.
- Eukaryotes have linear chromosomes that contain many origins of replication.
- Unwound strands are kept from reannealing or being degraded by single-stranded DNA-binding proteins.
- Supercoiling causes torsional strain on the DNA molecule, which can be released by DNA topoisomerases, which create nicks in the DNA molecule.
- DNA replication is semiconservative: one old parent strand and one new daughter strand is incorporated into each of the two new DNA molecules.
- DNA cannot be synthesized without an adjacent nucleotide to hook onto, so a small RNA primer is put down by primase.
- DNA polymerase III (prokaryotes) or **DNA polymerases α,δ, and*ε** *(eukaryotes) can then synthesize a new strand of DNA; they read the template DNA 3′ to 5′ and synthesize the new strand 5′ to 3′.
- The leading strand requires only one primer and can then be synthesized continuously in its entirety.
- The lagging strand requires many primers and is synthesized in discrete sections called Okazaki fragments.
- RNA primers can later be removed by DNA polymerase I (prokaryotes) or RNase H (eukaryotes), and filled in with DNA by DNA polymerase I (prokaryotes) or DNA polymerase δ (eukaryotes). DNA ligase can then fuse the DNA strands together to create one complete molecule.
DNA Repair
- Oncogenes develop from mutations of proto-oncogenes, and promote cell cycling. They may lead to cancer, which is defined by unchecked cell proliferation with the ability to spread by local invasion or metastasize (migrate to distant sites via the bloodstream or lymphatic system).
- Tumor suppressor genes code for proteins that reduce cell cycling or promote DNA repair; mutations of tumor suppressor genes can also lead to cancer.
- During replication, DNA polymerase proofreads its work and excises incorrectly matched bases. The daughter strand is identified by its lack of methylation and corrected accordingly.
- Mismatch repair also occurs during the G2 phase of the cell cycle, using the genes MSH2 and MLH1.
- Nucleotide excision repair fixes helix-deforming lesions of DNA (such as thymine dimers) via a cut-and-patch process that requires an excision endonuclease.
- Base excision repair fixes nondeforming lesions of the DNA helix (such as cytosine deamination) by removing the base, leaving an apurinic/apyrimidinic (AP) site. An AP endonuclease then removes the damaged sequence, which can be filled in with the correct bases.
Recombinant Biotechnology
- Recombinant DNA is DNA composed of nucleotides from two different sources.
- DNA cloning introduces a fragment of DNA into a vector plasmid. A restriction enzyme (restriction endonuclease) cuts both the plasmid and the fragment, which are left with sticky ends. Once the fragment binds to the plasmid, it can be introduced into a bacterial cell and permitted to replicate, generating many copies of the fragment of interest.
- Recombinant vectors contain an origin of replication, a polylinker that the fragment of interest is placed into, and at least one type of selection marker (to permit for selection of that colony after replication).
- Once replicated, the bacterial cells can be used to create a protein of interest, or can be lysed to allow for isolation of the recombinant DNA.
- DNA libraries are large collections of known DNA sequences.
- Genomic libraries contain large fragments of DNA, including both coding and noncoding regions of the genome. They cannot be used to make recombinant proteins or for gene therapy.
- cDNA libraries (expression libraries) contain smaller fragments of DNA, and only include the exons of genes expressed by the sample tissue. They can be used to make recombinant proteins or for gene therapy.
- Hybridization is the joining of complementary base pair sequences.
- Polymerase chain reaction (PCR) is an automated process by which millions of copies of a DNA sequence can be created from a very small sample by hybridization.
- Blotting is a method of transferring molecules from a gel onto a thin membrane for further analysis.
- Molecules can be separated by size using gel electrophoresis.
- Southern blotting can be used to detect the presence and relative quantity of various DNA strands in a sample. After electrophoresis, the sample is transferred to a membrane that can be probed with single-stranded oligonucleotides to look for a sequence of interest.
- Northern blotting can be used to detect the presence and relative quantity of various RNA strands in a sample. After electrophoresis, the sample is transferred to a membrane that can be probed with single-stranded oligonucleotides to look for a sequence of interest.
- Western blotting can be used to detect the presence and relative quantity of various proteins in a sample. After electrophoresis, the sample is transferred to a membrane that can be probed with primary and secondary antibodies to look for a protein of interest.
- Sanger sequencing is a method of DNA sequencing that uses dideoxyribonucleotides, which terminate the DNA chain because they lack a 3′ -OH group. The resulting fragments can be separated by gel electrophoresis, and the sequence can be read directly from the gel.
- Molecular labeling is used to track molecules.
- Fluorescent tags can be added onto molecules and then imaged using light.
- Autoradiography is a photographic method of detecting radioactive substances.
- Gene therapy is a method of curing genetic deficiencies by introducing a functional gene with a viral vector.
- Transgenic mice are created by integrating a gene of interest into the germ line or embryonic stem cells of a developing mouse.
- Organisms that contain cells from two different lineages (such as mice formed by integration of transgenic embryonic stem cells into a normal mouse blastocyst) are called chimeras.
- Transgenic mice can be mated to select for the transgene.
- Knockout mice are created by deleting a gene of interest.
- Biotechnology brings up a number of safety and ethical issues, including pathogen resistance and the ethics of choosing individuals for specific traits.
ANSWERS TO CONCEPT CHECKS
**6.1**
- Nucleosides contain a five-carbon sugar (pentose) and nitrogenous base. Nucleotides are composed of a nucleoside plus one to three phosphate groups.
- A pairs with T (in DNA) or U (in RNA), using two hydrogen bonds. C pairs with G, using three hydrogen bonds.
- DNA contains deoxyribose, while RNA contains ribose. DNA contains thymine, while RNA contains uracil. Usually, DNA is double-stranded, while RNA is single-stranded.
- The aromaticity of nucleic acids makes these compounds very stable and unreactive. Stability is important for storing genetic information and avoiding spontaneous mutations.
- This does not violate Chargaff’s rules. RNA is single-stranded, and thus the complementarity seen in DNA does not hold true. For single-stranded RNA, %C does not necessarily equal %G; %A does not necessarily equal %U.
**6.2**
- The five histone proteins are H1, H2A, H2B, H3, and H4. H1 is the only one not in the histone core.
-
Characteristic Heterochromatin Euchromatin Density of chromatin packing Dense Not dense (uncondensed)
Appearance under light microscopy Dark Light
Transcriptional activity Silent Active
- High GC-content increases hydrogen bonding, making the association between DNA strands very strong at telomeres and centromeres.
**6.3**
-
Enzyme Prokaryotes/Eukaryotes/Both Function Helicase Both Unwinds DNA double helix
Single-stranded DNA-binding protein Both Prevents reannealing of DNA double helix during replication
Primase Both Places ∼10-nucleotide RNA primer to begin DNA replication
DNA polymerase III Prokaryotes Adds nucleotides to growing daughter strand
**DNA polymerase *α*** Eukaryotes Adds nucleotides to growing daughter strand
DNA polymerase I Prokaryotes Fills in gaps left behind after RNA primer excision
RNase H Eukaryotes Excises RNA primer
DNA ligase Both Joins DNA strands (especially between Okazaki fragments)
DNA topoisomerases Both Reduces torsional strain from positive supercoils by introducing nicks in DNA strand
- The lagging strand is more prone to mutations because it must constantly start and stop the process of DNA replication. Additionally, it contains many more RNA primers, all of which must be removed and filled in with DNA.
- Telomeres are the ends of eukaryotic chromosomes and contain repetitive sequences of noncoding DNA. These protect the chromosome from losing important genes from the incomplete replication of the 5′ end of the DNA strand.
**6.4**
- Oncogenes (or, more properly, proto-oncogenes) code for cell cycle–promoting proteins; when mutated, a proto-oncogene becomes an oncogene, promoting rapid cell cycling. Tumor suppressor genes code for repair or cell cycle–inhibiting proteins; when mutated, the cell cycle is allowed to proceed unchecked. Oncogenes are like stepping on the gas pedal, mutated tumor suppressor genes are like cutting the brakes.
- The parent strand is more heavily methylated, whereas the daughter strand is barely methylated at all. This allows DNA polymerase to distinguish between the two strands during proofreading.
-
Repair Mechanism Phase of Cell Cycle Key Enzymes/Genes DNA polymerase (proofreading) S DNA polymerase
Mismatch repair G2 MSH2, MLH1(MutS and MutL in prokaryotes)
Nucleotide excision repair G1, G2 Excision endonuclease
Base excision repair G1, G2 Glycosylase, AP endonuclease
- Nucleotide excision repair corrects lesions that are large enough to distort the double helix; base excision repair corrects lesions that are small enough not to distort the double helix.
**6.5**
- Genomic libraries include all of the DNA in an organism’s genome, including noncoding regions. This may be useful for studying DNA in introns, centromeres, or telomeres. cDNA libraries only include expressed genes from a given tissue, but can be used to express recombinant proteins or to perform gene therapy.
- PCR increases the number of copies of a given DNA sequence and can be used for a sample containing very few copies of the DNA sequence. Southern blotting is useful when searching for a particular DNA sequence because it separates DNA fragments by length and then probes for a sequence of interest.
- Dideoxyribonucleotides lack the 3′ –OH group that is required for DNA strand elongation. Thus, once a dideoxyribonucleotide is added to a growing DNA molecule, no more nucleotides can be added because dideoxyribonucleotides have no 3′ –OH group with which to form a bond.
- Transgenic mice have a gene introduced into their germ line or embryonic stem cells to look at the effects of that gene; they are therefore best suited for studying the effects of dominant alleles. Knockout mice are those in which a gene of interest has been removed, rather than added.
SCIENCE MASTERY ASSESSMENT EXPLANATIONS
1. B
Nucleotides bond together to form polynucleotides. The 3′ hydroxyl group of one nucleotide’s sugar joins the 5′ hydroxyl group of the adjacent nucleotide’s sugar by a phosphodiester bond. Hydrogen bonding, (A), is important for holding complementary strands together, but does not play a role in the bonds formed between adjacent nucleotides on a single strand.
2. D
Because we are looking for the false statement, we have to read each choice to eliminate those that are true or find one that is overtly false. Let’s quickly review the main differences between DNA and RNA. In cells, DNA is double-stranded, with a deoxyribose sugar and the nitrogenous bases A, T, C, and G. RNA, on the other hand, is usually single-stranded, with a ribose sugar and the bases A, U, C, and G. (D) is false because both DNA replication and RNA synthesis proceed in a 5′ to 3′ direction.
3. A
The melting temperature of DNA is the temperature at which a DNA double helix separates into two single strands (denatures). To do this, the hydrogen bonds linking the base pairs must be broken. Cytosine binds to guanine with three hydrogen bonds, whereas adenine binds to thymine with two hydrogen bonds. The amount of heat needed to disrupt the bonding is proportional to the number of bonds. Thus, the higher the GC-content in a DNA segment, the higher the melting point.
4. C
Aromatic rings must contain conjugated π electrons, which require alternating single and multiple bonds, or lone pairs. In carbohydrate ring structures, only single bonds are present, thus preventing aromaticity. Nucleic acids contain aromatic heterocycles, while proteins will generally contain at least one aromatic amino acid (tryptophan, phenylalanine, or tyrosine).
5. C
For a compound to be aromatic, it must be cyclic, planar, conjugated, and contain 4n+ 2 πelectrons, where n is any integer. Conjugation requires that every atom in the ring have at least one unhybridized p-orbital. While most examples of aromatic compounds have alternating single and double bonds, compounds can be aromatic if they contain triple bonds as well; this would still permit at least one unhybridized p-orbital.
6. C
During DNA replication, the strands are separated by DNA helicase. At the replication fork, primase, (A), creates a primer for the initiation of replication, which is followed by DNA polymerase. On the lagging strand, Okazaki fragments form and are joined by DNA ligase, (B). After the chromosome has been processed, the ends, called telomeres, are replicated with the assistance of the enzyme telomerase, (D). RNA polymerase I is located in the nucleolus and synthesizes rRNA.
7. B
cDNA (complementary DNA) is formed from a processed mRNA strand by reverse transcription. cDNA is used in DNA libraries and contains only the exons of genes that are transcriptionally active in the sample tissue.
8. A
The polymerase chain reaction is used to clone a sequence of DNA using a DNA sample, a primer, free nucleotides, and enzymes. The polymerase from Thermus aquaticus is used because the reaction is regulated by thermal cycling, which would denature human enzymes.
9. D
Endonucleases are enzymes that cut DNA. They are used by the cell for DNA repair. They are also used by scientists during DNA analysis, as restriction enzymes are endonucleases. Restriction enzymes are used to cleave DNA before electrophoresis and Southern blotting, and to introduce a gene of interest into a viral vector for gene therapy.
10. D
Prokaryotic DNA is circular and lacks histone proteins, and thus does not form nucleosomes. Both prokaryotic and eukaryotic DNA are replicated by DNA polymerases, although these polymerases differ in identity. Eukaryotic DNA is organized into chromatin, which can condense to form linear chromosomes; only prokaryotes have circular chromosomes. Only eukaryotic DNA has telomeres.
11. C
One common DNA mutation is the transition from cytosine to uracil in the presence of heat. DNA repair enzymes recognize uracil and correct this error by excising the base and inserting cytosine. RNA exists only transiently in the cell, such that cytosine degradation is insignificant. Were uracil to be used in DNA under normal circumstances, it would be impossible to tell if a base should be uracil or if it is a damaged cytosine nucleotide.
12. A
Oncogenes are most likely to result in cancer through activation, (B), while tumor suppressor genes are most likely to result in cancer through inactivation.
13. B
One of the primary ethical concerns related to gene sequencing is the issue of consent and privacy. Because genetic screening provides information on direct relatives, there are potential violations of privacy in communicating this information to family members who may be at risk. There are not significant physical risks, eliminating (A), and gene sequencing is fairly accurate, eliminating (C) and (D).
14. D
Euchromatin has a classic “beads on a string” appearance that stains lightly, while heterochromatin is tightly packed and stains darkly. Heterochromatin is primarily composed of inactive genes or untranslated regions, while euchromatin is able to be expressed. All chromatin is found in the nucleus, not the cytoplasm.
15. D
Mismatch repair mechanisms are active during S phase (proofreading) and G2 phase (MSH2 and MLH1), eliminating (B) and (C). Nucleotide and base excision repair mechanisms are most active during the G1 and G2 phases, also eliminating (A). These mechanisms exist during interphase because they are aimed at preventing propagation of the error into daughter cells during M phase (mitosis).
GO ONLINE
Consult your online resources for additional practice.
SHARED CONCEPTS
- Biochemistry Chapter 4
- Carbohydrate Structure and Function
- Biochemistry Chapter 7
- RNA and the Genetic Code
- Biology Chapter 1
- The Cell
- Biology Chapter 2
- Reproduction
- Biology Chapter 3
- Embryogenesis and Development
- Biology Chapter 12
- Genetics and Evolution