Chapter 7: RNA and the Genetic Code

Chapter 7: RNA and the Genetic Code
SCIENCE MASTERY ASSESSMENT
Every pre-med knows this feeling: there is so much content I have to know for the MCAT! How do I know what to do first or what's important?
While the high-yield badges throughout this book will help you identify the most important topics, this Science Mastery Assessment is another tool in your MCAT prep arsenal. This quiz (which can also be taken in your online resources) and the guidance below will help ensure that you are spending the appropriate amount of time on this chapter based on your personal strengths and weaknesses. Don't worry though— skipping something now does not mean you'll never study it. Later on in your prep, as you complete full-length tests, you'll uncover specific pieces of content that you need to review and can come back to these chapters as appropriate.
How to Use This Assessment
If you answer 0–7 questions correctly:
Spend about 1 hour to read this chapter in full and take limited notes throughout. Follow up by reviewing all quiz questions to ensure that you now understand how to solve each one.
If you answer 8–11 questions correctly:
Spend 20–40 minutes reviewing the quiz questions. Beginning with the questions you missed, read and take notes on the corresponding subchapters. For questions you answered correctly, ensure your thinking matches that of the explanation and you understand why each choice was correct or incorrect.
If you answer 12–15 questions correctly:
Spend less than 20 minutes reviewing all questions from the quiz. If you missed any, then include a quick read-through of the corresponding subchapters, or even just the relevant content within a subchapter, as part of your question review. For questions you got correct, ensure your thinking matches that of the explanation and review the Concept Summary at the end of the chapter.
- What role does peptidyl transferase play in protein synthesis?
- It transports the initiator aminoacyl-tRNA complex.
- It helps the ribosome to advance three nucleotides along the mRNA in the 5′ to 3′ direction.
- It holds the protein in its tertiary structure.
- It catalyzes the formation of a peptide bond.
-
A mutation in which of the following components of the lac operon would lead to a significant reduction in the expression of lactase (one of the structural genes)?
- Operator
- Regulatory gene
- Promoter
- Structural genes
- Topoisomerases are enzymes involved in:
- DNA replication and transcription.
- posttranscriptional processing.
- RNA synthesis and translation.
- posttranslational processing.
- Val-tRNAValis the tRNA that carries valine to the ribosome during translation. Which of the following sequences gives an appropriate anticodon for this tRNA? (Note: Refer to Figure 7.6 for a genetic code table.)
- CAU
- AUC
- UAC
- GUG
- Enhancers are transcriptional regulatory sequences that function by enhancing the activity of:
- RNA polymerase at a single promoter site.
- RNA polymerase at multiple promoter sites.
- spliceosomes and lariat formation in the ribosome.
- transcription factors that bind to the promoter but not to RNA polymerase.
- In the genetic code of human nuclear DNA, one of the codons specifying the amino acid tyrosine is UAC. If one nucleotide is changed and the codon is mutated to UAG, what type of mutation will occur?
- Silent mutation
- Missense mutation
- Nonsense mutation
- Frameshift mutation
-
Which of the following is NOT used by eukaryotes to increase the transcription of a gene?
- Gene duplication
- Histone acetylation
- DNA methylation
- Enhancers
- When trypsin converts chymotrypsinogen to chymotrypsin, some molecules of chymotrypsin bind to a repressor, which in turn binds to an operator region and prevents further transcription of trypsin. This is most similar to which of the following operons?
- trp operon during lack of tryptophan
- trp operon during abundance of tryptophan
- lac operon during lack of lactose
- lac operon during abundance of lactose
- Which of the following RNA molecules or proteins is NOT found in the spliceosome during intron excision?
- snRNA
- hnRNA
- shRNA
- snRNPs
- A 4-year-old toddler with cystic fibrosis (CF) is seen by his physician for an upper respiratory infection. Prior genetic testing has shown that there has been a deletion of three base pairs in exon 10 of the CFTRgene that affects codons 507 and 508. The nucleotide sequence in this region for normal and mutant alleles is shown below (Xdenotes the missing nucleotide):
Codon Number 506 507 508 509 510 511 Normal gene (coding strand) ATC ATC TTT GGT GTT TCC
Mutant gene (coding strand) ATC ATX XXT GGT GTT TCC
What effect will this mutation have on the amino acid sequence of the protein encoded by the CFTRgene? (Note: Refer to Figure 7.6 for a genetic code table.)
- Deletion of a phenylalanine residue with no change in the C-terminus sequence
- Deletion of a leucine residue with no change in the C-terminus sequence
- Deletion of a phenylalanine residue with a change in the C-terminus sequence
- Deletion of a leucine residue with a change in the C-terminus sequence
- A gene encodes a protein with 150 amino acids. There is one intron of 1000 base pairs (bp), a 5′-untranslated region of 100 bp, and a 3′-untranslated region of 200 bp. In the final mRNA, about how many bases lie between the start AUG codon and final termination codon?
- 150
- 450
- 650
- 1750
- Peptidyl transferase connects the carboxylate group of one amino acid to the amino group of an incoming amino acid. What type of linkage is created in this peptide bond?
- Ester
- Amide
- Anhydride
- Ether
- A eukaryotic cell has been found to exhibit a truncation mutation that creates an inactive RNA polymerase I enzyme. Which type of RNA will be affected by this inactivation?
- rRNA
- tRNA
- snRNA
- hnRNA
- You have just sequenced a piece of DNA that reads as follows:
5′—TCTTTGAGACATCC—3′
What would the base sequence of the mRNA transcribed from this DNA be?
- 5′—AGAAACUCUGUAGG—3′
- 5′—GGAUGUCUCAAAGA—3′
- 5′—AGAAACTCTGTAGG—3′
- 5′—GGATCTCTCAAAGA—3′
- Double-stranded RNA cannot be translated by the ribosome and is marked for degradation in the cell. Which of the following strands of RNA would prevent mature mRNA in the cytoplasm from being translated?
- Identical mRNA to the one produced
- Antisense mRNA to the one produced
- mRNA with thymine substituted for uracil
- Sense mRNA to the one produced
Answer Key
- D
- C
- A
- C
- A
- C
- C
- B
- C
- A
- B
- B
- A
- B
- B
Chapter 7: RNA and the Genetic Code
CHAPTER 7
RNA AND THE GENETIC CODE
In This Chapter
7.1 The Genetic Code
Types of RNA
Codons
Mutations
7.2 Transcription
Mechanism of Transcription
Posttranscriptional Processing
7.3 Translation
The Ribosome
Mechanism of Translation
Posttranslational Processing
7.4 Control of Gene Expression in Prokaryotes
Operon Structure
Inducible Systems
Repressible Systems
7.5 Control of Gene Expression in Eukaryotes
Transcription Factors
Gene Amplification
Regulation of Chromatin Structure
Concept Summary
CHAPTER PROFILE
The content in this chapter should be relevant to about 9% of all questions about biochemistry on the MCAT.
This chapter covers material from the following AAMC content categories:
1B: Transmission of genetic information from the gene to the protein
5D: Structure, function, and reactivity of biologically-relevant molecules
Introduction
Hepatitis C virus (HCV) continues to be a major cause of cirrhosis and liver failure in the United States. Usually associated with intravenous drug use, hepatitis C causes ongoing damage and inflammation in the liver, leading to the formation of scar tissue that replaces the normal cells of the organ. Over time, this buildup of scar tissue makes the liver unable to keep up with the metabolic demands of the body, and liver failure ensues. To fight this virus, infected hepatocytes release interferon, a peptide signal that—as the name suggests—interferes with viral replication. Because viruses must hijack the host cell’s machinery to replicate, one way the body can limit the spread of the virus is by shutting off the processes of transcription and translation. Interferon not only curtails these processes in virally infected cells, but also induces the production of RNase L, which cleaves RNA in cells to further reduce the ability of the virus to replicate. Coupled with other immune defenses, interferon thus serves as an efficient mechanism to protect the body from viral pathogens.
Even in normal, healthy cells, the first step in expressing genetic information is transcription of the information in the base sequence of a double-stranded DNA molecule to form a single-stranded molecule of RNA. The second step is translating that nucleotide sequence into a protein. Not every cell, though, expresses every gene product, and control of gene expression leads to the differentiation of the totipotent zygote into all of the tissues of the body. In this chapter, we will discuss the process through which proteins are produced along with the controls that modulate each step of the path.
7.1 The Genetic Code
LEARNING OBJECTIVES
After Chapter 7.1, you will be able to:
- Differentiate between three different types of RNA: mRNA, tRNA, and rRNA
- Transcribe a DNA sequence like "GAATTCG" into its mRNA conjugate
- Define the concepts of wobble and degeneracy
- Identify the translation outcomes of key codons, including AUG, UAG, UAA, and UGA
- Predict the likely impact of different mutation types on the resulting peptide
An organism must be able to store and preserve its genetic information, pass that information along to future generations, and express that information as it carries out all the processes of life. We know that DNA and RNA share the same language: they both code using nitrogenous bases. Proteins, however, are composed of amino acids, which constitute a different language altogether. Therefore, we use the genetic code to translate this genetic information into proteins.
While nucleotides play a crucial role in maintaining our genetic identity from generation to generation, it is the proteins they encode that help organisms develop and perform the necessary functions of life. The major steps involved in the transfer of genetic information are illustrated in the central dogma of molecular biology, as shown in Figure 7.1. Classically, a gene is a unit of DNA that encodes a specific protein or RNA molecule, and through transcription and translation, that gene can be expressed. Although this sequence is now complicated by our increased knowledge of the ways in which genes and nucleic acids may be expressed, it is still useful as a general working definition of the processes of DNA replication, transcription, and translation. We have already discussed DNA synthesis, but will continue learning more about gene expression in the rest of this chapter.
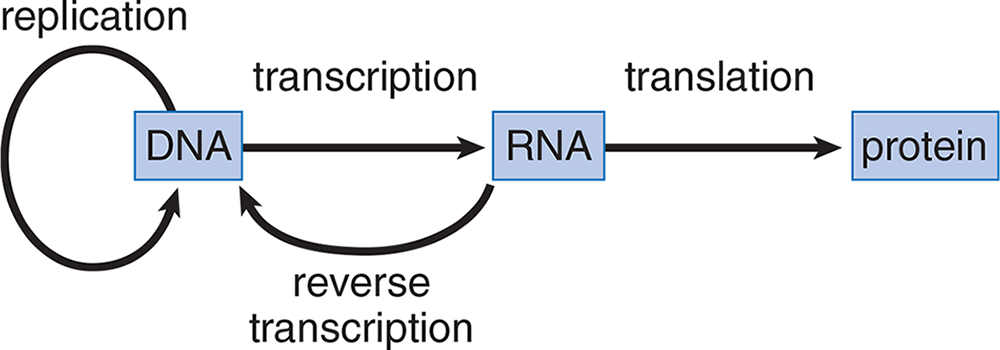
Figure 7.1 The Central Dogma of Molecular Biology
The relationship between the sequence found in double-stranded DNA, single-stranded RNA, and protein is illustrated in Figure 7.2 for a prototypical gene. Messenger RNA is synthesized in the 5′ → 3′ direction and is complementary and antiparallel to the DNA template strand. The ribosome translates the mRNA in the 5′ → 3′ direction, as it synthesizes the protein from the amino terminus (N-terminus) to the carboxy terminus (C-terminus).
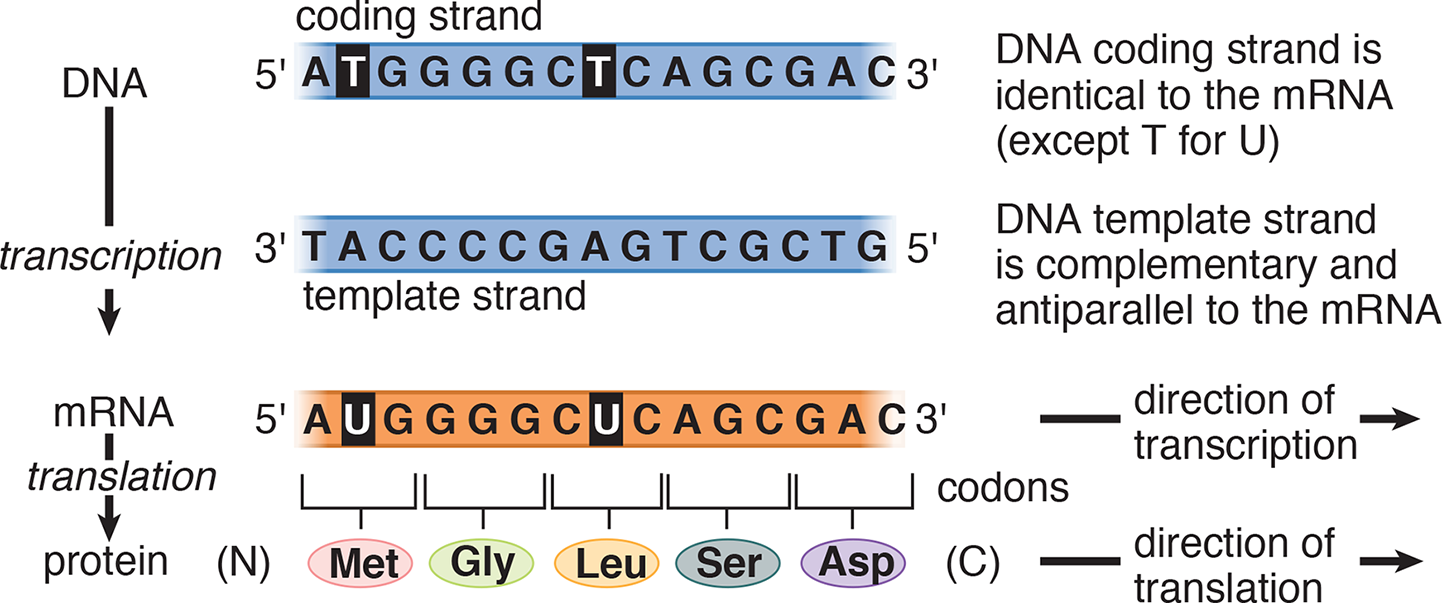
Figure 7.2 Flow of Genetic Information from DNA to Protein
Types of RNA
There are three main types of RNA found in cells: mRNA, tRNA, and rRNA. Each of the main types is described below, but regulatory and specialized forms of RNA are also described later in the chapter.
Messenger RNA (mRNA)
Messenger RNA (mRNA) carries the information specifying the amino acid sequence of the protein to the ribosome. mRNA is transcribed from template DNA strands by RNA polymerase enzymes in the nucleus of cells. Then, mRNA may undergo a host of posttranscriptional modifications prior to its release from the nucleus. mRNA is the only type of RNA that contains information that is translated into protein; to do so, it is read in three-nucleotide segments termed codons. In eukaryotes, mRNA is monocistronic, meaning that each mRNA molecule translates into only one protein product. Thus, in eukaryotes, the cell has a different mRNA molecule for each of the thousands of different proteins made by that cell. In prokaryotes, mRNA may be polycistronic, and starting the process of translation at different locations in the mRNA can result in different proteins. Figure 7.3 shows the difference between monocistronic and polycistronic mRNA. The process of creating mature mRNA will be discussed in the next section of this chapter.
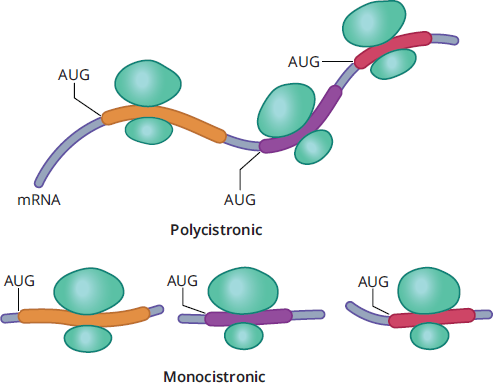
Figure 7.3 Monocistronic and Polycistronic mRNA
KEY CONCEPT
mRNA is the messenger of genetic information. DNA codes for proteins but cannot perform any of the important enzymatic reactions that proteins are responsible for in cells. mRNA takes the information from the DNA to the ribosomes, where creation of the primary protein structure occurs.
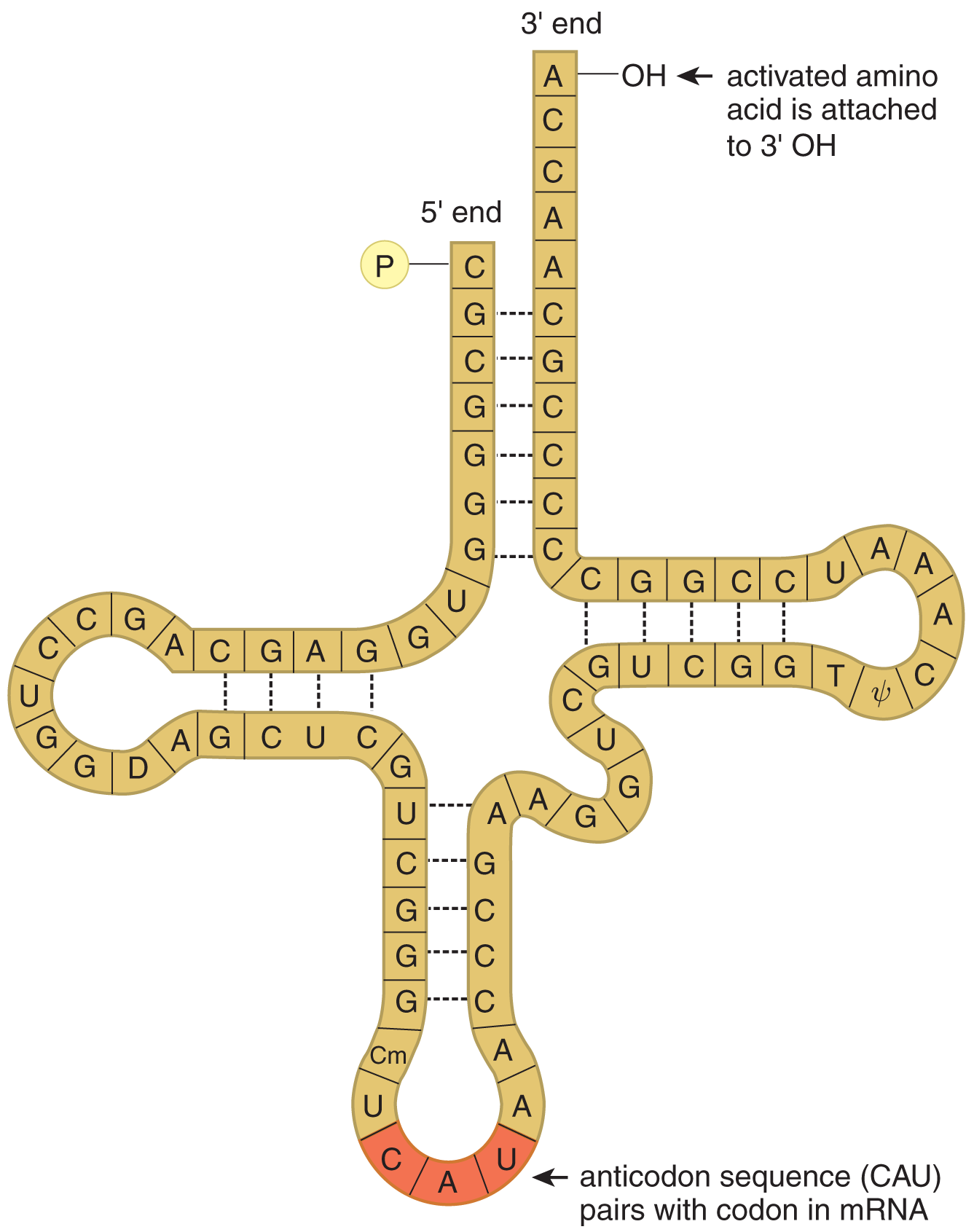
Figure 7.4 The Structure of tRNA
Transfer RNA (tRNA)
Transfer RNA (tRNA) is responsible for converting the language of nucleic acids to the language of amino acids and peptides. Each tRNA molecule contains a folded strand of RNA that includes a three-nucleotide anticodon, as shown in Figure 7.4. This anticodon recognizes and pairs with the appropriate codon on an mRNA molecule while in the ribosome. There are 20 amino acids in eukaryotic proteins, each of which is represented by at least one codon. To become part of a nascent polypeptide in the ribosome, amino acids are connected to a specific tRNA molecule; such tRNA molecules are said to be charged or activated with an amino acid, as shown in Figure 7.5. Mature tRNA is found in the cytoplasm.
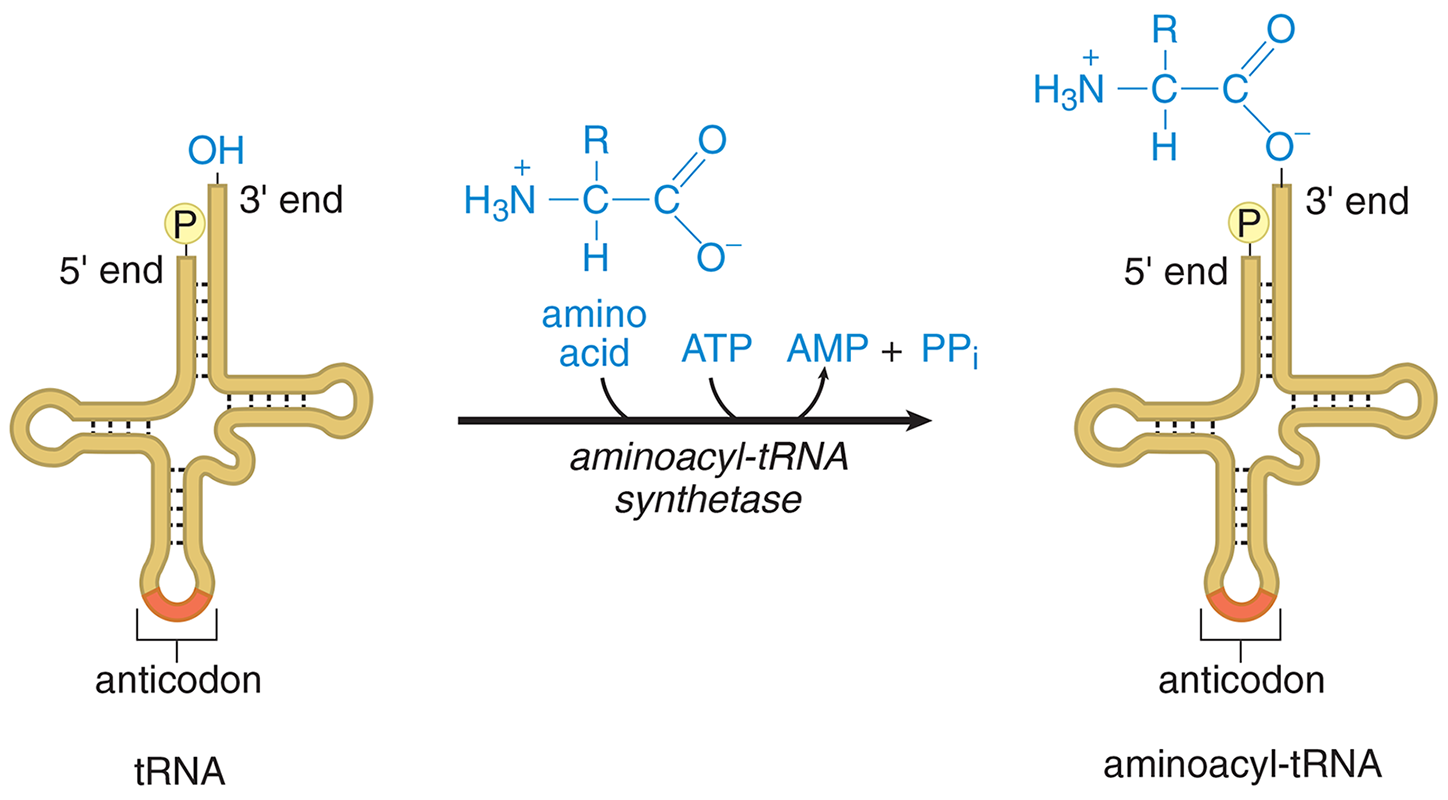
Figure 7.5 Activation of Amino Acid for Protein Synthesis
Each type of amino acid is activated by a different aminoacyl-tRNA synthetase that requires two high-energy bonds from ATP, implying that the attachment of the amino acid is an energy rich bond. The aminoacyl-tRNA synthetase transfers the activated amino acid to the 3′ end of the correct tRNA. Each tRNA has a CCA nucleotide sequence where the amino acid binds. The high-energy aminoacyl-tRNA bond will be used to supply the energy needed to create a peptide bond during translation.
Ribosomal RNA (rRNA)
Ribosomal RNA (rRNA) is synthesized in the nucleolus and functions as an integral part of the ribosomal machinery used during protein assembly in the cytoplasm. Many rRNA molecules function as ribozymes; that is, enzymes made of RNA molecules instead of peptides. rRNA helps catalyze the formation of peptide bonds and is also important in splicing out its own introns within the nucleus. The complex structure of the ribosome is described later in this chapter.
Codons
If a gene sequence is a “sentence” describing a protein, then its basic unit is a three-letter “word” known as the codon, which is translated into an amino acid. Genetic code tables, such as the one in Figure 7.6, serve as an easy way to determine the amino acid that is translated from each mRNA codon. Each codon consists of three bases; thus, there are 64 codons. Note how all codons are written in the 5′ → 3′ direction, and the code is unambiguous, in that each codon is specific for one and only one amino acid.
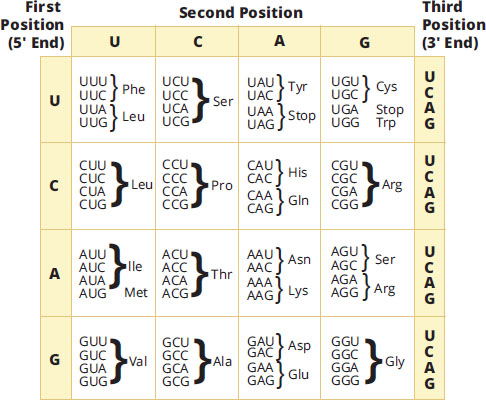
Figure 7.6 The Genetic Code
Note that 61 of the codons code for one of the 20 amino acids, while three codons encode for the termination of translation. This code is universal across species (although there are some exceptions in the mitochondria that are not necessary to know for the MCAT).
KEY CONCEPT
Each codon represents only one amino acid; however, most amino acids are represented by multiple codons.
During translation, the codon of the mRNA is recognized by a complementary anticodon on a transfer RNA (tRNA). The anticodon sequence allows the tRNA to pair with the codon in the mRNA. Because base-pairing is involved, the orientation of this interaction will be antiparallel. For example, the aminoacyl tRNA Ile-tRNAIle has an anticodon sequence 5′—GAU—3′, allowing it to pair with the isoleucine codon 5′—AUC—3′, as seen in Figure 7.7.
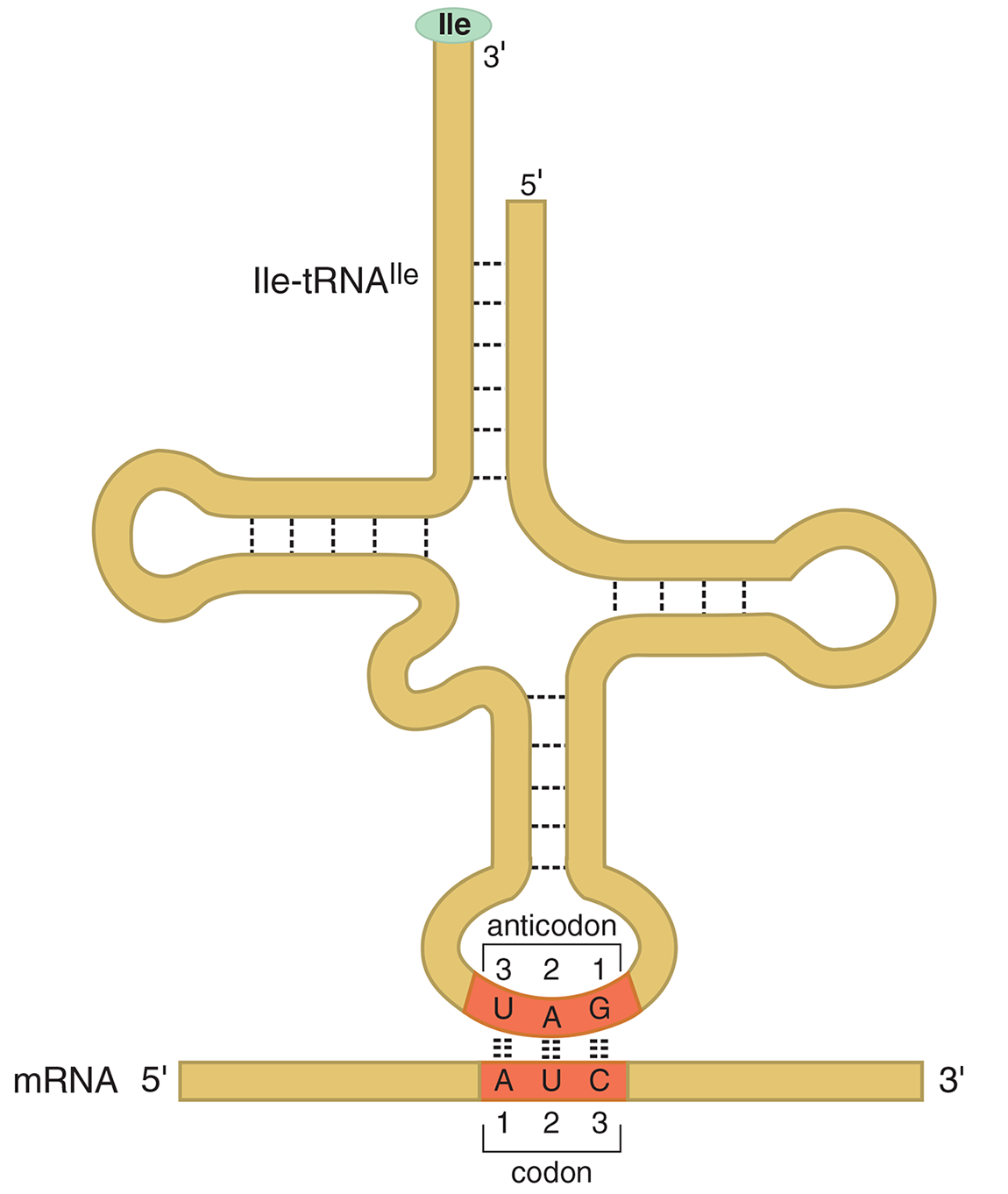
Figure 7.7 Base Pairing of an Aminoacyl-tRNA with a Codon in mRNA
Every preprocessed eukaryotic protein starts with the exact same amino acid: methionine. Because every protein begins with methionine, the codon for methionine (AUG) is considered the start codon for translation of the mRNA into protein. There are also three codons that encode for termination of protein translation; there are no charged tRNA molecules that recognize these codons, which leads to the release of the protein from the ribosome. The three stop codons are UGA, UAA, and UAG.
MNEMONIC
Stop codons:
- UAA—U Are Annoying
- UGA—U Go Away
- UAG—U Are Gone
Mutations
Degeneracy and Wobble
The genetic code is degenerate because more than one codon can specify a single amino acid. In fact, all amino acids, except for methionine and tryptophan, are encoded by multiple codons. Referring back to Figure 7.6, we can see that for the amino acids with multiple codons, the first two bases are usually the same, and the third base in the codon is variable. We refer to this variable third base in the codon as the wobble position. Wobble is an evolutionary development designed to protect against mutations in the coding regions of our DNA. Mutations in the wobble position tend to be called silent or degenerate, which means there is no effect on the expression of the amino acid and therefore no adverse effects on the polypeptide sequence. The amino acid glycine, for example, requires that only the first two nucleotides of the codon be GG. The third nucleotide could be A, C, G, or U, and the amino acid composition of the protein would remain the same. The sequence that codes for protein can have genetic variance. If the variant is below 1% of allele frequency, it is generally termed a mutation. If the variant occurs in greater than 1% of the population, then it is termed as a polymorphism.
KEY CONCEPT
The degeneracy of the genetic code allows for mutations in DNA that do not always result in altered protein structure or function. Usually, a mutation within an intron will also not change the protein sequence because introns are cleaved out of the mRNA transcript prior to translation.
Missense and Nonsense Mutations
If a mutation occurs and it affects one of the nucleotides in a codon, it is known as a point mutation. Although we’ve already discussed the silent point mutation in the wobble position, other point mutations can have a severe detrimental effect depending on where the mutation occurs in the genome. Because these point mutations can affect the primary amino acid sequence of the protein, they are called expressed mutations. Expressed point mutations fall into two categories: missense and nonsense.
- Missense mutation—a mutation where one amino acid substitutes for another
- Nonsense mutation—a mutation where the codon now encodes for a premature stop codon (also known as a truncation mutation)
Frameshift Mutations
The three nucleotides of a codon are referred to as the reading frame. Point mutations occur when one nucleotide is changed, but a frameshift mutation occurs when some number of nucleotides are added to or deleted from the mRNA sequence. Insertion or deletion of nucleotides will shift the reading frame, usually resulting in changes in the amino acid sequence or premature truncation of the protein. The effects of frameshift mutations are typically more serious than point mutations, although it is heavily dependent on where within the DNA sequence the mutation actually occurred. A synopsis of the different types of mutations can be found in Figure 7.8.
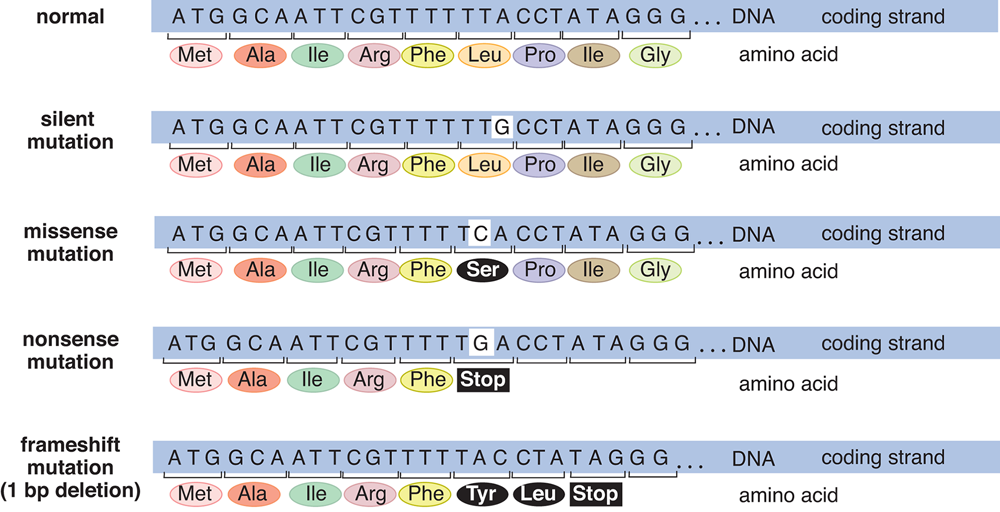
Figure 7.8 Some Common Types of Mutations in DNA
REAL WORLD
Cystic fibrosis is most commonly caused by a frameshift mutation: a deletion at codon 508 in the polypeptide chain of the CFTRchloride channel gene. The subsequent loss of a phenylalanine residue at this position results in a defective chloride ion channel. This altered protein never reaches the cell membrane, leading to blocked passage of salt and water into and out of cells. As a result of this blockage, cells that line the passageways of the lungs, pancreas, and other organs produce an abnormally thick, sticky mucus that traps bacteria, increasing the likelihood of infection in patients.
MCAT CONCEPT CHECK 7.1:
Before you move on, assess your understanding of the material with these questions.
- What are the roles of the three main types of RNA?
- mRNA: ______________________________
- tRNA: ______________________________
- rRNA: ______________________________
- The three-base sequences listed below are DNA sequences. Using Figure 7.6, which amino acid is encoded by each of these sequences, after transcription and translation?
- GAT: ______________________________
- ATT: ______________________________
- CGC: ______________________________
- CCA: ______________________________
- Which mRNA codon is the start codon, and what amino acid does it code for? Which mRNA codons are the stop codons?
- Start codon: ________; codes for: ________
- Stop codons: __________
- What is wobble, and what role does it serve?
________________________________________
- For each of the mutations listed below, what changes in DNA sequence are observed, and what effect do they have on the encoded peptide?
Type of Mutation Change in DNA Sequence Effect on Encoded Protein Silent (degenerate) Missense Nonsense Frameshift
7.2 Transcription
LEARNING OBJECTIVES
After Chapter 7.2, you will be able to:
- Explain how each of the eukaryotic RNA polymerases (I, II, and III) impacts transcription
- Identify where RNA polymerase would bind to start transcription on a DNA strand
- Determine the mRNA that results from a given hnRNA molecule:
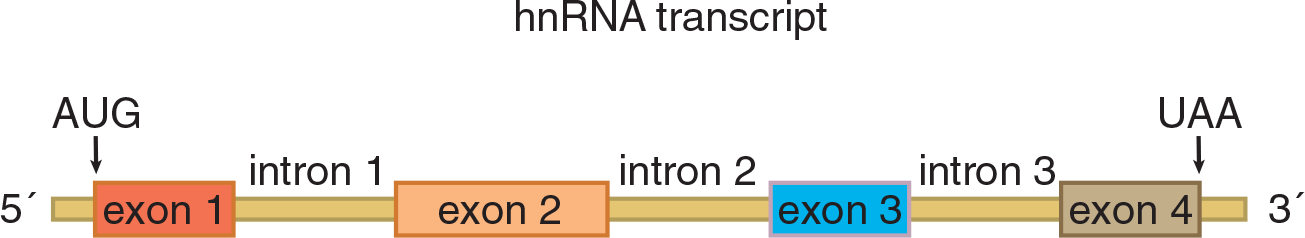
Although DNA contains the actual coding sequence for a protein, the machinery to generate that protein is located in the cytoplasm. DNA cannot leave the nucleus, as it will be quickly degraded, so it must use RNA to transmit genetic information. The creation of mRNA from a DNA template is known as transcription, and while mRNA is the only type of RNA that carries information from DNA directly, there are many other types of RNA that exist, two of which will play important roles during protein translation: transfer RNA (tRNA) and ribosomal RNA (rRNA).
Mechanism of Transcription
Transcription proceeds via three general steps:
Initiation: Several enzymes, including helicase and topoisomerase, are involved in unwinding the double-stranded DNA and preventing formation of super coils, as described in Chapter 6 of MCAT Biochemistry Review. This step is important in allowing the transcriptional machinery access to the DNA and the particular gene of interest. RNA is synthesized by a DNA-dependent RNA polymerase; RNA polymerase locates genes by searching for specialized DNA regions known as promoter regions, shown in Figure 7.9. In eukaryotes, RNA polymerase II is the main player in transcribing mRNA, and one of its major binding sites in the promoter region is known as the TATA box, named for its high concentration of thymine and adenine bases. Transcription factors help the RNA polymerase locate and bind to this promoter region of the DNA, helping to establish where transcription will start. Unlike DNA polymerase III, which was reviewed during DNA replication, RNA polymerase does not require a primer to start generating a transcript.
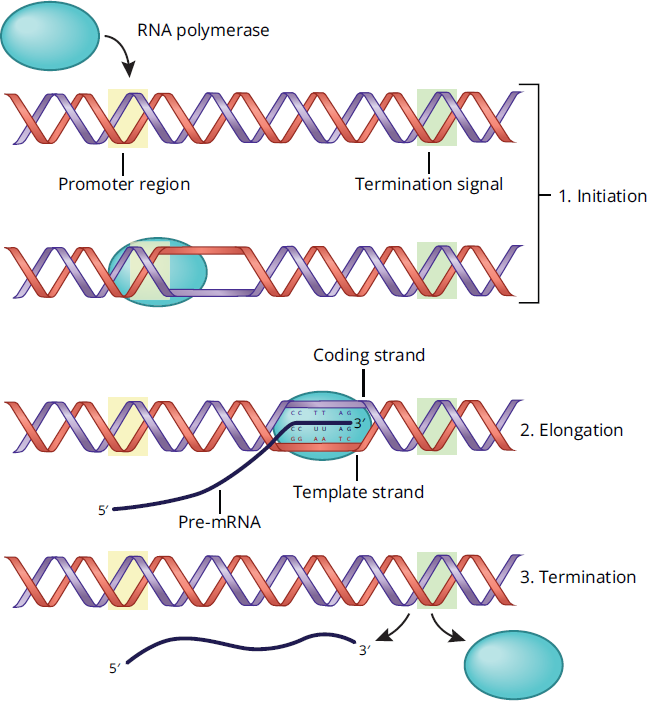
Figure 7.9 Steps in Transcription
Elongation: RNA polymerase travels along the template strand (also known as the antisense strand) in the 3′ → 5′ direction, which allows for the construction of newly transcribed mRNA in the 5′ → 3′ direction. Unlike DNA polymerase, RNA polymerase does not have proofreading capabilities, so the synthesized transcript will not be edited. The coding strand (or sense strand) of DNA is not used as a template during transcription. Because the coding strand is also complementary to the template strand, it is identical to the mRNA transcript except that all the thymine nucleotides in DNA have been replaced with uracil in the mRNA molecule.
KEY CONCEPT
Transcription is subject to the 5′ → 3′ rule, just like DNA synthesis. Synthesis of nucleic acids always occurs in the 5′ → 3′ direction.
Termination: Transcription will continue along the DNA coding region until the RNA polymerase reaches a termination sequence or stop signal, which results in the termination of transcription. The DNA double helix then re-forms, and the primary transcript formed is termed heterogeneous nuclear RNA (hnRNA), sometimes also known as pre-mRNA. mRNA is derived from hnRNA via posttranscriptional modifications, as described below.
In eukaryotes, there are three types of RNA polymerases, but only one is involved in the transcription of mRNA:
- RNA polymerase I is located in the nucleolus and synthesizes rRNA.
- RNA polymerase II is located in the nucleus and synthesizes hnRNA (pre-processed mRNA) and some small nuclear RNA (snRNA).
- RNA polymerase III is located in the nucleus and synthesizes tRNA and some rRNA.
In the vicinity of a gene, a numbering system is used to identify the location of important bases in the DNA strand, as shown in Figure 7.10. The first base transcribed from DNA to RNA is defined as the +1 base of that gene region. Bases to the left of this start point (upstream, or toward the 5′ end) are given negative numbers: –1, –2, –3, and so on. Bases to the right (downstream, or toward the 3′ end) are denoted with positive numbers: +2, +3, +4, and so on. Thus, no nucleotide in the gene is numbered 0. The TATA box, where RNA polymerase II binds, usually falls around –25.
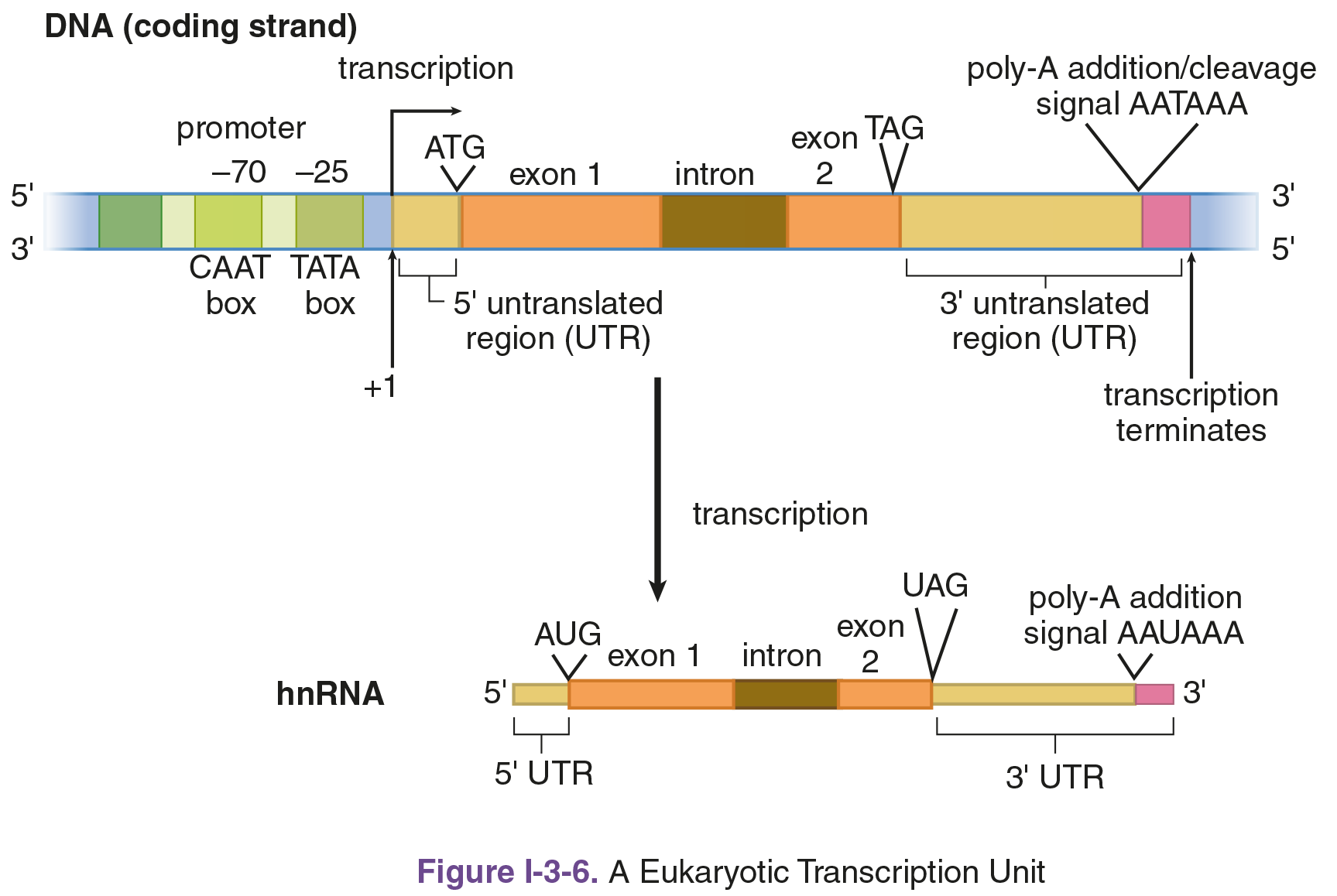
Figure 7.10 Transcription of DNA to hnRNA
Posttranscriptional Processing
Before the hnRNA can leave the nucleus and be translated to protein, it must undergo three specific processes to allow it to interact with the ribosome and survive the conditions of the cytoplasm, as demonstrated in Figure 7.11. You can think of the nucleus as the happy home of the cell; the DNA strands are the caregivers, and the hnRNA is their child. The child must mature in order to survive.
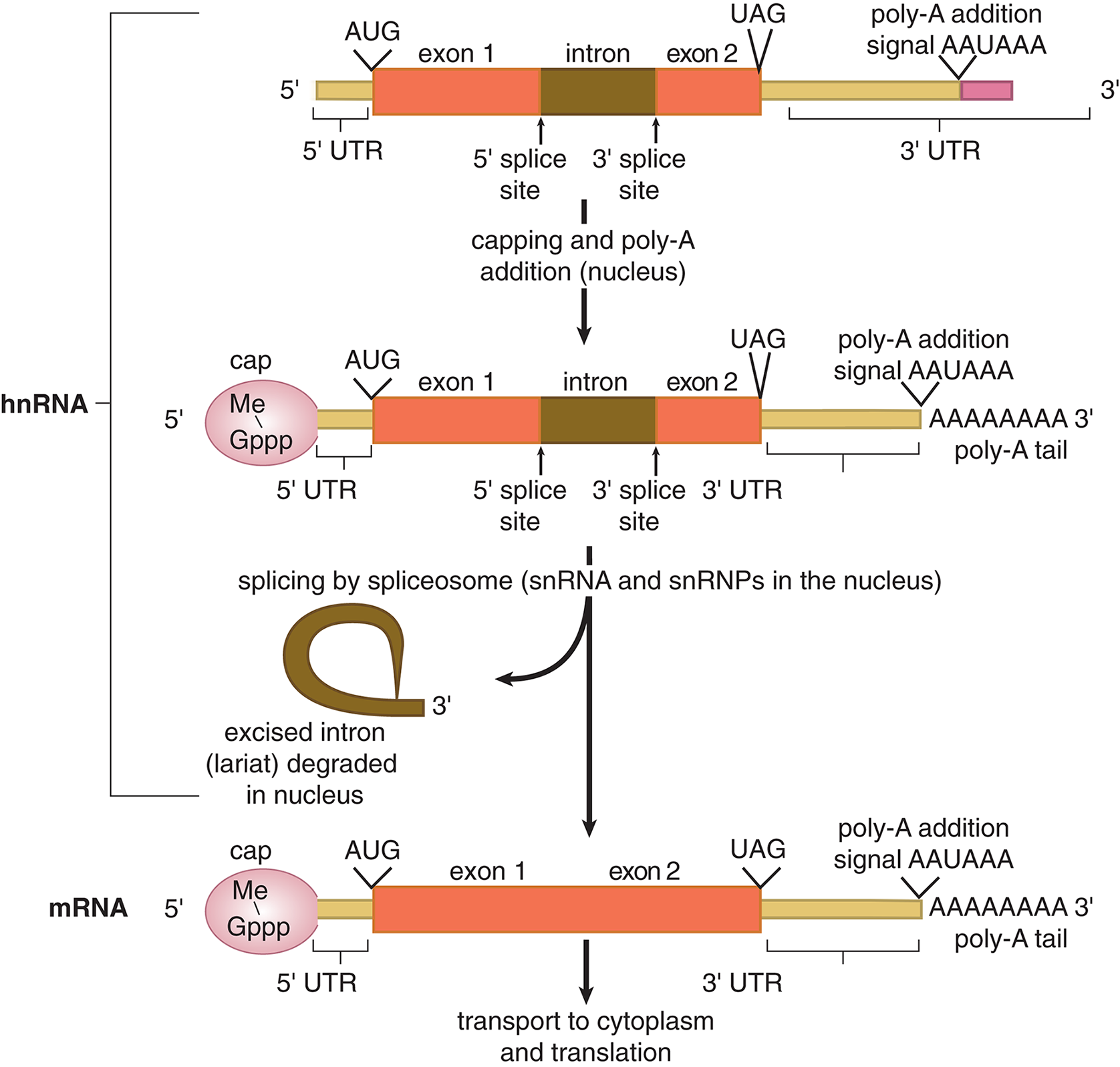
Figure 7.11 Processing Eukaryotic hnRNA to Form mRNA
KEY CONCEPT
The MCAT commonly tests post-transcriptional processing:
- Intron/exon splicing
- 5′ cap
- 3′ poly-A tail
Splicing: Introns and Exons
Maturation of the hnRNA includes splicing of the transcript to remove noncoding sequences (introns) and ligate coding sequences (exons) together. Splicing is accomplished by the spliceosome. In the spliceosome, small nuclear RNA (snRNA) molecules couple with proteins known as small nuclear ribonucleoproteins (also known as snRNPs, or “snurps”). The snRNP/snRNA complex recognizes both the 5′ and 3′ splice sites of the introns. These noncoding sequences are excised in the form of a lariat (lasso-shaped structure) and then degraded.
MNEMONIC
In trons stay in the nucleus; ex ons will ex it the nucleus as part of the mRNA.
The evolutionary function of introns in eukaryotic cells is not currently well-understood; however, scientists hypothesize that introns play an important role in the regulation of cellular gene expression levels and in maintaining the size of our genome. The existence of introns has also been hypothesized to allow for rapid protein evolution. Many eukaryotic proteins share peptide sequences in common, suggesting that the genes encoding for these particular peptides may employ a modular function; that is, they contain standard sequences that can be swapped in and out, depending on the needs of the cell.
5′ Cap
At the 5′ end of the hnRNA molecule, a 7-methylguanylate triphosphate cap is added. The cap is actually added during the process of transcription and is recognized by the ribosome as the binding site. It also protects the mRNA from degradation in the cytoplasm.
3′ Poly-A Tail
A polyadenosyl (poly-A) tail is added to the 3′ end of the mRNA transcript and protects the message against rapid degradation. It is composed of adenine bases. Think of the poly-A tail as a fuse for a “time bomb” for the mRNA transcript: as soon as the mRNA leaves the nucleus, it will start to get degraded from its 3′ end. The longer the poly-A tail, the more time the mRNA will be able to survive before being digested in the cytoplasm. The poly-A tail also assists with export of the mature mRNA from the nucleus.
At this point, when only the exons remain and the cap and tail have been added, the cell has created the mature mRNA that can now be transported into the cytoplasm for protein translation. Untranslated regions of the mRNA (UTRs) will still exist at the 5′ and 3′ edges of the transcript because the ribosome initiates translation at the start codon (AUG) and will end at a stop codon (UAA, UGA, UAG).
For some genes in eukaryotic cells, however, the primary transcript of hnRNA may be spliced together in different ways to produce multiple variants of proteins encoded by the same original gene. This process is known as alternative splicing, and it is illustrated in Figure 7.12. By utilizing alternative splicing, an organism can make many more different proteins from a limited number of genes. For reference, humans are estimated to make at least 100,000 proteins, but the number of human genes is only about 20,000–25,000. Don’t worry about memorizing these numbers, though; they are constantly changing with new research. Alternative splicing is also known to function in the regulation of gene expression, in addition to generating protein diversity.
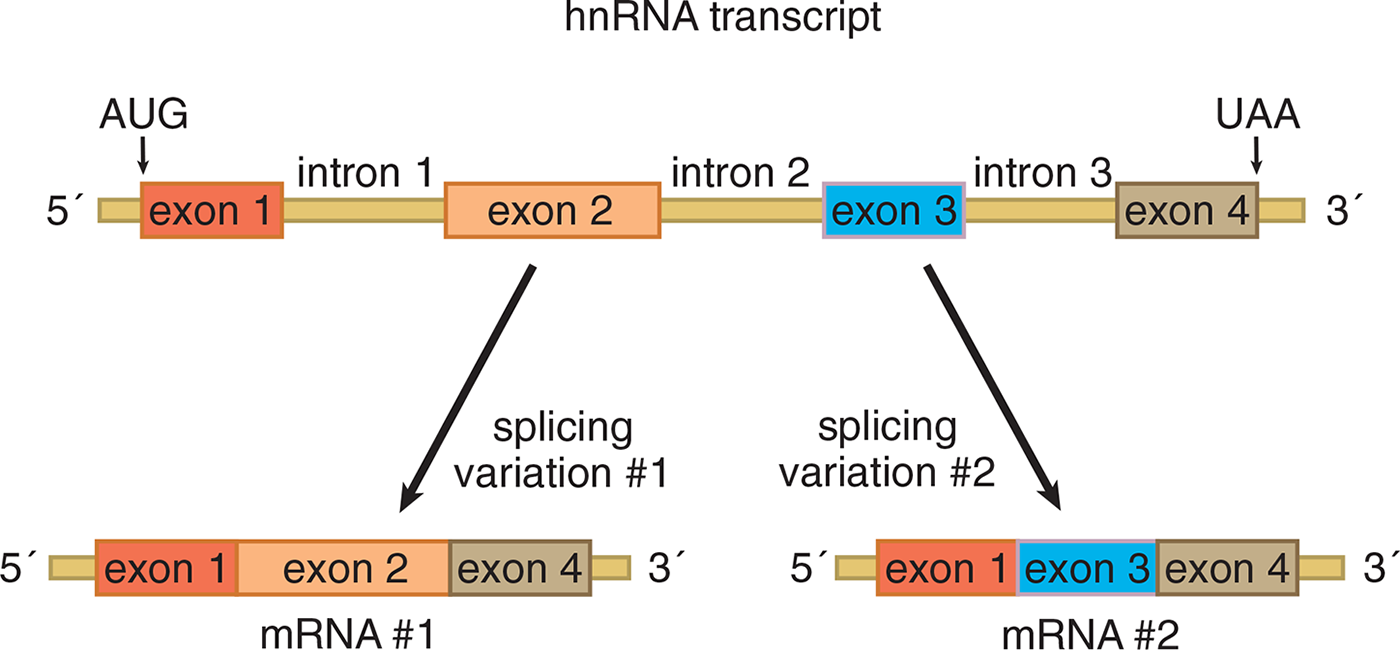
Figure 7.12 Alternative Splicing of Eukaryotic hnRNA to Produce Different Proteins
REAL WORLD
Mutations in splice sites can lead to abnormal proteins. For example, mutations that interfere with proper splicing of β-globulin mRNA are responsible for some cases of β-thalassemia, a group of blood disorders that hinder the production and efficacy of hemoglobin in the blood. Splice site mutations are one of the few mutations in noncoding DNA that may still have an effect on the translated protein.
MCAT CONCEPT CHECK 7.2:
Before you move on, assess your understanding of the material with these questions.
- What is the role of each eukaryotic RNA polymerase?
- RNA polymerase I: ______________________________________
- RNA polymerase II: _____________________________________
______________________________________
- RNA polymerase III:
- When starting transcription, where does RNA polymerase bind?
_____________________________
- What are the three major posttranscriptional modifications that turn hnRNA into mature mRNA?
- ____________________________________
- ____________________________________
- ____________________________________
- What is alternative splicing, and what does it accomplish?
_______________________________________
7.3 Translation
LEARNING OBJECTIVES
After Chapter 7.3, you will be able to:
- Describe the steps of translation: initiation, elongation, and termination
- Distinguish different types of posttranslational modifications, such as phosphorylation and glycosylation
- Explain the role of the functional sites in a ribosome:
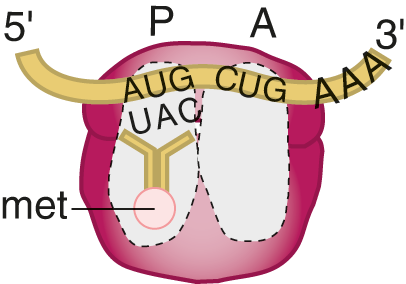
Once the mRNA transcript is created and processed, it can exit the nucleus through nuclear pores. Once in the cytoplasm, mRNA finds a ribosome to begin the process of translation—converting the mRNA transcript into a functional protein. Translation is a complex process that requires mRNA, tRNA, ribosomes, amino acids, and energy in the form of GTP.
KEY CONCEPT
Terminology and 5′ → 3′
- DNA → DNA = replication: new DNA synthesized in 5′ → 3′ direction
- DNA → RNA = transcription: new RNA synthesized in 5′ → 3′ direction (template is read 3′ → 5′)
- RNA → protein = translation: mRNA read in 5′ → 3′ direction
The Ribosome
As mentioned earlier, the anticodon of the tRNA binds to the codon on the mature mRNA in the ribosome. The ribosome is composed of proteins and rRNA. In both prokaryotes and eukaryotes, there are large and small subunits; the subunits only bind together during protein synthesis. The structure of the ribosome dictates its main function, which is to bring the mRNA message together with the charged aminoacyl-tRNA complex to generate the protein. There are three binding sites in the ribosome for tRNA: the A site (aminoacyl), P site (peptidyl), and E site (exit). These are described further in the section on translation below.
Eukaryotic ribosomes contain four strands of rRNA, designated the 28S, 18S, 5.8S, and the 5S rRNAs; the “S” values indicate the size of the strand. The genes for some of the rRNAs (28S, 18S, and 5.8S rRNAs) used to construct the ribosome are found in the nucleolus. RNA polymerase I transcribes the 28S, 18S, and 5.8S rRNAs as a single unit within the nucleolus, which results in a 45S ribosomal precursor RNA. This 45S pre-rRNA is processed to become the 18S rRNA of the 40S (small) ribosomal subunit and the 28S and 5.8S rRNAs of the 60S (large) ribosomal subunit. RNA polymerase III transcribes the 5S rRNA, which is also found in the 60S ribosomal subunit; this process takes place outside of the nucleolus. The ribosomal subunits created are the 60S and 40S subunits; these subunits join during protein synthesis to form the whole 80S ribosome.
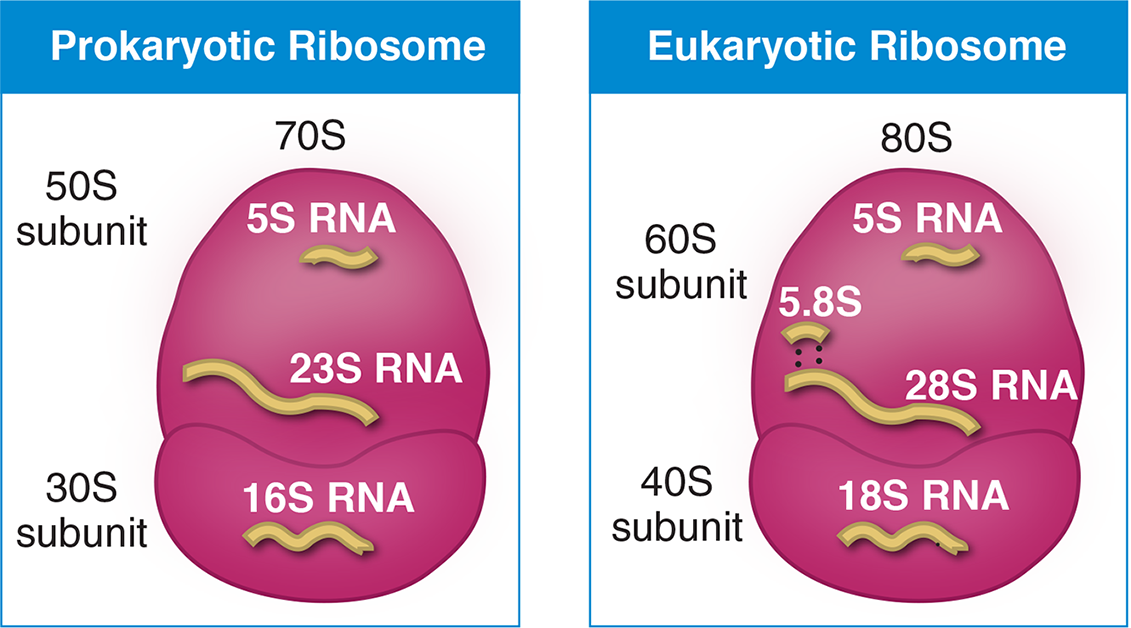
Figure 7.13 The Composition of Prokaryotic and Eukaryotic Ribosomes
In comparison with eukaryotes, prokaryotes have 50S and 30S large and small subunits, which assemble to create the complete 70S ribosome. Note that the “S” value is determined experimentally by studying the behavior of particles in an ultracentrifuge; thus, the numbers of each subunit and each rRNA are not additive because they are based on size and shape, not size alone. The structure of eukaryotic and prokaryotic ribosomes are shown in Figure 7.13.
REAL WORLD
The fact that prokaryotic and eukaryotic ribosomes have slightly different structures is no small fact. This difference allows us to target antibiotics, like macrolides (azithromycin, erythromycin), tetracyclines (doxycycline), vancomycin, and others to bacterial cells with fewer side effects to humans.
Mechanism of Translation
Translation occurs in the cytoplasm in prokaryotes and eukaryotes. In prokaryotes, the ribosomes start translating before the mRNA is complete; in eukaryotes, however, transcription and translation occur at separate times and in separate locations within the cell. The process of translation occurs in three stages, as shown in Figure 7.14: initiation, elongation, and termination. While the names of the three steps are identical to the names of the steps of transcription, the processes involved are quite distinct. In translation, specialized factors for initiation (initiation factors, IF), elongation (elongation factors, EF), and termination (release factors, RF), as well as GTP are required for each step.
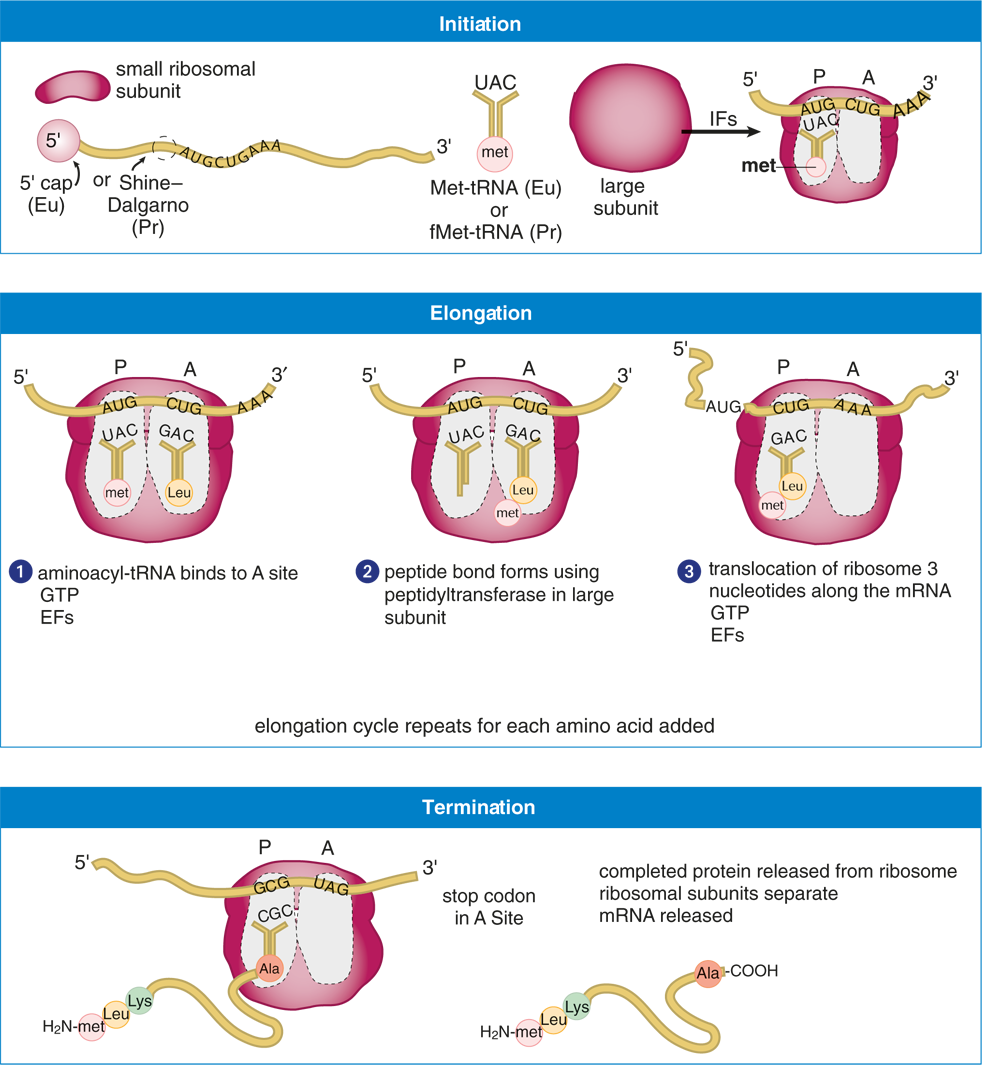
Figure 7.14 Steps in Translation
Initiation
The small ribosomal subunit binds to the mRNA. In prokaryotes, the small subunit binds to the Shine–Dalgarno sequence in the 5′ untranslated region of the mRNA. In eukaryotes, the small subunit binds to the 5′ cap structure. The charged initiator tRNA binds to the AUG start codon through base-pairing with its anticodon within the P site of the ribosome. The initial amino acid in prokaryotes is N-formylmethionine (fMet); in eukaryotes, it’s methionine.
The large subunit then binds to the small subunit, forming the completed initiation complex. This is assisted by initiation factors (IF) that are not permanently associated with the ribosome.
Elongation
Elongation is a three-step cycle that is repeated for each amino acid added to the protein after the initiator methionine. During elongation, the ribosome moves in the 5′ to 3′ direction along the mRNA, synthesizing the protein from its amino (N-) to carboxyl (C-) terminus. The ribosome contains three very important binding sites:
- The A site holds the incoming aminoacyl-tRNA complex. This is the next amino acid that is being added to the growing chain, and is determined by the mRNA codon within the A site.
- The P site holds the tRNA that carries the growing polypeptide chain. It is also where the first amino acid (methionine) binds because it is starting the polypeptide chain. A peptide bond is formed as the polypeptide is passed from the tRNA in the P site to the tRNA in the A site. This requires peptidyl transferase, an enzyme that is part of the large subunit. GTP is used for energy during the formation of this bond.
- The E site (not shown in Figure 7.14) is where the now inactivated (uncharged) tRNA pauses transiently before exiting the ribosome. As the now-uncharged tRNA enters the E site, it quickly unbinds from the mRNA and is ready to be recharged.
MNEMONIC
Order of sites in the ribosome during translation: APE.
Elongation factors (EF) assist by locating and recruiting aminoacyl-tRNA along with GTP, while helping to remove GDP once the energy has been used.
Some eukaryotic proteins contain signal sequences, which designate a particular destination for the protein, as shown in Figure 7.15. For peptides that will be secreted, such as hormones and digestive enzymes, a signal sequence directs the ribosome to move to the endoplasmic reticulum (ER), so that the protein can be translated directly into the lumen of the rough ER. From there, the protein can be sent to the Golgi apparatus and be secreted from a vesicle via exocytosis. Other signal sequences direct proteins to the nucleus, lysosomes, or cell membrane.
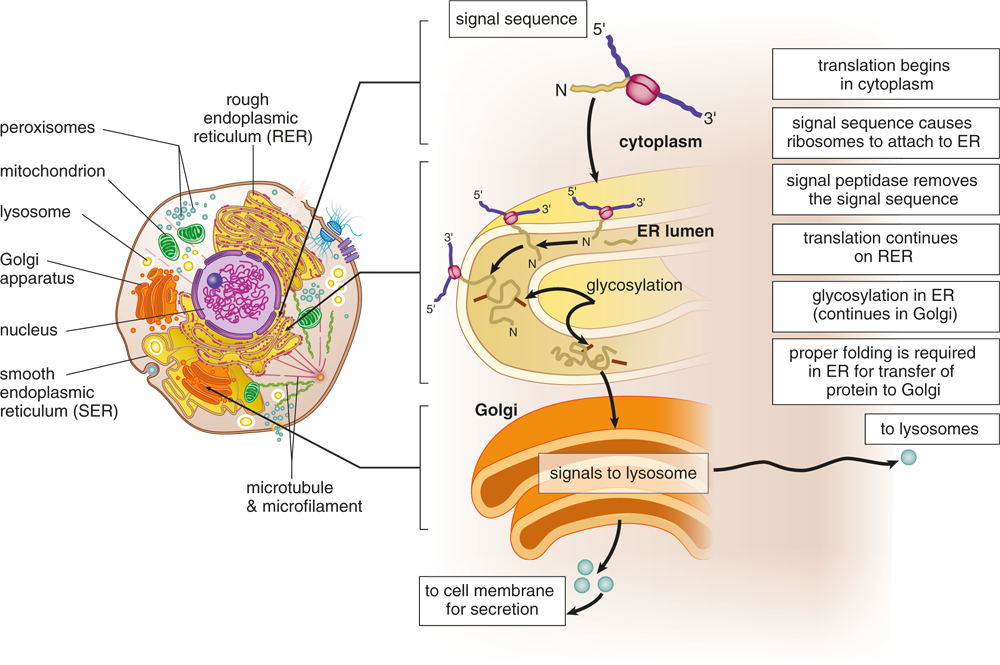
Figure 7.15 Synthesis of Secretory, Membrane, and Lysosomal Proteins
BIOCHEMISTRY GUIDED EXAMPLE WITH EXPERT THINKING
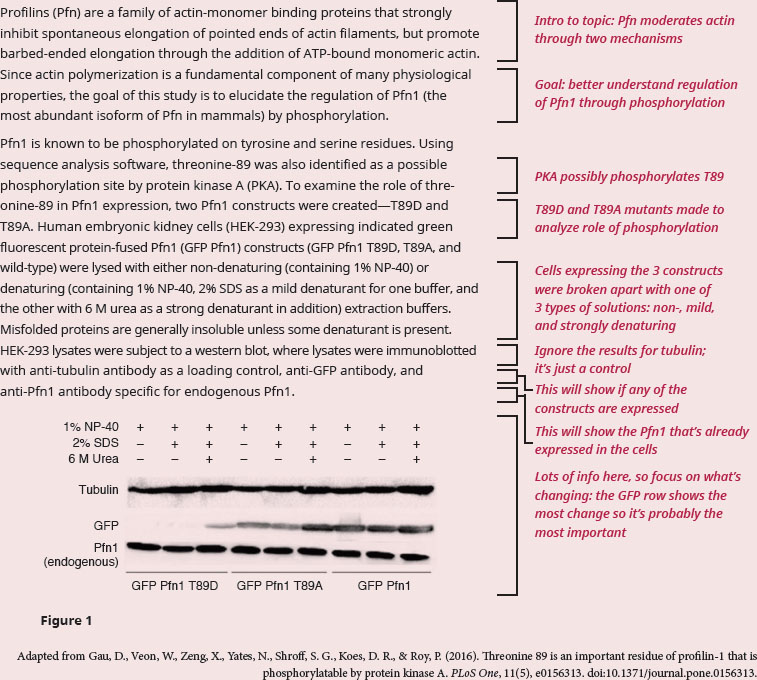
What is the purpose of mutating the potential phosphorylation site to aspartic acid and alanine, respectively? What is the implication for expression of Pfn1?
Since the question is asking for a reason behind the experimental design, our focus should be on understanding the way the experiment was designed. The passage is focused on clarifying how phosphorylation regulates Pfn1. We know from our content background that phosphorylation of an amino acid (specifically serine, threonine, and tyrosine) will turn a polar amino acid into one that is negatively charged. If we didn't remember that, we could also infer it by thinking about the charge on a phosphate group, which is highly negative. We have enough information to answer the first question. Mutating a threonine phosphorylation site to aspartic acid (which is negatively charged) would mimic the site being constantly phosphorylated, while mutating to alanine would mimic the site being constantly dephosphorylated, since alanine residues cannot be phosphorylated.
One possible phosphorylation site, T89, was identified, and three GFP constructs were made—one mimicking phosphorylation (T89D, lanes 1–3), one eliminating phosphorylation (T89A, lanes 4–6), and one unmodified (lanes 7–9). The GFP tag allows the experimenters to see whether the GFP-fused construct is being expressed (whether the protein is being made). We’re told this experimental procedure is a western blot, a technique in which a specific antibody of interest will bind to proteins that were separated on a gel. In western blots, a band appears when the antibody successfully binds to protein. The rows above the gel image display experimental conditions for each of the lanes. All lanes have 1% NP-40, which we’re told is non-denaturing. We should recall that the word denaturing means that a substance changes the 3-D structure of a protein. So, non-denaturing means that NP-40 won't change the shape of the protein by itself. The lanes with 2% SDS are in mild denaturing conditions, and the lanes with 2% SDS and 6 M urea are in strong denaturing conditions. Continuing by that same definition, we know that strong denaturing conditions will unfold and solubilize all proteins, greatly changing their 3-D structure. If we look at the row probing for GFP, we see that there is no band for non- and mild denaturing conditions, but there is a faint band under strong denaturing conditions for only the T89D construct. We can see that the lanes for both the wild type and T89A, regardless of condition, are mostly the same. Finally, recall that phosphorylation is a post-translational event, so a change in the phosphorylation state of the protein would occur after Pfn1 has been translated, and wouldn’t work to affect its expression.
Therefore, we can conclude that mutating the threonine residue can mimic the effect of constantly phosphorylated or dephosphorylated Pfn. Second, since we only see a GFP band when T89D is in the presence of a strong denaturant, phosphorylation of Pfn1 may cause the protein to fold in a way that makes it insoluble, thereby targeting the protein for degradation. Follow-up studies are required to verify the exact mechanism of action by which this occurs. Therefore, because T89D represents the "always phosphorylated" version of Pfn1, and T89D only appears on the gel in strongly denaturing conditions, we can conclude that Pfn1, when phosphorylated, isn't available/expressed. We can thus conclude that phosphorylation of Pfn1 can potentially be used to down-regulate the availability of Pfn1 for usage with actin polymerization.
Termination
When any of the three stop codons moves into the A site, a protein called release factor (RF) binds to the termination codon, causing a water molecule to be added to the polypeptide chain. The addition of this water molecule allows peptidyl transferase and termination factors to hydrolyze the completed polypeptide chain from the final tRNA. The polypeptide chain will then be released from the tRNA in the P site, and the two ribosomal subunits will dissociate.
Posttranslational Processing
The nascent polypeptide chain is subject to posttranslational modifications before it will become a functioning protein, similar to how hnRNA is modified prior to being released from the nucleus. One essential step for the final synthesis of the protein is proper folding. There is a specialized class of proteins called chaperones, the main function of which is to assist in the protein-folding process.
Many proteins are also modified by cleavage events. A common example of this is insulin, which needs to be cleaved from a larger, inactive peptide to achieve its active form. In peptides with signal sequences, the signal sequence must be cleaved if the protein is to enter the organelle and accomplish its function.
In peptides with quaternary structure, subunits come together to form the functional protein. A classic example is hemoglobin, which is composed of two alpha chains and two beta chains.
Other biomolecules may be added to the peptide via the following processes:
- Phosphorylation—addition of a phosphate group (PO42–) by protein kinases to activate or deactivate proteins; phosphorylation in eukaryotes is most commonly seen with serine, threonine, and tyrosine
- Carboxylation—addition of carboxylic acid groups, usually to serve as calcium-binding sites
- Glycosylation—addition of oligosaccharides as proteins pass through the ER and Golgi apparatus to determine cellular destination
- Prenylation—addition of lipid groups to certain membrane-bound enzymes
REAL WORLD
Posttranslational modifications are often important for proper protein functioning. For example, several clotting factors, including prothrombin, require posttranslational carboxylation of some of their glutamic acid residues in order to function properly. Vitamin K is required as a cofactor for these reactions; thus, vitamin K deficiency may result in a bleeding disorder.
MCAT CONCEPT CHECK 7.3:
Before you move on, assess your understanding of the material with these questions.
- What are the three steps of translation?
- __________________________________
- __________________________________
- __________________________________
- What are the roles of each site in the ribosome?
- A site: ______________________________
- P site: ______________________________
- E site: ______________________________
- What are the major posttranslational modifications that occur in proteins?
_______________________________________
7.4 Control of Gene Expression in Prokaryotes
LEARNING OBJECTIVES
After Chapter 7.4, you will be able to:
- Recognize the transcriptional controls on key operons such as the lacand trpoperons
- Differentiate between positive and negative control systems
- Explain the role of the different sections of a standard operon
An organism’s DNA encodes all of the RNA and protein molecules required to construct its cells. Yet organisms are able to differentially express their genes to make cell-specific products necessary for cellular development at specific times. In the next section, we’ll look at these processes in eukaryotic cells; for now, we’ll focus on the regulatory processes governing gene expression in prokaryotes—rules that are necessary in determining which subset of genes are selectively expressed or silenced in the prokaryotic cell.
Operon Structure
The simplest example of an on–off switch that regulates gene expression levels in prokaryotes was discovered in *E. coli**,which regulates the expression of many genes according to food sources that are available in the environment. For example, five genes inE. coliencode for enzymes that manufacture the amino acid tryptophan, and these are arranged in a cluster on the chromosome. By sharing a single common promoter region on the DNA sequence, these genes are transcribed as a group. This type of structure is called an operon—a cluster of genes transcribed as a single mRNA; this particular cluster inE. coliis known as thetrp* operon. Operons are incredibly common in the prokaryotic cell.
The Jacob–Monod model is used to describe the structure and function of operons. In this model, operons contain structural genes, an operator site, a promoter site, and a regulator gene, as shown in Figure 7.16. The structural gene codes for the protein of interest. Upstream of the structural gene is the operator site, a nontranscribable region of DNA that is capable of binding a repressor protein. Further upstream is the promoter site, which is similar in function to promoters in eukaryotes: it provides a place for RNA polymerase to bind. Furthest upstream is the regulator gene, which codes for a protein known as the repressor. There are two types of operons: inducible systems and repressible systems.
KEY CONCEPT
Operons include both inducible and repressible systems, and offer a simple on–off switch for gene control in prokaryotes.
Inducible Systems
In inducible systems, the repressor is bonded tightly to the operator system and thereby acts as a roadblock. RNA polymerase is unable to get from the promoter to the structural gene because the repressor is in the way. Such systems—in which the binding of a protein reduces transcriptional activity—are called negative control mechanisms. To remove that block, an inducer must bind the repressor protein so that RNA polymerase can move down the gene, as shown in Figure 7.16. Inducible systems operate on a principle analogous to competitive inhibition for enzyme activity: as the concentration of the inducer increases, it will pull more copies of the repressor off of the operator region, freeing up those genes for transcription. This system is useful because it allows gene products to be produced only when they are needed.
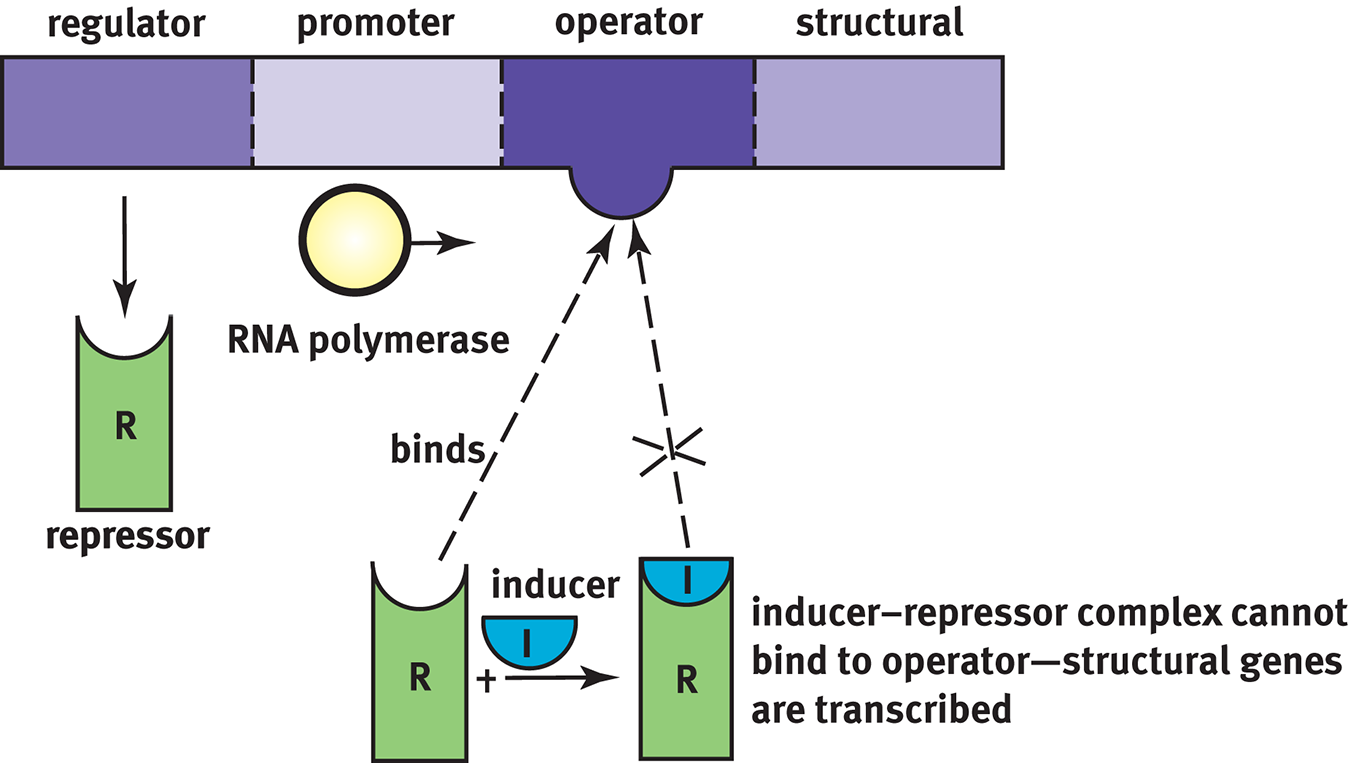
Figure 7.16 Inducible Systems Allow for gene transcription only when an inducer is present to bind the otherwise present repressor protein
A classic example of an inducible system is the lac operon, which contains the gene for lactase, as demonstrated in Figure 7.17. Bacteria can digest lactose, but it is more energetically expensive than digesting glucose. Therefore, bacteria only want to use this option if lactose is high and glucose is low. The lac operon is induced by the presence of lactose; thus, these genes are only transcribed when it is useful to the cell.
The lac operon is assisted by binding of the catabolite activator protein (CAP). CAP is a transcriptional activator used by E. coli when glucose levels are low to signal that alternative carbon sources should be used. Falling levels of glucose cause an increase in the signaling molecule cyclic AMP (cAMP), which binds to CAP. This induces a conformational change in CAP that allows it to bind the promoter region of the operon, further increasing transcription of the lactase gene. Such systems—in which the binding of a molecule increases transcription of a gene—are called positive control mechanisms.
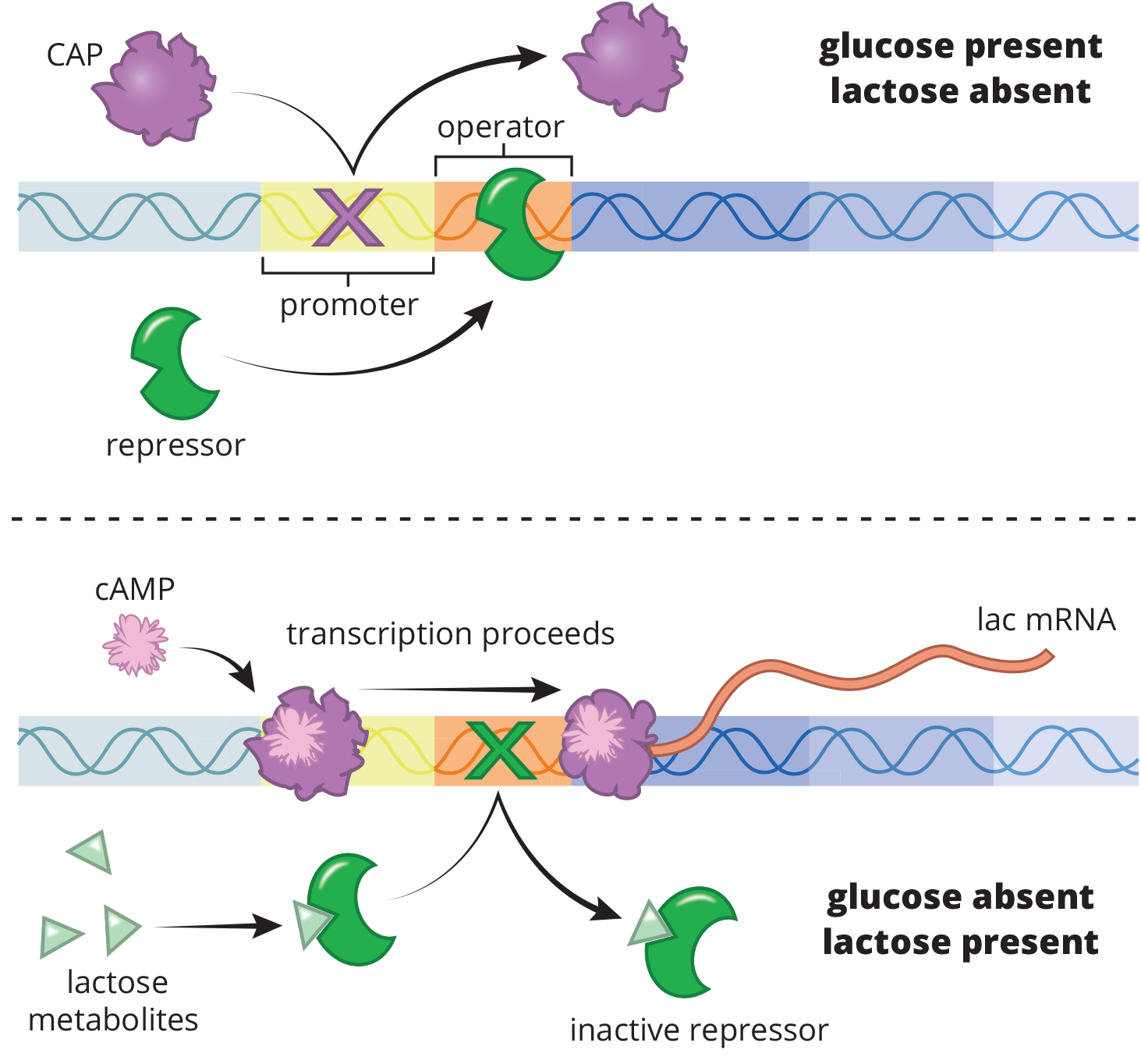
Figure 7.17 The lac Operon An example of an inducible system
Repressible Systems
Repressible systems allow constant production of a protein product. In contrast to the inducible system, the repressor made by the regulator gene is inactive until it binds to a corepressor. This complex then binds the operator site to prevent further transcription, as shown in Figure 7.18. Repressible systems tend to serve as negative feedback; often, the final structural product can serve as a corepressor. Thus, as its levels increase, it can bind the repressor, and the complex will attach to the operator region to prevent further transcription of the same gene.
KEY CONCEPT
Negative control—The binding of a protein to DNA stops transcription. Positive control—The binding of a protein to DNA increases transcription. Inducible system—The system is normally “off” but can be made to turn “on,” given a particular signal. Repressible system—The system is normally “on” but can be made to turn “off,” given a particular signal.
Any combination of control and system are possible; the lac operon is a negative inducible system whereas the trp operon is a negative repressible system.
The trp operon, described above, operates in this way as a negative repressible system. When tryptophan is high in the local environment, it acts as a corepressor. The binding of two molecules of tryptophan to the repressor causes the repressor to bind to the operator site. Thus, the cell turns off its machinery to synthesize its own tryptophan, which is an energetically expensive process because of its easy availability in the environment.
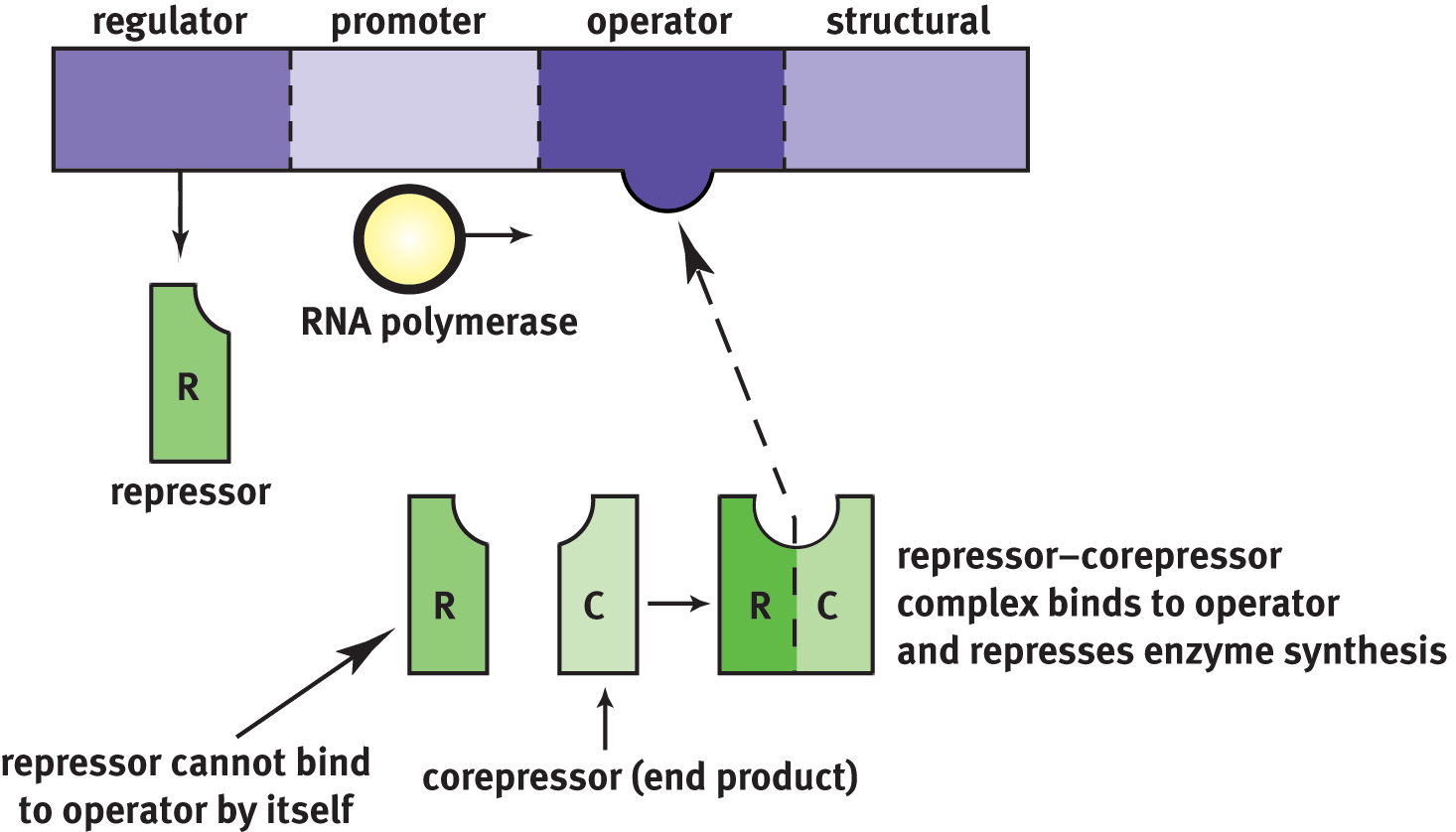
Figure 7.18 Repressible Systems Continually allow gene transcription unless a corepressor binds to the repressor to stop transcription
MCAT CONCEPT CHECK 7.4:
Before you move on, assess your understanding of the material with these questions.
- What type of operon is the trp operon? The lac operon?
- trp: _________________________________
- lac: ________________________________
- From 5′ to 3′, what are the components of the operon, and what are their roles?
Component Role
- What is a positive control system? What is a negative control system?
- Positive control system: _________________________________
- Negative control system: _________________________________
7.5 Control of Gene Expression in Eukaryotes
LEARNING OBJECTIVES
After Chapter 7.5, you will be able to:
- Identify the different mechanisms that can be used to regulate or amplify the expression of a gene
- Predict how histone and DNA modification will affect the ratio of heterochromatin to euchromatin
Genomic expression in eukaryotes is considerably more complex than in prokaryotes, and you will need to know those differences for Test Day. Regulation of gene expression is an essential feature that helps in maintaining the overall functionality of cells. In addition to basic transcriptional enzymes, however, there are a host of other regulatory proteins that play a prominent role in controlling gene expression levels in the cell.
Transcription Factors
Transcription factors are transcription-activating proteins that search the DNA looking for specific DNA-binding motifs. Transcription factors tend to have two recognizable domains: a DNA-binding domain and an activation domain. The DNA-binding domain binds to a specific nucleotide sequence in the promoter region or to a DNA response element (a sequence of DNA that binds only to specific transcription factors) to help in the recruitment of transcriptional machinery. The activation domain allows for the binding of several transcription factors and other important regulatory proteins, such as RNA polymerase and histone acetylases, which function in the remodeling of the chromatin structure.
Gene Amplification
Once the transcription complex is formed, basal (or low-level) transcription can begin and maintain moderate, but adequate, levels of the protein encoded by this gene in the cell. There are times, however, when the expression must be increased, or amplified, in response to specific signals such as hormones, growth factors, and other intracellular conditions. Eukaryotic cells accomplish this through enhancers and gene duplication.
KEY CONCEPT
The DNA regulatory base sequences (such as promoters, enhancers, and response elements) are known as cis regulators because they are in the same vicinity as the gene they control. Transcription factors, however, have to be produced and translocated back to the nucleus; thus they are called trans regulators because they travel through the cell to their point of action.
Enhancers
Response elements outside the normal promoter regions can be recognized by specific transcription factors to enhance transcription levels. Several response elements may be grouped together to form an enhancer, which allows for the control of one gene’s expression by multiple signals. Figure 7.19 demonstrates a eukaryotic example of an enhancer. Signal molecules, such as cyclic AMP (cAMP), cortisol, and estrogen, bind to specific receptors. For the examples given, these receptors are cyclic AMP response element-binding protein (CREB), the glucocorticoid (cortisol) receptor, and the estrogen receptor, respectively; all are transcription factors that bind to their respective response elements within the enhancer. Other proteins are involved in this process, but are outside the scope of the MCAT. Note that the large distance between the enhancer and promoter regions for a given gene means that DNA often must bend into a hairpin loop to bring these elements together spatially.
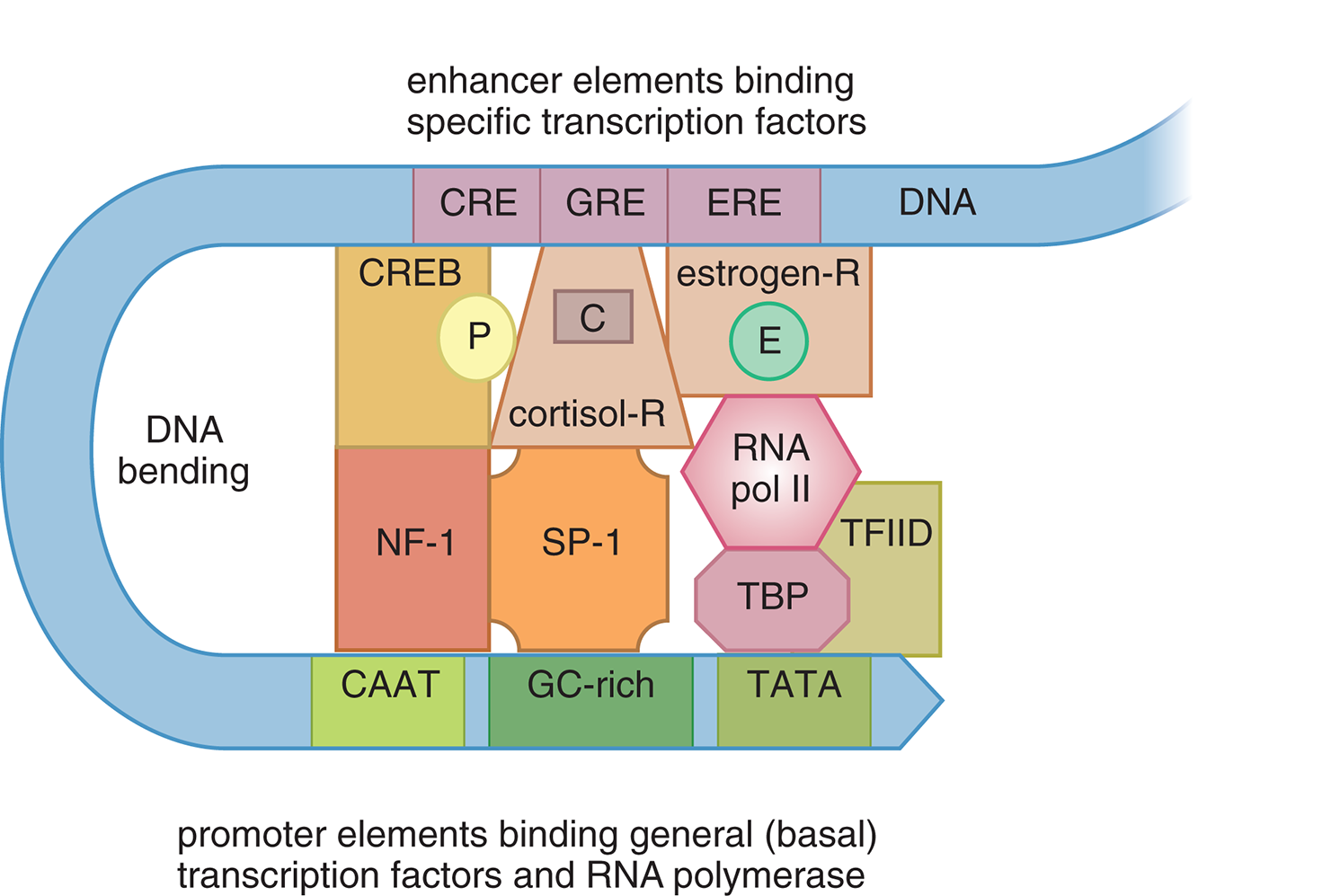
Figure 7.19 Stimulation of Transcription by an Enhancer and Its Associated Transcription Factors
Enhancer regions in the DNA can be up to 1000 base pairs away from the gene they regulate and can even be located within an intron, or noncoding region, of the gene. They differ from upstream promoter elements in their locations because upstream promoter elements must be within 25 bases of the start of a gene. By utilizing enhancer regions, genes have an increased likelihood to be amplified because of the variety of signals that can increase transcription levels.
Gene Duplication
Cells can also increase the expression of a gene product by duplicating the relevant gene. Genes can be duplicated in series on the same chromosome, yielding many copies in a row of the same genetic information. Genes can also be duplicated in parallel by opening the gene with helicases and permitting DNA replication only of that gene; cells can continue replicating the gene until hundreds of copies of the gene exist in parallel on the same chromosome.
Regulation of Chromatin Structure
In eukaryotic cells, DNA is packaged in the nucleus as chromatin, which requires chromatin remodeling to allow transcription factors and the transcriptional machinery easier access to the DNA. Heterochromatin is tightly coiled DNA that appears dark under the microscope; its tight coiling makes it inaccessible to the transcription machinery, so these genes are inactive. Euchromatin, on the other hand, is looser and appears light under the microscope; the transcription machinery can access the genes of interest, so these genes are active. Remodeling of the chromatin structures regulates gene expression levels in the cell.
Histone Acetylation
Transcription factors that bind to the DNA can recruit other coactivators such as histone acetylases. These proteins are involved in chromatin remodeling, as shown in Figure 7.20, because they acetylate lysine residues found in the amino terminal tail regions of histone proteins. Acetylation of histone proteins decreases the positive charge on lysine residues and weakens the interaction of the histone with DNA, resulting in an open chromatin conformation that allows for easier access of the transcriptional machinery to the DNA.
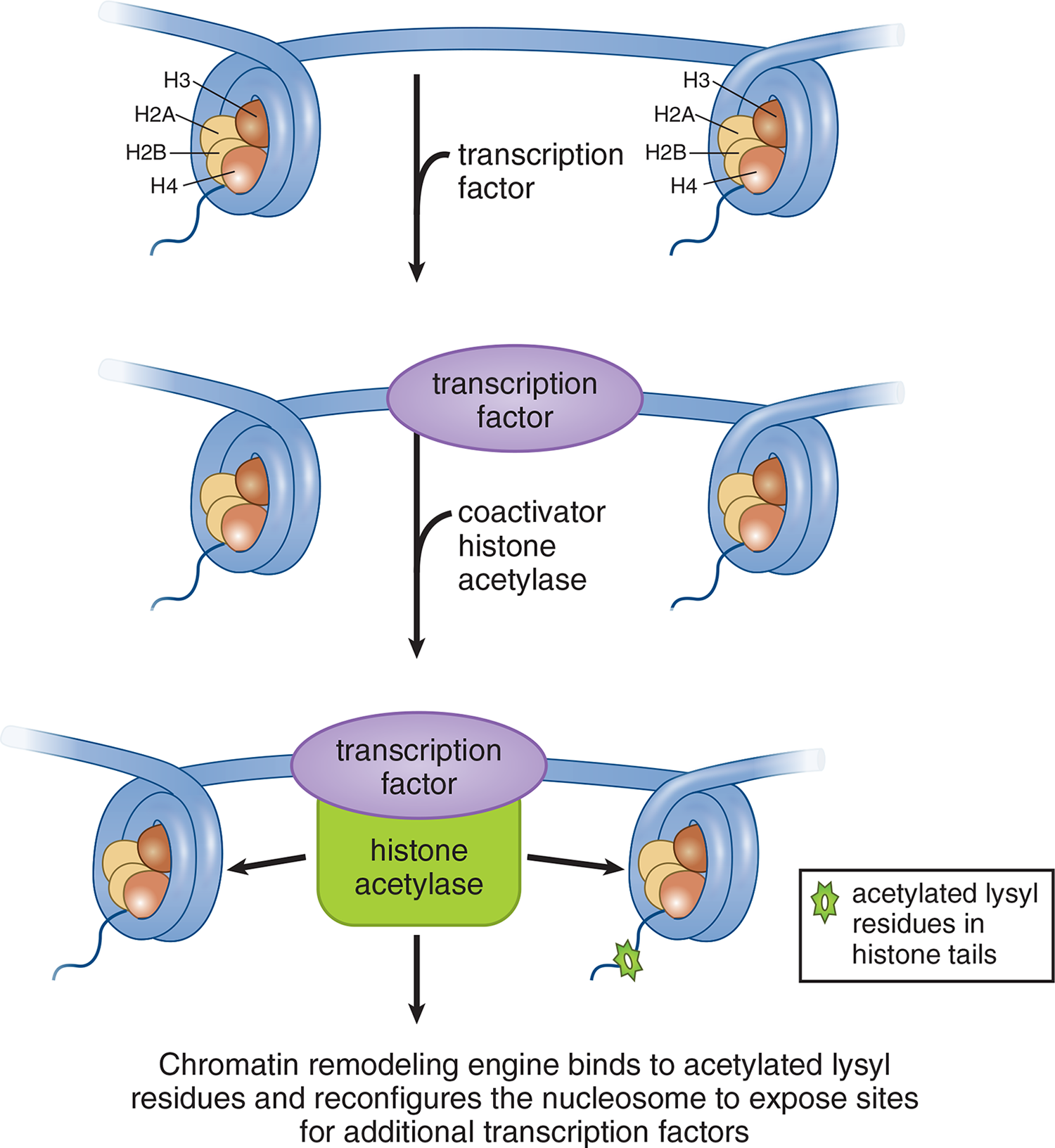
Figure 7.20 Chromatin Remodeling by Acetylation Increases space between histones, allowing better access to DNA for transcription factors
Specific patterns of histone acetylation can lead to increased gene expression levels. On the other hand, gene silencing can occur just as easily with chromatin remodeling. Histone deacetylases are proteins that function to remove acetyl groups from histones, which results in a closed chromatin conformation and overall decrease in gene expression levels in the cell.
DNA Methylation
DNA methylation is also involved in chromatin remodeling and regulation of gene expression levels in the cell. DNA methylases add methyl groups to cytosine and adenine nucleotides, with an example of methylated DNA shown in Figure 7.21; methylation of genes is often linked with the silencing of gene expression. During development, methylation plays an important role in silencing genes that no longer need to be activated. Heterochromatin regions of the DNA are much more heavily methylated, hindering access of the transcriptional machinery to the DNA.
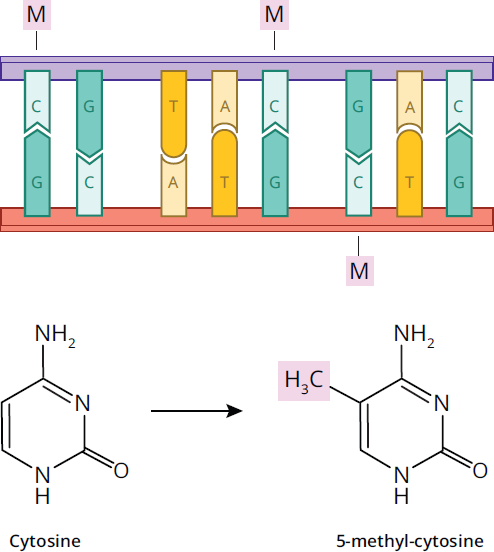
Figure 7.21 DNA Methylation
MCAT CONCEPT CHECK 7.5:
Before you move on, assess your understanding of the material with these questions.
- In an enhancer, what are the differences between signal molecules, transcription factors, and response elements?
______________________________________
- By what histone and DNA modifications can genes be silenced in eukaryotic cells? Would these processes increase the proportion of heterochromatin or euchromatin?
________________________________________
Conclusion
To carry out the functions of life, we must produce around 100,000 different proteins using our 20,000–25,000 available genes. Every protein in the biosphere is produced through the central dogma of molecular biology, as genes in DNA are transcribed into mRNA and then translated into a functional protein. This is a complex, highly regulated process in both prokaryotes and eukaryotes, and questions on transcription and translation, and their regulation, are frequent on the MCAT.
The last two chapters focused on the important roles played by many organelles in the cell, including the nucleus, nucleolus, ribosome, rough endoplasmic reticulum, and Golgi apparatus. After secreted proteins such as hormones and digestive enzymes are produced, they make their way to the plasma membrane for exocytosis. It is this last stop that we’ll examine in the next chapter: the structure, function, and biochemistry of biological membranes.
GO ONLINE!
You've reviewed the content, now test your knowledge and critical thinking skills by completing a test-like passage set in your online resources!
CONCEPT SUMMARY
The Genetic Code
- The central dogma states that DNA is transcribed to RNA, which is translated to protein.
- A degenerate code allows multiple codons to encode for the same amino acid.
- Initiation (start) codon: AUG
- Termination (stop) codons: UAA, UGA, UAG
- Redundancy and wobble (third base in the codon) allows mutations to occur without effects in the protein.
- Point mutations can cause:
- Silent mutations with no effect on protein synthesis.
- Nonsense (truncation) mutations that produce a premature stop codon.
- Missense mutations that produce a codon that codes for a different amino acid.
- Frameshift mutations result from nucleotide addition or deletion, and change the reading frame of subsequent codons.
- RNA is structurally similar to DNA except:
- Substitution of a ribose sugar for deoxyribose
- Substitution of uracil for thymine
- It is single-stranded instead of double-stranded
- There are three types of RNA with separate jobs in transcription:
- Messenger RNA (mRNA) carries the message from DNA in the nucleus via transcription of the gene; it travels into the cytoplasm to be translated.
- Transfer RNA (tRNA) brings in amino acids and recognizes the codon on the mRNA using its anticodon.
- Ribosomal RNA (rRNA) makes up the ribosome and is enzymatically active.
Transcription
-
- Helicase unwinds the DNA double helix.
- RNA polymerase II binds to the TATA box within the promoter region of the gene (25 base pairs upstream from first transcribed base).
- hnRNA is synthesized from the DNA template (antisense) strand.
- Posttranscriptional modifications include:
- A 7-methylguanylate triphosphate cap is added to the 5′ end.
- A polyadenosyl (poly-A) tail is added to the 3′ end.
- Splicing is done by snRNA and snRNPs in the spliceosome; introns are removed in a lariat structure, and exons are ligated together.
- Prokaryotic cells can increase the variability of gene products from one transcript through polycistronic genes (in which starting transcription in different sites within the gene leads to different gene products).
- Eukaryotic cells can increase variability of gene products through alternative splicing (combining different exons in a modular fashion to acquire different gene products).
Translation
- tRNA translates the codon into the correct amino acid.
- Ribosomes are the factories where translation (protein synthesis) occurs.
- There are three stages of translation.
- Initiation in prokaryotes occurs when the 30S ribosome attaches to the Shine–Dalgarno sequence and scans for a start codon; it lays down N-formylmethionine in the P site of the ribosome.
- Initiation in eukaryotes occurs when the 40S ribosome attaches to the 5′ cap and scans for a start codon; it lays down methionine in the P site of the ribosome.
- Elongation involves the addition of a new aminoacyl-tRNA into the A site of the ribosome and transfer of the growing polypeptide chain from the tRNA in the P site to the tRNA in the A site. The now uncharged tRNA pauses in the E site before exiting the ribosome.
- Termination occurs when the codon in the A site is a stop codon; a release factor places a water molecule on the polypeptide chain and thus releases the protein.
- Initiation, elongation, and release factors help with each step in recruitment and assembly/disassembly of the ribosome.
- Posttranslational modifications include:
- Folding by chaperones
- Formation of quaternary structure
- Cleavage of proteins or signal sequences
- Covalent addition of other biomolecules (phosphorylation, carboxylation, glycosylation, prenylation)
Control of Gene Expression in Prokaryotes
- The Jacob–Monod model of repressors and activators explains how operons work.
- Operons are inducible or repressible clusters of genes transcribed as a single mRNA.
- Inducible systems (such as the lac operon) are bonded to a repressor under normal conditions; they can be turned on by an inducer pulling the repressor from the operator site.
- Repressible systems (such as the trp operon) are transcribed under normal conditions; they can be turned off by a corepressor coupling with the repressor and the binding of this complex to the operator site.
Control of Gene Expression in Eukaryotes
- Transcription factors search for promoter and enhancer regions in the DNA.
- Promoters are within 25 base pairs of the transcription start site.
- Enhancers are more than 25 base pairs away from the transcription start site.
- Modification of chromatin structure affects the ability of transcriptional enzymes to access the DNA through histone acetylation (increases accessibility) or DNA methylation (decreases accessibility).
ANSWERS TO CONCEPT CHECKS
**7.1**
- mRNA carries information from DNA by traveling from the nucleus (where it is transcribed) to the cytoplasm (where it is translated). tRNA translates nucleic acids to amino acids by pairing its anticodon with mRNA codons; it is charged with an amino acid, which can be added to the growing peptide chain. rRNA forms much of the structural and catalytic component of the ribosome, and acts as a ribozyme to create peptide bonds between amino acids.
-
- GAT: mRNA codon = AUC; Isoleucine (Ile)
- ATT: mRNA codon = AAU; Asparagine (Asn)
- CGC: mRNA codon = GCG; Alanine (Ala)
- CCA: mRNA codon = UGG; Tryptophan (Trp)
- The start codon is AUG, which codes for methionine; the stop codons are UAA, UGA, and UAG.
- Wobble refers to the fact that the third base in a codon often plays no role in determining which amino acid is translated from that codon. For example, any codon starting with “CC” codes for proline, regardless of which base is in the third (wobble) position. This is protective because mutations in the wobble position will not have any effect on the protein translated from that gene.
-
Type of Mutation Change in DNA Sequence Effect on Encoded Protein Silent (degenerate) Substitution of bases in the wobble position, introns, or noncoding DNA No change observed
Missense Substitution of one base, creating an mRNA codon that matches a different amino acid One amino acid is changed in the protein; variable effects on function depending on specific change
Nonsense Substitution of one base, creating a stop codon Early truncation of protein; variable effects on function, but usually more severe than missense mutations
Frameshift Insertion or deletion of bases, creating a shift in the reading frame of the mRNA Change in most amino acids after the site of insertion or deletion; usually the most severe of the types listed here
**7.2**
- RNA polymerase I synthesizes most rRNA. RNA polymerase II synthesizes mRNA (hnRNA) and snRNA. RNA polymerase III synthesizes tRNA and some rRNA.
- RNA polymerase II binds to the TATA box, which is located within the promoter region of a relevant gene, at about –25.
- The major posttranscriptional modifications are:
- Splicing: removal of introns, joining of exons; uses snRNA and snRNPs in the spliceosome to create a lariat, which is then degraded; exons are ligated together
- 5′ cap: addition of a 7-methylguanylate triphosphate cap to the 5′ end of the transcript
- 3′ poly-A tail: addition of adenosine bases to the 3′ end to protect against degradation
- Alternative splicing is the ability of some genes to use various combinations of exons to create multiple proteins from one hnRNA transcript. This increases protein diversity and allows a species to maximize the number of proteins it can create from a limited number of genes.
**7.3**
- Initiation, elongation, and termination
P site: holds growing polypeptide until peptidyl transferase forms peptide bond and polypeptide is handed to A site E site: transiently holds uncharged tRNA as it exits the ribosome
- A site: binds incoming aminoacyl-tRNA using codon–anticodon pairing
- Posttranslational modifications include proper folding by chaperones, formation of quaternary structure, cleavage of proteins or signal sequences, and addition of other biomolecules (phosphorylation, carboxylation, glycosylation, prenylation).
**7.4**
- The trp operon is a negative repressible system; the lac operon is a negative inducible system.
-
Component Role Regulator gene Transcribed to form repressor protein
Promoter site Site of RNA polymerase binding (similar to promoters in eukaryotes)
Operator site Binding site for repressor protein
Structural gene The gene of interest; its transcription is dependent on the repressor being absent from the operator site
- Positive control systems require the binding of a protein to the operator site to increase transcription. Negative control systems require the binding of a protein to the operator site to decrease transcription.
**7.5**
- Signal molecules include steroid hormones and second messengers, which bind to their receptors in the nucleus. These receptors are transcription factors that use their DNA-binding domain to attach to a particular sequence in DNA called a response element. Once bonded to the response element, these transcription factors can then promote increased expression of the relevant gene.
- Histone deacetylation and DNA methylation will both downregulate the transcription of a gene. These processes allow the relevant DNA to be clumped more tightly, increasing the proportion of heterochromatin.
SCIENCE MASTERY ASSESSMENT EXPLANATIONS
1. D
Peptidyl transferase is an enzyme that catalyzes the formation of a peptide bond between the incoming amino acid in the A site and the growing polypeptide chain in the P site. Initiation and elongation factors help transport charged tRNA molecules into the ribosome and advance the ribosome down the mRNA transcript, as in (A) and (B). Chaperones maintain a protein’s three-dimensional shape as it is formed, as in (C).
2. C
The promoter functions as a recruitment site for RNA polymerase and is required for gene expression. A mutation in this region would hinder RNA polymerase recruitment, resulting in a decrease in gene expression. These observations support (C) as the correct answer. By contrast, mutations in the operator and regulatory genes would render the repressor less able to block transcription, leading to the opposite effect, i.e. an increase in lactase expression. This result eliminates both (A) and (B). Finally, mutations in the structural genes could lead to the production of a nonfunctional enzyme, but are unlikely to affect gene expression, eliminating (D).
3. A
Topoisomerases, such as prokaryotic DNA gyrase, are involved in DNA replication and mRNA synthesis (transcription). DNA gyrase is a type of topoisomerase that enhances the action of helicase enzymes by the introduction of negative supercoils into the DNA molecule. These negative supercoils facilitate DNA replication by keeping the strands separated and untangled.
4. C
There are four different codons for valine: GUU, GUC, GUA, and GUG. Through base-pairing, we can determine that the proper anticodon must end with “AC.” Remember that the codon and anticodon are antiparallel to each other, and that nucleic acids are always written 5′ → 3′ on the MCAT. Therefore, we are looking for an answer that ends with “AC” (rather than starting with “CA”).
5. A
Specific transcription factors bind to a specific DNA sequence, such as an enhancer, and to RNA polymerase at a single promoter sequence. They enable the RNA polymerase to transcribe the specific gene for that enhancer more efficiently.
6. C
UAG is one of the three known stop codons, so changing tyrosine to a stop codon must be a nonsense (or truncation) mutation.
7. C
In this question, the correct answer is the answer that is notassociated with increased transcription. DNA methylation, (C), is associated with silencing of gene expression. Regions of DNA with poor gene expression, called heterochromatin, are heavily methylated. These modifications significantly decrease the ability of RNA polymerase to access DNA.
8. B
The example given is a sample of repression due to the abundance of a corepressor. In other words, this is a repressible system that is currently blocking transcription. For the trp operon, an abundance of tryptophan in the environment allows for the repressor to bind tryptophan and then to the operator site. This blocks transcription of the genes required to synthesize tryptophan within the cell. The system described is a repressible system; the lac operon is an inducible system, in which an inducer binds to the repressor, thus permitting transcription.
9. C
shRNA (short hairpin RNA) is a useful biotechnology tool used in RNA interference. It is not, however, produced in the nucleus for use in the spliceosome. It targets mRNA to be degraded in the cytoplasm; it is not utilized in splicing of the hnRNA (heterogeneous nuclear RNA). snRNA (small nuclear RNA) and snRNPs (small nuclear ribonucleoproteins), however, do bind to the hnRNA to induce splicing.
10. A
In this table, we are given the sequence of the sense (coding) DNA strand. This will be identical to the mRNA transcript, except all thymine nucleotides will be replaced with uracil. With the deletion of these three bases, codon 507 changes from AUC to AUU in the transcript; these both code for isoleucine due to wobble. However, codon 508 (UUU in the transcript) has been lost. UUU codes for phenylalanine. The C-terminus sequence will remain unchanged because the deletion of three bases (exactly one codon) will not throw off the reading frame. For reference, the mutant reading frames would be:
(Note: refer back to Figure 7.6 for a table of the genetic code)
AUC AUU GGU GUU UCC
11. B
The intron will not be a part of the final, processed mRNA, and the untranslated regions of the mRNA will not be turned into amino acids. Translation will begin with codon 1 (which would be AUG). Because there are 150 amino acids, we can surmise that there will be 151 codons. Each codon will use 3 nucleotides, so 150 × 3 = 450 because codon 151 will be the stop codon.
12. B
Peptidyl transferase connects the incoming amino terminal to the previous carboxyl terminal; the only functional group listed here with a carbonyl and amino group is the amide. Peptide bonds are thus amide linkages, and the correct answer is (B).
13. A
RNA polymerase I in eukaryotes is found in the nucleolus and is in charge of transcribing most of the rRNA for use during ribosomal creation. RNA polymerase II is responsible for hnRNA and snRNA. RNA polymerase III is responsible for tRNA and the 5S rRNA.
14. B
To answer this question correctly, we must remember that mRNA will be antiparallel to DNA. Our answer should be 5′ to 3′ mRNA, with the 5′ end complementary to the 3′ end of the DNA that is being transcribed. Thus, the mRNA transcribed from this strand will be 5′—GGAUGUCUCAAAGA—3′. mRNA contains uracil, rather than thymine.
15. B
The mRNA produced has the same structure as the sense strand of DNA (with uracils instead of thymines). Because bonding of nucleic acids is always complementary but antiparallel, the antisense strand of mRNA would be the one that binds to the produced mRNA, creating double-stranded RNA that is then degraded once found in the cytoplasm.
GO ONLINE
Consult your online resources for additional practice.
SHARED CONCEPTS
- Biochemistry Chapter 1
- Amino Acids, Peptides, and Proteins
- Biochemistry Chapter 2
- Enzymes
- Biochemistry Chapter 6
- DNA and Biotechnology
- Biology Chapter 1
- The Cell
- Biology Chapter 3
- Embryogenesis and Development
- Biology Chapter 12
- Genetics and Evolution