U1 · Microorganisms & microbiology
- Three domains
- Bacteria, Archaea, Eukarya — Carl Woese's rRNA-based tree (1977). Archaea are sister to Eukarya, not Bacteria.
- Koch's postulates
- (1) Microbe present in disease, absent in healthy. (2) Isolated in pure culture. (3) Causes disease when introduced. (4) Re-isolated from new host. Modernized for non-culturable + viral pathogens.
- Pasteur
- Disproved spontaneous generation; pioneered fermentation, pasteurization, vaccines (rabies, anthrax).
- Microbial sizes
- Most bacteria 1-5 µm; viruses 20-300 nm; Mycoplasma ~0.3 µm; Thiomargarita up to 750 µm (visible!).
- Gram stain
- Crystal violet → iodine mordant → ethanol decolorize → safranin counter. Gram-positive retain purple (thick peptidoglycan); Gram-negative pink (thin PG + outer membrane).
U2 · Microbial cell structure


- Peptidoglycan
- Murein. Polymer of NAG-NAM cross-linked by tetrapeptides. Bacteria only — Archaea have pseudopeptidoglycan (NAG-NAT) or other walls.
- Teichoic acids
- Glycerol/ribitol-phosphate polymers anchored in Gram-positive cell wall; antigen + ion homeostasis.
- LPS (lipopolysaccharide)
- Gram-negative outer-membrane outer leaflet; Lipid A (endotoxin) + core + O-antigen. Triggers TLR4 / septic shock.
- Periplasm
- Gel-filled space between inner + outer membranes in Gram-negatives; site of secreted enzymes (β-lactamases).
- Capsule / S-layer
- Outer polysaccharide capsule (virulence factor; resists phagocytosis). S-layer = paracrystalline protein layer (especially Archaea).
- Bacterial flagellum
- Helical filament rotated by H⁺- or Na⁺-driven motor. Run-and-tumble chemotaxis. Right-handed helix in E. coli.
- Pili / fimbriae
- Surface filaments. Type IV pili for twitching motility + biofilm; F-pilus for conjugation.
- Endospore
- Dormant heat/chemical-resistant structure of Bacillus, Clostridium. Dipicolinic acid + Ca²⁺ in core. Triggered by starvation.
- Archaeal membrane
- Ether-linked isoprenoid lipids (vs ester-linked fatty acids in Bacteria/Eukarya); often monolayer in hyperthermophiles.
U3 · Microbial growth
- Binary fission
- One cell → two via FtsZ ring at midcell + septum formation.
- Generation time
- Time for population to double. E. coli ~20 min in rich media; M. tuberculosis ~24 hr.
- Growth curve
- Lag → exponential (log) → stationary → death. N = N₀ × 2^n.
- Continuous culture (chemostat)
- Steady-state growth at controllable rate by limiting one nutrient.
- Optimal temperature classes
- Psychrophile (<15°C), mesophile (20-45°C), thermophile (45-80°C), hyperthermophile (>80°C).
- pH classes
- Acidophile (<5), neutrophile (5-8), alkaliphile (>9). Stomach H. pylori = neutrophile that buffers via urease.
- Oxygen classes
- Obligate aerobe, facultative anaerobe (best in O₂ but tolerates), microaerophile (low O₂), aerotolerant anaerobe, obligate anaerobe.
- Water activity (a_w)
- Free water available for growth. Halophiles thrive at low a_w (high salt).
U4 · Metabolism — energy & biosynthesis
- Catabolism vs anabolism
- Catabolism breaks down to make energy (ATP) + reducing power (NADH/NADPH). Anabolism builds biomass.
- Chemoorganotroph
- Organic compounds = energy + carbon source. Most cultured bacteria.
- Chemolithotroph
- Inorganic e⁻ donors (H₂, NH₃, NO₂⁻, H₂S, Fe²⁺) for energy. CO₂ fixation for carbon (often).
- Phototroph
- Light-driven energy. Oxygenic (cyanobacteria, plants) split H₂O → O₂. Anoxygenic photosynthesis uses H₂S, etc.
- Fermentation
- Substrate-level phosphorylation only; organic e⁻ acceptor. Products: lactate, ethanol, butyrate, acetate, etc.
- Aerobic respiration
- Glycolysis + TCA + ETC; O₂ as terminal e⁻ acceptor → H₂O. Highest ATP yield.
- Anaerobic respiration
- ETC with non-O₂ acceptor: NO₃⁻ (denitrification), SO₄²⁻ (Desulfovibrio), Fe³⁺ (Geobacter), CO₂ (methanogens), fumarate.
- Substrate-level vs oxidative phosphorylation
- SLP: phosphate from substrate to ADP (e.g., glycolysis step 7). Oxidative: ATP synthase using H⁺ gradient.
U5 · Metabolic diversity
- Calvin cycle
- RuBisCO fixes CO₂ in autotrophs. Found in cyanobacteria, plants, many lithotrophs.
- Reverse TCA / 3-hydroxypropionate / Wood-Ljungdahl
- Alternative CO₂ fixation pathways used by various Archaea + Bacteria. Wood-Ljungdahl in acetogens + methanogens.
- Methanogenesis
- Archaea-only (Euryarchaeota): CO₂ + H₂ → CH₄ + H₂O OR acetate → CO₂ + CH₄. Coenzyme F420, methyl-CoM.
- Sulfate reduction
- Desulfovibrio: SO₄²⁻ → H₂S using organic H donors. Marine sediments.
- Nitrification
- NH₃ → NO₂⁻ (Nitrosomonas) → NO₃⁻ (Nitrobacter). Two genera typically.
- Denitrification
- NO₃⁻ → NO₂⁻ → NO → N₂O → N₂. Anaerobic respiration; major N-cycle return flux.
- Nitrogen fixation
- N₂ → 2 NH₃ via nitrogenase (Mo-Fe protein). High ATP cost. Rhizobium + legumes; cyanobacteria; free-living.
U6 · Microbial genomes
- Bacterial genome
- Usually one circular chromosome in cytoplasm (nucleoid); supercoiled by gyrase + Topo I. ~few Mbp typically.
- Plasmid
- Extrachromosomal circular dsDNA; replicates independently. Carries optional genes (resistance, virulence, conjugation).
- Origin of replication (oriC)
- Site where replication initiates; DnaA binds DnaA boxes; helicase loads.
- Replication enzymes
- DnaB helicase, DnaG primase, DNA pol III holoenzyme (replicative), DNA pol I (Okazaki primer removal), ligase.
- Topoisomerases
- Type I single-strand cuts (Topo I); Type II double-strand cuts (DNA gyrase introduces neg supercoils, Topo IV decatenates daughters). Quinolone target.
- Transposable elements
- IS elements (insertion sequences) + composite transposons. Carry transposase + flanked by inverted repeats.
- Pan-genome
- Core genes shared across all strains + accessory genes in some. Highlights HGT.
U7 · Horizontal gene transfer

- Transformation
- Uptake of naked DNA from environment by competent cells (natural: Streptococcus pneumoniae, Bacillus subtilis; artificial: heat shock, electroporation).
- Transduction
- Phage-mediated DNA transfer. Generalized: any host DNA packaged. Specialized: prophage excision picks up flanking genes.
- Conjugation
- Cell-to-cell DNA transfer through pilus. F+ donor transfers F plasmid; Hfr integrates F into chromosome and transfers chromosomal genes.
- Restriction-modification
- Defense system: restriction enzyme cuts unmethylated foreign DNA; methylase protects host. Type II = molecular cloning workhorse.
- CRISPR-Cas
- Adaptive bacterial immunity. Spacers (records of past invaders) → crRNA + Cas9/12/13 → cleaves matching foreign DNA. Now genome-editing tool.
- Site-specific recombination
- Integrase between attP + attB (e.g., λ phage); recombines specific sites only.
U8 · Regulation of gene expression
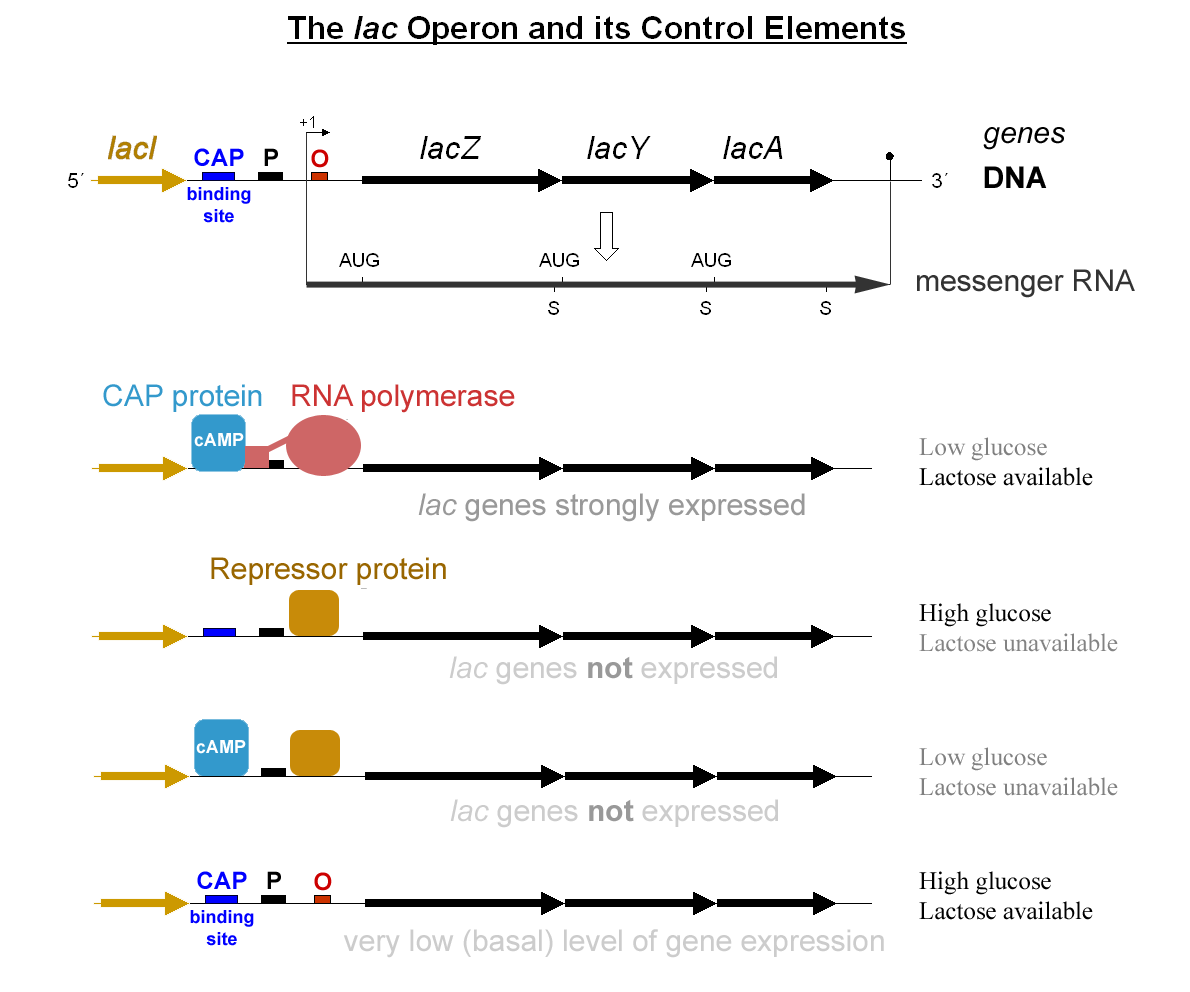
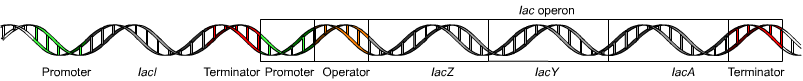
- Operon
- Polycistronic mRNA from one promoter; coordinately regulated. Prokaryote-specific.
- lac operon
- Catabolic. Repressor (LacI) blocks transcription unless allolactose binds. CAP+cAMP activates when glucose low. Inducible operon.
- trp operon
- Anabolic. Repressor + tryptophan (corepressor) blocks transcription. Repressible operon. Also attenuation: leader peptide ribosome stalling.
- Sigma (σ) factor
- Subunit of bacterial RNA polymerase that recognizes specific promoter classes. E. coli: σ70 (housekeeping), σ32 (heat shock), σS (stationary), σ54 (N), σF (flagella), σE (extracytoplasmic stress).
- Two-component system
- Membrane sensor histidine kinase autophosphorylates; transfers phosphate to cytoplasmic response regulator → DNA binding. Bacteria-wide environmental sensing.
- Quorum sensing
- Cell-density-dependent gene regulation via diffusible autoinducers. AHLs (LuxI/LuxR) in Gram-negative; AIPs in Gram-positive. Controls biofilm + virulence.
- Riboswitch
- 5'-UTR mRNA element binds metabolite directly → conformational change → premature termination or RBS occlusion. No protein needed.
- sRNA / antisense RNA
- Trans-encoded small RNAs (often Hfq-dependent) bind mRNA → block translation or recruit RNase E.
U9 · Viruses
- Virus
- Obligate intracellular acellular agent. Genome (DNA/RNA, ds/ss) + capsid (± envelope).
- Capsid symmetries
- Helical (TMV), icosahedral (T=1, 3, 7…), complex (T4 phage with head + tail).
- Lytic cycle
- Adsorption → penetration → replication → assembly → lysis (release).
- Lysogenic cycle
- Phage integrates (prophage) into host chromosome; replicates passively until induction (UV, etc.) → lytic cycle.
- One-step growth curve
- Plot of phage titer over time after synchronous infection; eclipse → latent → burst.
- Retrovirus
- +ssRNA → reverse transcriptase → dsDNA → integrase → provirus. HIV.
- Baltimore classification
- I dsDNA, II ssDNA, III dsRNA, IV +ssRNA, V −ssRNA, VI +ssRNA-RT (retro), VII dsDNA-RT (HBV).
- CRISPR vs phages
- Spacer acquisition from phage; later phages with matching protospacer cleaved.
U10 · Bacterial diversity
- Proteobacteria
- Largest, most diverse Gram-negative phylum. α (Rhizobium, Rickettsia), β, γ (E. coli, Pseudomonas, Vibrio), δ (Desulfovibrio, myxobacteria), ε (Helicobacter, Campylobacter).
- Firmicutes
- Gram-positive low-GC. Bacillus, Clostridium, Staphylococcus, Streptococcus, Lactobacillus.
- Actinobacteria
- Gram-positive high-GC. Mycobacterium (TB), Streptomyces (antibiotic source), Corynebacterium (diphtheria).
- Cyanobacteria
- Oxygenic photosynthesis; ancestor of chloroplasts. Heterocysts fix N₂ in some.
- Spirochaetes
- Long thin spirals with periplasmic flagella. Treponema (syphilis), Borrelia (Lyme), Leptospira.
- Bacteroidetes
- Anaerobic gut residents — gut microbiome dominant.
U11 · Archaea
- Distinguishing features
- Ether-linked isoprenoid membrane lipids; pseudopeptidoglycan or S-layer walls; unique RNA pol with 8-12 subunits (more like eukaryotic); histones in some.
- Crenarchaeota
- Mostly extreme thermophiles + acidophiles. Sulfolobus, Pyrococcus, Thermoproteus.
- Euryarchaeota
- Methanogens (Methanocaldococcus), extreme halophiles (Halobacterium), thermoacidophiles (Thermoplasma).
- Thaumarchaeota
- Ammonia oxidizers in marine + soil; major component of microbial N cycling.
- Halophiles
- Require >1.5 M NaCl. Use compatible solutes (K⁺, glycine betaine) for osmotic balance. Halobacterium uses bacteriorhodopsin proton pump.
- Hyperthermophiles
- Optimum T > 80°C. Pyrolobus fumarii grows at 113°C. Hydrothermal vents.
U12 · Microbial ecology
- Microbiome
- Microbial community in a defined habitat (gut, skin, soil, ocean). Studied via 16S rRNA + shotgun metagenomics.
- 16S rRNA
- Universal phylogenetic marker; conserved + variable regions allow species ID without culture.
- Biofilm
- Surface-attached community in EPS matrix. Stages: attachment → microcolony → maturation → dispersal. Highly resistant to antibiotics + immune attack.
- Quorum sensing in biofilms
- AHL or AIP signals reach threshold density → coordinated gene expression (virulence, EPS, dispersion). Pseudomonas LasR + RhlR.
- Carbon cycle role
- Decomposers mineralize organic C → CO₂. Anaerobes ferment + methanogens produce CH₄ in wetlands + cattle.
- Nitrogen cycle
- Fixation (Rhizobium, cyanobacteria) → ammonification → nitrification → denitrification. Bacterial monopoly aside from lightning + Haber-Bosch.
- Sulfur cycle
- Sulfate reducers (Desulfovibrio) → H₂S; sulfide oxidizers (Beggiatoa, Thiobacillus) → SO₄²⁻.
- Symbioses
- Rumen microbiome digests cellulose; root nodules fix N₂; insect endosymbionts supply vitamins.
U13 · Pathogenesis — Rowen focus
- Virulence factor
- Microbial product that contributes to disease: toxins, capsules, adhesins, secretion systems, immune evasion.
- Adhesion + colonization
- Pili, fimbriae, surface adhesins bind host receptors. E. coli P-pili to UTI; Vibrio cholerae TCP to gut.
- Exotoxins
- Secreted proteins. Diphtheria (ADP-ribosylates EF-2), cholera (constitutive Gαs cAMP), botulinum + tetanus (cleave SNAREs), Shiga (depurinates 28S rRNA).
- Endotoxin (LPS)
- Lipid A of Gram-negative outer membrane. Released on cell lysis; TLR4 → cytokine storm → septic shock.
- Type III secretion system (T3SS) [Rowen specialty]
- Needle-like injectisome that delivers effectors directly into host cell cytosol. Yersinia, Salmonella, Pseudomonas aeruginosa, EPEC.
- Type IV secretion (T4SS)
- Conjugation-related secretion; Helicobacter CagA, Agrobacterium T-DNA, Legionella Dot/Icm.
- Type VI secretion (T6SS)
- Bacteriophage-derived; injects toxins into competing bacteria + sometimes host.
- Pseudomonas aeruginosa [Rowen's organism]
- Opportunistic pathogen; biofilms in CF lungs. Mucoid conversion (alginate overproduction) marks chronic infection. Multidrug resistance via efflux + porin loss + β-lactamases.
- Mucoid conversion in P. aeruginosa
- Mutation in mucA (anti-σ factor) releases σ22 (AlgT/U) → activates alginate biosynthesis genes → mucoid phenotype. Hallmark of CF lung adaptation.
- Biofilm resistance
- Slow growth, persister cells, EPS diffusion barrier, altered gene expression, quorum sensing → 100-1000× more antibiotic-tolerant.
U14 · Antimicrobials & resistance
- Cell wall inhibitors
- β-lactams (penicillin, cephalosporin, carbapenem) inhibit transpeptidase / PBPs. Vancomycin binds D-Ala-D-Ala. Bactericidal.
- Protein synthesis inhibitors
- 30S: aminoglycosides (streptomycin, gentamicin), tetracyclines. 50S: macrolides (erythromycin), chloramphenicol, lincosamides, oxazolidinones (linezolid).
- Nucleic acid inhibitors
- Quinolones (ciprofloxacin) inhibit gyrase/Topo IV. Rifampin binds RpoB → blocks transcription. Metronidazole damages DNA in anaerobes.
- Folate antagonists
- Sulfonamides + trimethoprim block folate synthesis (sequential steps); synergistic combo (TMP-SMX).
- MIC
- Minimum inhibitory concentration: lowest [drug] that prevents visible growth. Standard antibiotic susceptibility metric.
- Resistance mechanisms
- (1) Enzymatic inactivation (β-lactamase, AME). (2) Efflux pumps (tetracycline, fluoroquinolone). (3) Target modification (PBP2a in MRSA, ribosomal methylation, DNA gyrase mutation). (4) Reduced uptake (porin loss).
- MRSA, VRE, ESBL, CRE
- MRSA = methicillin-resistant S. aureus (mecA → PBP2a). VRE = vancomycin-resistant Enterococcus (D-Ala-D-Lac). ESBL = extended-spectrum β-lactamase. CRE = carbapenem-resistant Enterobacterales (KPC, NDM-1).
- Horizontal spread of resistance
- Plasmids, integrons, transposons carry multiple resistance genes — co-selection drives MDR.
U15 · Applied / clinical microbiology
- Industrial fermentation
- Lactic acid (yogurt, sauerkraut), ethanol (beer, fuel), penicillin (Penicillium), insulin (recombinant E. coli), citric acid (Aspergillus).
- Bioremediation
- Microbes degrade pollutants: Pseudomonas on hydrocarbons, Geobacter on uranium, Dehalococcoides on chlorinated solvents.
- Vaccine types
- Live attenuated (MMR, OPV), inactivated (IPV, flu), subunit (HBV), toxoid (DT, TT), conjugate (Hib, pneumococcal), mRNA (COVID).
- Sterilization vs disinfection
- Sterilization eliminates ALL microbes (autoclave 121°C 15 psi 15 min). Disinfection reduces pathogens on inanimate surfaces. Antisepsis on living tissue.
- Lab diagnosis
- Culture + Gram stain + biochemical tests + MALDI-TOF + 16S rRNA + PCR. Rapid Ag/Ab tests for select pathogens.
Rowen-targeted exam tips
- Know P. aeruginosa in detail: Gram-negative, motile, biofilm-forming, mucoid in CF (mucA → AlgT → alginate), MDR via efflux + porin + β-lactamases.
- Be ready to diagram T3SS / T4SS / T6SS; know representative pathogens for each.
- For lac vs trp: know inducible vs repressible, allolactose vs Trp as effector, attenuation in trp.
- Differentiate Bacteria vs Archaea: peptidoglycan vs pseudopep, ester vs ether lipids, σ factor vs eukaryotic-like RNA pol.
- Memorize 4 antibiotic classes by target + 3-4 resistance mechanisms with examples (MRSA, VRE, ESBL, CRE).